Abstract
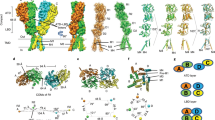
Ionotropic glutamate receptors (iGluRs) are ligand-gated ion channels that mediate most excitatory synaptic transmission in the central nervous system. The free energy of neurotransmitter binding to the ligand-binding domains (LBDs) of iGluRs is converted into useful work to drive receptor activation. We have computed the principal thermodynamic contributions from ligand docking and ligand-induced closure of LBDs for nine ligands of GluA2 using all-atom molecular dynamics free energy simulations. We have validated the results by comparison with experimentally measured apparent affinities to the isolated LBD. Features in the free energy landscapes that govern closure of LBDs are key determinants of binding free energies. An analysis of accessible LBD conformations transposed into the context of an intact GluA2 receptor revealed that the relative displacement of specific diagonal subunits in the tetrameric structure may be key to the action of partial agonists.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Mayer, M.L. Glutamate receptors at atomic resolution. Nature 440, 456–462 (2006).
Sobolevsky, A.I., Rosconi, M.P. & Gouaux, E. X-ray structure, symmetry and mechanism of an AMPA-subtype glutamate receptor. Nature 462, 745–756 (2009).
Armstrong, N. & Gouaux, E. Mechanisms for activation and antagonism of an AMPA-sensitive glutamate receptor: crystal structures of the GluR2 ligand binding core. Neuron 28, 165–181 (2000).
Inanobe, A., Furukawa, H. & Gouaux, E. Mechanism of partial agonist action at the NR1 subunit of NMDA receptors. Neuron 47, 71–84 (2005).
Frydenvang, K. et al. Full domain closure of the ligand-binding core of the ionotropic glutamate receptor iGluR5 induced by the high affinity agonist dysiherbaine and the functional antagonist 8,9-dideoxyneodysiherbaine. J. Biol. Chem. 284, 14219–14229 (2009).
Holm, M.M., Lunn, M.L., Traynelis, S.F., Kastrup, J.S. & Egebjerg, J. Structural determinants of agonist-specific kinetics at the ionotropic glutamate receptor 2. Proc. Natl. Acad. Sci. USA 102, 12053–12058 (2005).
Bjerrum, E.J. & Biggin, P.C. Rigid body essential X-ray crystallography: distinguishing the bend and twist of glutamate receptor ligand binding domains. Proteins 72, 434–446 (2008).
Birdsey-Benson, A., Gill, A., Henderson, L.P. & Madden, D.R. Enhanced efficacy without further cleft closure: reevaluating twist as a source of agonist efficacy in AMPA receptors. J. Neurosci. 30, 1463–1470 (2010).
Ahmed, A.H. et al. Mechanisms of antagonism of the GluR2 AMPA receptor: structure and dynamics of the complex of two willardiine antagonists with the glutamate binding domain. Biochemistry 48, 3894–3903 (2009).
Robert, A., Armstrong, N., Gouaux, J.E. & Howe, J.R. AMPA receptor binding cleft mutations that alter affinity, efficacy, and recovery from desensitization. J. Neurosci. 25, 3752–3762 (2005).
Weston, M.C., Gertler, C., Mayer, M.L. & Rosenmund, C. Interdomain interactions in AMPA and kainate receptors regulate affinity for glutamate. J. Neurosci. 26, 7650–7658 (2006).
Zhang, W., Cho, Y., Lolis, E. & Howe, J.R. Structural and single-channel results indicate that the rates of ligand binding domain closing and opening directly impact AMPA receptor gating. J. Neurosci. 28, 932–943 (2008).
Maltsev, A.S., Ahmed, A.H., Fenwick, M.K., Jane, D.E. & Oswald, R.E. Mechanism of partial agonism at the GluR2 AMPA receptor: measurements of lobe orientation in solution. Biochemistry 47, 10600–10610 (2008).
Woo, H.J. & Roux, B. Calculation of absolute protein-ligand binding free energy from computer simulations. Proc. Natl. Acad. Sci. USA 102, 6825–6830 (2005).
Abele, R., Keinänen, K. & Madden, D.R. Agonist-induced isomerization in a glutamate receptor ligand-binding domain. J. Biol. Chem. 275, 21355–21363 (2000).
Cheng, Q., Du, M., Ramanoudjame, G. & Jayaraman, V. Evolution of glutamate interactions during binding to a glutamate receptor. Nat. Chem. Biol. 1, 329–332 (2005).
Fenwick, M.K. & Oswald, R.E. On the mechanisms of alpha-amino-3-hydroxy-5-methylisoxazole-4-propionic acid (AMPA) receptor binding to glutamate and kainate. J. Biol. Chem. 285, 12334–12343 (2010).
Cheng, Q. & Jayaraman, V. Chemistry and conformation of the ligand-binding domain of GluR2 subtype of glutamate receptors. J. Biol. Chem. 279, 26346–26350 (2004).
Lau, A.Y. & Roux, B. The free energy landscapes governing conformational changes in a glutamate receptor ligand-binding domain. Structure 15, 1203–1214 (2007).
Turski, L. et al. ZK200775: a phosphonate quinoxalinedione AMPA antagonist for neuroprotection in stroke and trauma. Proc. Natl. Acad. Sci. USA 95, 10960–10965 (1998).
Das, U., Kumar, J., Mayer, M.L. & Plested, A.J.R. Domain organization and function in GluK2 subtype kainate receptors. Proc. Natl. Acad. Sci. USA 107, 8463–8468 (2010).
Hogner, A. et al. Structural basis for AMPA receptor activation and ligand selectivity: crystal structures of five agonist complexes with the GluR2 ligand-binding core. J. Mol. Biol. 322, 93–109 (2002).
Nielsen, B.B. et al. Exploring the GluR2 ligand-binding core in complex with the bicyclical AMPA analogue (S)-4-AHCP. FEBS J. 272, 1639–1648 (2005).
Hogner, A. et al. Competitive antagonism of AMPA receptors by ligands of different classes: crystal structure of ATPO bound to the GluR2 ligand-binding core, in comparison with DNQX. J. Med. Chem. 46, 214–221 (2003).
Fiser, A. & Sali, A. ModLoop: automated modeling of loops in protein structures. Bioinformatics 19, 2500–2501 (2003).
Krivov, G.G., Shapovalov, M.V. & Dunbrack, R.L.. Jr Improved prediction of protein side-chain conformations with SCWRL4. Proteins 77, 778–795 (2009).
Wang, J., Wang, W., Kollman, P.A. & Case, D.A. Automatic atom type and bond type perception in molecular mechanical calculations. J. Mol. Graph. Model. 25, 247–260 (2006).
Wang, J., Wolf, R.M., Caldwell, J.W., Kollman, P.A. & Case, D.A. Development and testing of a general AMBER force field. J. Comput. Chem. 25, 1157–1174 (2004).
MacKerell, A.D. Jr. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998).
Frisch, M.J. et al. Gaussian 03, Revision C.02. (2004).
Brooks, B.R. et al. CHARMM: the biomolecular simulation program. J. Comput. Chem. 30, 1545–1614 (2009).
Kumar, S., Rosenberg, J.M., Bouzida, D., Swendsen, R.H. & Kollman, P.A. The weighted histogram analysis method for free-energy calculations on biomolecules. I. The method. J. Comput. Chem. 13, 1011–1021 (1992).
Souaille, M. & Roux, B. Extension to the weighted histogram analysis method: combining umbrella sampling with free energy calculations. Comput. Phys. Commun. 135, 40–57 (2001).
Acknowledgements
We thank E. Gouaux, V. Jayaraman, C. Landes, M. Mayer, R. Oswald and H. Weinstein for review of the manuscript, and W. Gan for discussions. This work was supported by grant MCB-0920261 from the National Science Foundation and grant GM062342 from the US National Institutes of Health. Computational resources were provided by the National Center for Supercomputing Applications (NCSA) through grant MCA01S018.
Author information
Authors and Affiliations
Contributions
A.Y.L. and B.R. designed the research, analyzed the data and wrote the manuscript. A.Y.L. performed the computations.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1–5 and Supplementary Methods (PDF 1074 kb)
Rights and permissions
About this article
Cite this article
Lau, A., Roux, B. The hidden energetics of ligand binding and activation in a glutamate receptor. Nat Struct Mol Biol 18, 283–287 (2011). https://doi.org/10.1038/nsmb.2010
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2010
This article is cited by
-
Automation of absolute protein-ligand binding free energy calculations for docking refinement and compound evaluation
Scientific Reports (2021)
-
Mechanism of partial agonism in AMPA-type glutamate receptors
Nature Communications (2017)
-
Role of the Ion Channel Extracellular Collar in AMPA Receptor Gating
Scientific Reports (2017)
-
Structural mechanisms of activation and desensitization in neurotransmitter-gated ion channels
Nature Structural & Molecular Biology (2016)
-
Error-based Extraction of States and Energy Landscapes from Experimental Single-Molecule Time-Series
Scientific Reports (2015)