Abstract
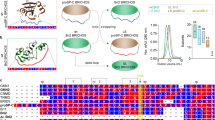
The ring-forming AAA+ protein ClpB cooperates with the DnaK chaperone system to refold aggregated proteins in Escherichia coli. The M domain, a ClpB-specific coiled-coil structure with two wings, motif 1 and motif 2, is essential to disaggregation, but the positioning and mechanistic role of M domains in ClpB hexamers remain unresolved. We show that M domains nestle at the ClpB ring surface, with both M-domain motifs contacting the first ATPase domain (AAA-1). Both wings contribute to maintaining a repressed ClpB activity state. Motif 2 docks intramolecularly to AAA-1 to regulate ClpB unfolding power, and motif 1 contacts a neighboring AAA-1 domain. Mutations that stabilize motif 2 docking repress ClpB, whereas destabilization leads to derepressed ClpB activity with greater unfolding power that is toxic in vivo. Our results underline the vital nature of tight ClpB activity control and elucidate a regulated M-domain toggle control mechanism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Accession codes
References
Tyedmers, J., Mogk, A. & Bukau, B. Cellular strategies for controlling protein aggregation. Nat. Rev. Mol. Cell Biol. 11, 777–788 (2010).
Glover, J.R. & Lindquist, S. Hsp104, Hsp70, and Hsp40: a novel chaperone system that rescues previously aggregated proteins. Cell 94, 73–82 (1998).
Goloubinoff, P., Mogk, A., Peres Ben Zvi, A., Tomoyasu, T. & Bukau, B. Sequential mechanism of solubilization and refolding of stable protein aggregates by a bichaperone network. Proc. Natl. Acad. Sci. USA 96, 13732–13737 (1999).
Zolkiewski, M. ClpB cooperates with DnaK, DnaJ, and GrpE in suppressing protein aggregation. A novel multi-chaperone system from Escherichia coli. J. Biol. Chem. 274, 28083–28086 (1999).
Motohashi, K., Watanabe, Y., Yohda, M. & Yoshida, M. Heat-inactivated proteins are rescued by the DnaK.J-GrpE set and ClpB chaperones. Proc. Natl. Acad. Sci. USA 96, 7184–7189 (1999).
Hong, S.W. & Vierling, E. Mutants of Arabidopsis thaliana defective in the acquisition of tolerance to high temperature stress. Proc. Natl. Acad. Sci. USA 97, 4392–4397 (2000).
Queitsch, C., Hong, S.W., Vierling, E. & Lindquist, S. Heat shock protein 101 plays a crucial role in thermotolerance in Arabidopsis. Plant Cell 12, 479–492 (2000).
Sanchez, Y. & Lindquist, S.L. HSP104 required for induced thermotolerance. Science 248, 1112–1115 (1990).
Squires, C.L., Pedersen, S., Ross, B.M. & Squires, C. ClpB is the Escherichia coli heat shock protein F84.1. J. Bacteriol. 173, 4254–4262 (1991).
Lum, R., Tkach, J.M., Vierling, E. & Glover, J.R. Evidence for an unfolding/threading mechanism for protein disaggregation by Saccharomyces cerevisiae Hsp104. J. Biol. Chem. 279, 29139–29146 (2004).
Weibezahn, J. et al. Thermotolerance requires refolding of aggregated proteins by substrate translocation through the central pore of ClpB. Cell 119, 653–665 (2004).
Lee, S. et al. The structure of ClpB. A molecular chaperone that rescues proteins from an aggregated state. Cell 115, 229–240 (2003).
Mogk, A. et al. Roles of individual domains and conserved motifs of the AAA+ chaperone ClpB in oligomerization, ATP-hydrolysis and chaperone activity. J. Biol. Chem. 278, 17615–17624 (2003).
Kedzierska, S., Akoev, V., Barnett, M.E. & Zolkiewski, M. Structure and function of the middle domain of ClpB from Escherichia coli. Biochemistry 42, 14242–14248 (2003).
Sielaff, B. & Tsai, F.T. The M-domain controls Hsp104 protein remodeling activity in an Hsp70/Hsp40-dependent manner. J. Mol. Biol. 402, 30–37 (2010).
Miot, M. et al. Species-specific collaboration of heat shock proteins (Hsp) 70 and 100 in thermotolerance and protein disaggregation. Proc. Natl. Acad. Sci. USA 108, 6915–6920 (2011).
Haslberger, T. et al. M domains couple the ClpB threading motor with the DnaK chaperone activity. Mol. Cell 25, 247–260 (2007).
Lee, S., Choi, J.M. & Tsai, F.T. Visualizing the ATPase cycle in a protein disaggregating machine: structural basis for substrate binding by ClpB. Mol. Cell 25, 261–271 (2007).
Wendler, P. et al. Atypical AAA+ subunit packing creates an expanded cavity for disaggregation by the protein-remodeling factor Hsp104. Cell 131, 1366–1377 (2007).
Lee, S., Sielaff, B., Lee, J. & Tsai, F.T. CryoEM structure of Hsp104 and its mechanistic implication for protein disaggregation. Proc. Natl. Acad. Sci. USA 107, 8135–8140 (2010).
Wales, T.E. & Engen, J.R. Hydrogen exchange mass spectrometry for the analysis of protein dynamics. Mass Spectrom. Rev. 25, 158–170 (2006).
Schlee, S., Groemping, Y., Herde, P., Seidel, R. & Reinstein, J. The chaperone function of ClpB from Thermus thermophilus depends on allosteric interactions of its two ATP-binding sites. J. Mol. Biol. 306, 889–899 (2001).
Kim, K.I. et al. Heptameric ring structure of the heat-shock protein ClpB, a protein- activated ATPase in Escherichia coli. J. Mol. Biol. 303, 655–666 (2000).
Akoev, V., Gogol, E.P., Barnett, M.E. & Zolkiewski, M. Nucleotide-induced switch in oligomerization of the AAA+ ATPase ClpB. Protein Sci. 13, 567–574 (2004).
del Castillo, U., Fernandez-Higuero, J.A., Perez-Acebron, S., Moro, F. & Muga, A. Nucleotide utilization requirements that render ClpB active as a chaperone. FEBS Lett. 584, 929–934 (2010).
Zietkiewicz, S., Slusarz, M.J., Slusarz, R., Liberek, K. & Rodziewicz-Motowidlo, S. Conformational stability of the full-atom hexameric model of the ClpB chaperone from Escherichia coli. Biopolymers 93, 47–60 (2010).
Haslberger, T. et al. Protein disaggregation by the AAA+ chaperone ClpB involves partial threading of looped polypeptide segments. Nat. Struct. Mol. Biol. 15, 641–650 (2008).
Doyle, S.M. et al. Asymmetric deceleration of ClpB or Hsp104 ATPase activity unleashes protein-remodeling activity. Nat. Struct. Mol. Biol. 14, 114–122 (2007).
Winkler, J., Tyedmers, J., Bukau, B. & Mogk, A. Hsp70 targets Hsp100 chaperones to substrates for protein disaggregation and prion fragmentation. J. Cell Biol. 198, 387–404 (2012).
Maglica, Z., Striebel, F. & Weber-Ban, E. An intrinsic degradation tag on the ClpA C-terminus regulates the balance of ClpAP complexes with different substrate specificity. J. Mol. Biol. 384, 503–511 (2008).
Schlothauer, T., Mogk, A., Dougan, D.A., Bukau, B. & Turgay, K. MecA, an adaptor protein necessary for ClpC chaperone activity. Proc. Natl. Acad. Sci. USA 100, 2306–2311 (2003).
Martin, A., Baker, T.A. & Sauer, R.T. Rebuilt AAA+ motors reveal operating principles for ATP-fuelled machines. Nature 437, 1115–1120 (2005).
Seyffer, F. et al. Hsp70 proteins bind Hsp100 regulatory M domains to activate AAA+ disaggregase at aggregate surfaces. Nat. Struct. Mol. Biol. advance online publication, doi:10.1038/nsmb.2442 (18 November 2012).
del Castillo, U. et al. A quantitative analysis of the effect of nucleotides and the M domain on the association equilibrium of ClpB. Biochemistry 50, 1991–2003 (2011).
Sauer, R.T. & Baker, T.A. AAA+ proteases: ATP-fueled machines of protein destruction. Annu. Rev. Biochem. 80, 587–612 (2011).
Schirmer, E.C., Homann, O.R., Kowal, A.S. & Lindquist, S. Dominant gain-of-function mutations in Hsp104p reveal crucial roles for the middle region. Mol. Biol. Cell 15, 2061–2072 (2004).
Lee, U. et al. Genetic analysis reveals domain interactions of Arabidopsis Hsp100/ClpB and cooperation with the small heat shock protein chaperone system. Plant Cell 17, 559–571 (2005).
Rist, W., Jorgensen, T.J., Roepstorff, P., Bukau, B. & Mayer, M.P. Mapping temperature-induced conformational changes in the Escherichia coli heat shock transcription factor sigma 32 by amide hydrogen exchange. J. Biol. Chem. 278, 51415–51421 (2003).
Zhang, Z. & Marshall, A.G. A universal algorithm for fast and automated charge state deconvolution of electrospray mass-to-charge ratio spectra. J. Am. Soc. Mass Spectrom. 9, 225–233 (1998).
Sali, A. & Blundell, T.L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008).
MacKerell, A.D., Banavali, N. & Foloppe, N. Development and current status of the CHARMM force field for nucleic acids. Biopolymers 56, 257–265 (2001).
Onufriev, A., Bashford, D. & Case, D.A. Exploring protein native states and large-scale conformational changes with a modified generalized born model. Proteins 55, 383–394 (2004).
Monticelli, L. et al. The MARTINI coarse grained forcefield: extension to proteins. J. Chem. Theory Comput. 4, 819–834 (2008).
Periole, X., Cavalli, M., Marrink, S.J. & Ceruso, M. Combining an elastic network with a coarse-grained molecular force field: structure, dynamics and intermolecular recognition. J. Chem. Theory Comput. 5, 2531–2543 (2009).
Lindahl, E., Azuara, C., Koehl, P. & Delarue, M. NOMAD-Ref: visualization, deformation and refinement of macromolecular structures based on all-atom normal mode analysis. Nucleic Acids Res. 34, W52–W56 (2006).
Kulinski, T. et al. Conformational analysis of galanin using end to end distance distribution observed by Forster resonance energy transfer. Eur. Biophys. J. 26, 145–154 (1997).
Acknowledgements
This work was supported by a grant from the Deutsche Forschungsgemeinschaft (BU617-17) to B. Bukau and A. Mogk. We thank T. Ruppert and J. Fiaux for help in HX experiments and L. Guilbride for editing the manuscript. E. Kummer and F. Seyffer were supported by the Hartmut Hoffmann-Berling International Graduate School of Molecular and Cellular Biology. Y. Oguchi was supported by a Humboldt fellowship. M.B. and R.C.W. acknowledge the support of the Klaus Tschira Foundation.
Author information
Authors and Affiliations
Contributions
Y.O., E.K., F.S., M.B., R.C.W., A.M. and B.B. conceived and designed experiments. Y.O., E.K., F.S., M.B., B.A. and R.Z. carried out experiments. Y.O., E.K., F.S., M.B., B.A., R.Z., R.C.W., A.M. and B.B. analyzed the data. R.C.W., A.M. and B.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 4341 kb)
Rights and permissions
About this article
Cite this article
Oguchi, Y., Kummer, E., Seyffer, F. et al. A tightly regulated molecular toggle controls AAA+ disaggregase. Nat Struct Mol Biol 19, 1338–1346 (2012). https://doi.org/10.1038/nsmb.2441
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2441
This article is cited by
-
Cytoplasmic molecular chaperones in Pseudomonas species
Journal of Microbiology (2022)
-
A Review: Molecular Chaperone-mediated Folding, Unfolding and Disaggregation of Expressed Recombinant Proteins
Cell Biochemistry and Biophysics (2021)
-
Processive extrusion of polypeptide loops by a Hsp100 disaggregase
Nature (2020)
-
Genome-wide analysis of the HSP101/CLPB gene family for heat tolerance in hexaploid wheat
Scientific Reports (2020)
-
AtHsp101 research sets course of action for the genetic improvement of crops against heat stress
Journal of Plant Biochemistry and Biotechnology (2020)