Abstract
Multidrug and toxic compound extrusion (MATE) transporters contribute to multidrug resistance by coupling the efflux of drugs to the influx of Na+ or H+. Known structures of Na+-coupled, extracellular-facing MATE transporters from the NorM subfamily revealed 12 membrane-spanning segments related by a quasi–two-fold rotational symmetry and a multidrug-binding cavity situated near the membrane surface. Here we report the crystal structure of an H+-coupled MATE transporter from Bacillus halodurans and the DinF subfamily at 3.2-Å resolution, unveiling a surprisingly asymmetric arrangement of 12 transmembrane helices. We also identified a membrane-embedded substrate-binding chamber by combining crystallographic and biochemical analyses. Our studies further suggested a direct competition between H+ and substrate during DinF-mediated transport and implied how a MATE transporter alternates between its extracellular- and intracellular-facing conformations to propel multidrug extrusion. Collectively, our results demonstrated heretofore-unrecognized mechanistic diversity among MATE transporters.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Higgins, C.F. Multiple molecular mechanisms for multidrug resistance transporters. Nature 446, 749–757 (2007).
Fischbach, M.A. & Walsh, C.T. Antibiotics for emerging pathogens. Science 325, 1089–1093 (2009).
Brown, M.H., Paulsen, I.T. & Skurray, R.A. The multidrug efflux protein NorM is a prototype of a new family of transporters. Mol. Microbiol. 31, 394–395 (1999).
Omote, H., Miasa, M., Matsumoto, T., Otsuka, M. & Moroyama, Y. The MATE proteins as fundamental transporters of metabolic and xenobiotic organic cations. Trends Pharmacol. Sci. 27, 587–593 (2006).
Kuroda, T. & Tsuchiya, T. Multidrug efflux transporters in the MATE family. Biochim. Biophys. Acta 1794, 763–768 (2009).
Morita, Y., Kataoka, A., Shiota, S., Mizushima, T. & Tsuchiya, T. NorM of Vibrio parahaemolyticus is an Na+-driven multidrug efflux pump. J. Bacteriol. 182, 6694–6697 (2000).
Chen, J. et al. VmrA, a member of a novel class of Na+-coupled multidrug efflux pumps from Vibrio parahaemolyticus. J. Bacteriol. 184, 572–576 (2002).
Long, F., Rouquette-Loughlin, C., Shafer, W.M. & Yu, E.W. Functional cloning and characterization of the multidrug efflux pumps NorM from Neisseria gonorrhoeae and YdhE from Escherichia coli. Antimicrob. Agents Chemother. 52, 3052–3060 (2008).
Hr, G.X. et al. An H+-coupled multidrug efflux pump, PmpM, a member of the MATE family of transporters, from Pseudomonas aeruginosa. J. Bacteriol. 186, 262–265 (2004).
Su, X.Z., Chen, J., Mizushima, T., Kuroda, T. & Tsuchiya, T. AbeM, an H+-coupled Acinetobacter baumannii multidrug efflux pump belonging to the MATE family of transporters. Antimicrob. Agents Chemother. 49, 4362–4364 (2005).
Li, L., He, Z., Pandey, G.K., Tsuchiya, T. & Luan, S. Functional cloning and characterization of a plant efflux carrier for multidrug and heavy metal detoxification. J. Biol. Chem. 277, 5360–5368 (2002).
Otsuka, M. et al. A human transporter protein that mediates the final excretion step for toxic organic cations. Proc. Natl. Acad. Sci. USA 102, 17923–17928 (2005).
Masuda, S. et al. Identification and functional characterization of a new human kidney-specific H+/organic cation antiporter, kidney-specific multidrug and toxin extrusion 2. J. Am. Soc. Nephrol. 17, 2127–2135 (2006).
He, X. et al. Structure of a cation-bound multidrug and toxic compound extrusion transporter. Nature 467, 991–994 (2010).
Lu, M. et al. Structures of a Na+-coupled, substrate-bound MATE multidrug transporter. Proc. Natl. Acad. Sci. USA 110, 2099–2104 (2013).
Mitchell, P. A general theory of membrane transport from studies of bacteria. Nature 180, 134–136 (1957).
Jardetzky, O. Simple allosteric model for membrane pumps. Nature 211, 969–970 (1966).
Huda, M.N. et al. Gene cloning and characterization of VcrM, a Na+-coupled multidrug efflux pump, from Vibrio cholerae non-O1. Microbiol. Immunol. 47, 419–427 (2003).
Dridi, L., Tankoviv, J. & Petit, J.C. CdeA of Clostridium difficile, a new multidrug efflux transporter of the MATE family. Microb. Drug Resist. 10, 191–196 (2004).
Brown, D.G., Swanson, J.K. & Allen, C. The host-induced Ralstonia solanacearum genes, acrA and dinF, encode multidrug efflux pumps and contribute to bacterial wilt virulence. Appl. Environ. Microbiol. 73, 2777–2786 (2007).
Rodríguez-Beltrán, J., Rodríguez-Rojas, A., Guelfo, J.R., Couce, A. & Blázquez, J. The Escherichia coli SOS gene dinF protects against oxidative stress and bile salts. PLoS ONE 7, e34791 (2012).
Otsuka, M. et al. Identification of essential amino acid residues of the NorM Na+/multidrug antiporter in Vibrio parahaemolyticus. J. Bacteriol. 187, 1552–1558 (2005).
Matsumoto, T., Kanamoto, T., Otsuka, M., Omote, H. & Moriyama, Y. Role of glutamate residues in substrate recognition by human MATE1 polyspecific H+/organic cation exporter. Am. J. Physiol. Cell Physiol. 294, C1074–C1078 (2008).
Radchenko, M.V. et al. Potassium/proton antiporter system of Escherichia coli. J. Biol. Chem. 281, 19822–19829 (2006).
Fluman, N., Ryan, C.M., Whitelegge, J.P. & Bibi, E. Dissection of mechanistic principles of a secondary multidrug efflux protein. Mol. Cell 47, 777–787 (2012).
Chen, Y.J. et al. X-ray structure of EmrE supports dual topology model. Proc. Natl. Acad. Sci. USA 104, 18999–19004 (2007).
Bas, D.C., Rogers, D.M. & Jensen, J.H. Very fast prediction and rationalization of pKa values for protein-ligand complexes. Proteins 73, 765–783 (2008).
Adam, Y., Tayer, N., Rotem, D., Schteiber, G. & Schudiner, S. The fast release of sticky protons: kinetics of substrate binding and proton release in a multidrug transporter. Proc. Natl. Acad. Sci. USA 104, 17989–17994 (2007).
Gao, X. et al. Mechanism of substrate recognition and transport by an amino acid antiporter. Nature 463, 828–832 (2010).
Nakashima, R., Sakurai, K., Yamasaki, S., Nishino, K. & Yamaguchi, A. Structures of the multidrug exporter AcrB reveal a proximal multisite drug-binding pocket. Nature 480, 565–569 (2011).
Perez, C., Koshy, C., Yildiz, O. & Ziegler, C. Alternating-access mechanism in conformationally asymmetric trimers of the betaine transporter BetP. Nature 490, 126–130 (2012).
Tanaka, Y. et al. Structural basis for the drug extrusion mechanism by a MATE multidrug transporter. Nature 496, 247–251 (2013).
Yamanaka, H., Kobayashi, H., Takahashi, E. & Okamoto, K. MacAB is involved in the secretion of Escherichia coli heat-stable enterotoxin II. J. Bacteriol. 190, 7693–7698 (2008).
Wiegand, I., Hilpert, K. & Hancock, R.E.W. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 3, 163–175 (2008).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Read, R.J. Pushing the boundaries of molecular replacement with maximum likelihood. Acta Crystallogr. D Biol. Crystallogr. 57, 1373–1382 (2001).
De La Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–494 (1997).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Acknowledgements
We thank the beamline staff at 23-ID and 22-ID of Argonne National Laboratory for assistance during data collection. We also thank D. Fu, R. Kaplan, S. Smith, C. Correll and M. Glucksman for comments on the manuscript. This work was supported by the US National Institutes of Health (R01-GM094195 to M.L.) and Rosalind Franklin University of Medicine and Science (M.L.).
Author information
Authors and Affiliations
Contributions
M.L. conceived of the project. M.L. and Y.G. expressed, purified and crystallized the proteins. M.L. and J.S. collected and analyzed X-ray diffraction data. M.R., R.N. and M.L. conducted mutagenesis and functional studies. M.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Stereo views of the experimental electron density for apo–DinF-BH.
The map was calculated to 3.2 Å resolution using density-modified MIRAS phases. Density modification included solvent flattening, histogram matching, cross-crystal averaging and phase extension. (a) The electron density map (cyan mesh, 1.5 σ) was viewed from the membrane plane and the Cα backbone was colored magenta. Notably, the experimental map shows no gap in protein backbone electron density and thus had enabled chain tracing of the entire protein molecule during model building. (b) The featured slice of electron density map (cyan wire, 1.2 σ) depicts a portion of transmembrane helix 1 (TM1, magenta), revealing the amino-acid side-chain details. The quality of the experimental phases ensured the placement of D40, which is essential for the transport function, into the electron density map with confidence.
Supplementary Figure 2 Structural asymmetry in DinF-BH.
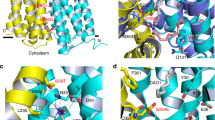
(a) Ribbon rendition of DinF-BH as viewed parallel to the membrane. Left and right views are related by an 180° rotation around the membrane normal. Residues 3-227 and 228-448 were colored cyan and yellow, respectively. Red arrow indicates the wide crevice. (b) Structure as viewed from the periplasmic side. Red arrow highlights the fenestration between TM7 and TM8. (c) View of the protein surface from the periplasm, which was colored according to electrostatic potentials from -20 (red) to +20 kTe-1 (blue). Red arrow indicates the solvent-exposed crevice. (d) Structural overlay of the amino and carboxyl halves as viewed from the membrane plane. Left and right views are related by an 180o rotation around the membrane normal. Residues 3-227 and 228-448 were colored cyan and yellow, respectively, except for residues 253-288, which are in red.
Supplementary Figure 3 Identification of the R6G-binding site.
(a) A slice of electron density (cyan mesh) for the R6G-soaked crystals was overlaid onto the final structural model, with the Cα backbone drawn as black ribbons and R6G as magenta sticks. The electron density map was calculated to 3.7 Å resolution using solvent-flattened MIRAS phases and contoured at 1.5 σ. (b) As a comparison, electron density map (cyan wire) for the apo DinF-BH was calculated to 3.2 Å resolution using density-modified MIRAS phases, contoured at 1.5 σ σ and overlaid onto the final structural model (black). R6G taken from the R6G-bound structure was shown to highlight the absence of the electron density for R6G. (c) Stereo view of the fitting of bound R6G to the electron density. The electron density map (cyan mesh) was calculated to 3.7 Å resolution using native amplitudes and density-modified MIRAS phases combined with model-derived phases. The electron density was overlaid onto the final model of R6G (magenta) and contoured at 1.5 σ.
Supplementary Figure 4 The membrane-embedded, drug-binding aspartates.
(a) Ribbon diagram of DinF-BH as viewed from the membrane plane. Residues 3-227 and 228-448 were colored cyan and yellow, respectively, except for residues 253-288, which were colored red. R6G (magenta) and D40 were drawn as stick models. (b) Structure of R6G-bound NorM-NG (PDB 4HUN)15 as viewed from the membrane plane. Residues 5-230 and 231-459 were colored cyan and yellow, respectively. R6G (magenta) and NorM-NGD41 are shown in stick representation. Blue arrows in (a) and (b) highlight the directions of drug export. (c and d) [H+]-dependence of R6G binding by DinF-BH and DinF-BHD40N. Dissociation constants (Kd) of R6G binding by DinF-BH was plotted against the common logarithm (Log) of H+ concentrations. The dotted line in (c) represents a nonlinear regression fit to an equation describing competitive binding between R6G and H+. The data fitting yielded a Ki of 5.8 ± 0.3 × 10-8 M for H+, which corresponds to a pKa of 7.24 ± 0.02. The error bars represent standard deviations (n=3).
Supplementary Figure 5 Putative intracellular transport pathway in NorM-NG.
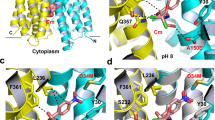
(a) The inward-facing model as viewed from the membrane plane, residues 5-230 and 231-459 were colored cyan and yellow, respectively. (b) Close-up of the intracellular transport path as viewed from the cytoplasmic side. TPP (magenta) and relevant amino acids are shown in stick representation. Functionally critical residues were labeled in red. (c and d) Functional consequences of mutations along the putative transport route. (c) Attenuance (A600nm) measurement of bacteria expressing NorM-NG variants in the presence of 0.5 mg/ml ethidium. (d) Time course of fractional fluorescence reduction (ΔF) as a result of R6G extrusion mediated by NorM-NG variants. Reactions were initiated by the addition of 200 mM NaCl. Error bars indicate standard deviations (n=3). Single mutant of T42, which is located along the known extracellular transport path, had been used as a control.
Supplementary Figure 6 Proposed transport mechanism for NorM-NG.
The protein is shown in ribbon representation, with TPP (magenta) as a stick model and Na+ as a green sphere. The amino (residues 5-230) and carboxyl (231-459) domains of NorM-NG were colored cyan and yellow, respectively. Na+ binding to outward-facing, drug-bound NorM-NG (state I, PDB 4HUK15) triggers drug release, Na+-bound, outward-facing NorM-NG (state II) then switches to the Na+-bound, inward-facing state (III). Drug binding to NorM-NG subsequently promotes the dissociation of Na+ to yield the drug-bound, inward-facing state (IV), which eventually returns to the drug-bound, outward-facing conformation (state I). Magenta and green circles highlight the spatially well-separated substrate- and Na+-binding sites.
Supplementary Figure 7 Structural comparisons of DinF-BH and NorM-NG with pfMATE.
(a and b) Different solvent accessibilities of membrane-embedded aspartates between DinF-BH and pfMATE. Structures of DinF-BH (a) and pfMATE (b, PDB 3VVN)32 as viewed from the membrane plane (left) or the periplasmic side (right). The amino and carboxyl domains were colored cyan and yellow, respectively; except for the extracellular halves of the TM7 and TM8, which were shown in red. D40 in DinF-BH and its counterpart in pfMATE (pfMATED41) were both drawn as sphere models. Compared with D40 in DinF-BH, pfMATED41 is more readily accessible from the solvent, as highlighted by the black arrows. (c) Electrostatic surface potentials of ligand-binding sites in the MATE transporters. Protein surface of NorM-NG (PDB 4HUK)15 was viewed from the periplasmic side of the membrane; and those of DinF-BH and pfMATE (PDB 3VVP)32 were viewed within the lipid bilayer from the carboxyl-terminal domain of the protein. Protein surfaces were colored according to electrostatic potentials from -10 (red) to +10 kTe-1 (blue). The bound ligands were drawn as stick models.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1 and 2 (PDF 893 kb)
Rights and permissions
About this article
Cite this article
Lu, M., Radchenko, M., Symersky, J. et al. Structural insights into H+-coupled multidrug extrusion by a MATE transporter. Nat Struct Mol Biol 20, 1310–1317 (2013). https://doi.org/10.1038/nsmb.2687
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2687
This article is cited by
-
Engineered MATE multidrug transporters reveal two functionally distinct ion-coupling pathways in NorM from Vibrio cholerae
Communications Biology (2021)
-
Structure and mechanism of a redesigned multidrug transporter from the Major Facilitator Superfamily
Scientific Reports (2020)
-
Structure of an engineered multidrug transporter MdfA reveals the molecular basis for substrate recognition
Communications Biology (2019)
-
Genome-wide analysis of the MATE gene family in potato
Molecular Biology Reports (2019)
-
Multidrug efflux pumps: structure, function and regulation
Nature Reviews Microbiology (2018)