Abstract
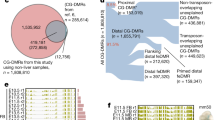
CpG island–like sequences are commonly thought to provide the sole signals for designating constitutively unmethylated regions in the genome, thus generating open chromatin domains within a sea of global repression. Using a new database obtained from comprehensive microarray analysis, we show that unmethylated regions (UMRs) seem to be formed during early embryogenesis, not as a result of CpG-ness, but rather through the recognition of specific sequence motifs closely associated with transcription start sites. This same system probably brings about the resetting of pluripotency genes during somatic cell reprogramming. The data also reveal a new class of nonpromoter UMRs that become de novo methylated in a tissue-specific manner during development, and this process may be involved in gene regulation. In short, we show that UMRs are an important aspect of genome structure that have a dynamic role in development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kafri, T. et al. Developmental pattern of gene-specific DNA methylation in the mouse embryo and germline. Genes Dev. 6, 705–714 (1992).
Monk, M., Boubelik, M. & Lehnert, S. Temporal and regional changes in DNA methylation in the embryonic, extraembryonic and germ cell lineages during mouse embryo development. Development 99, 371–382 (1987).
Brandeis, M. et al. Sp1 elements protect a CpG island from de novo methylation. Nature 371, 435–438 (1994).
Frank, D. et al. Demethylation of CpG islands in embryonic cells. Nature 351, 239–241 (1991).
Macleod, D., Charlton, J., Mullins, J. & Bird, A.P. Sp1 sites in the mouse Aprt gene promoter are required to prevent methylation of the CpG island. Genes Dev. 8, 2282–2292 (1994).
Yamada, Y. et al. A comprehensive analysis of allelic methylation status of CpG islands on human chromosome 11q: comparison with chromosome 21q. DNA Seq. 17, 300–306 (2006).
Yamada, Y. et al. A comprehensive analysis of allelic methylation status of CpG islands on human chromosome 21q. Genome Res. 14, 247–266 (2004).
Shen, L. et al. Genome-wide profiling of DNA methylation reveals a class of normally methylated CpG island promoters. PLoS Genet. 3, 2023–2036 (2007).
Weber, M. et al. Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nat. Genet. 39, 457–466 (2007).
Illingworth, R. et al. A novel CpG island set identifies tissue-specific methylation at developmental gene loci. PLoS Biol. 6, e22 (2008).
Rakyan, V.K. et al. An integrated resource for genome-wide identification and analysis of human tissue-specific differentially methylated regions (tDMRs). Genome Res. 18, 1518–1529 (2008).
Rauch, T.A., Wu, X., Zhong, X., Riggs, A.D. & Pfeifer, G.P. A human B cell methylome at 100-base pair resolution. Proc. Natl. Acad. Sci. USA 106, 671–678 (2009).
Siegfried, Z. et al. DNA methylation represses transcription in vivo. Nat. Genet. 22, 203–206 (1999).
Lorincz, M.C. & Schubeler, D. RNA polymerase II: just stopping by. Cell 130, 16–18 (2007).
Meissner, A. et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454, 766–770 (2008).
Gal-Yam, E.N. et al. Frequent switching of Polycomb repressive marks and DNA hypermethylation in the PC3 prostate cancer cell line. Proc. Natl. Acad. Sci. USA 105, 12979–12984 (2008).
Keshet, I. et al. Evidence for an instructive mechanism of de novo methylation in cancer cells. Nat. Genet. 38, 149–153 (2006).
Weber, M. et al. Chromosome-wide and promoter-specific analyses identify sites of differential DNA methylation in normal and transformed human cells. Nat. Genet. 37, 853–862 (2005).
Guenther, M.G., Levine, S.S., Boyer, L.A., Jaenisch, R. & Young, R.A. A chromatin landmark and transcription initiation at most promoters in human cells. Cell 130, 77–88 (2007).
Zhao, X.D. et al. Whole-genome mapping of histone H3 Lys4 and 27 trimethylations reveals distinct genomic compartments in human embryonic stem cells. Cell Stem Cell 1, 286–298 (2007).
Ooi, S.K. et al. DNMT3L connects unmethylated lysine 4 of histone H3 to de novo methylation of DNA. Nature 448, 714–717 (2007).
Eden, E., Lipson, D., Yogev, S. & Yakhini, Z. Discovering motifs in ranked lists of DNA sequences. PLOS Comput. Biol. 3, e39 (2007).
Bock, C., Walter, J., Paulsen, M. & Lengauer, T. CpG island mapping by epigenome prediction. PLOS Comput. Biol. 3, e110 (2007).
Bock, C. et al. CpG island methylation in human lymphocytes is highly correlated with DNA sequence, repeats, and predicted DNA structure. PLoS Genet. 2, e26 (2006).
Gidekel, S. & Bergman, Y. A unique developmental pattern of Oct-3/4 DNA methylation is controlled by a cis-demodification element. J. Biol. Chem. 277, 34521–34530 (2002).
Eckhardt, F. et al. DNA methylation profiling of human chromosomes 6, 20 and 22. Nat. Genet. 38, 1378–1385 (2006).
Koslowski, M. et al. Frequent nonrandom activation of germ-line genes in human cancer. Cancer Res. 64, 5988–5993 (2004).
Simpson, A.J., Caballero, O.L., Jungbluth, A., Chen, Y.T. & Old, L.J. Cancer/testis antigens, gametogenesis and cancer. Nat. Rev. Cancer 5, 615–625 (2005).
Hajkova, P. et al. Epigenetic reprogramming in mouse primordial germ cells. Mech. Dev. 117, 15–23 (2002).
Oakes, C.C., La Salle, S., Smiraglia, D.J., Robaire, B. & Trasler, J.M. Developmental acquisition of genome-wide DNA methylation occurs prior to meiosis in male germ cells. Dev. Biol. 307, 368–379 (2007).
Hellman, A. & Chess, A. Gene body-specific methylation on the active X chromosome. Science 315, 1141–1143 (2007).
Lock, L.F., Melton, D.W., Caskey, C.T. & Martin, G.R. Methylation of the mouse Hprt gene differs on the active and inactive X chromosomes. Mol. Cell. Biol. 6, 914–924 (1986).
Antequera, F., Boyes, J. & Bird, A. High levels of de novo methylations and altered chromatin structure at CpG islands in cell lines. Cell 62, 503–514 (1990).
Feldman, N. et al. G9a-mediated irreversible epigenetic inactivation of Oct-3/4 during early embryogenesis. Nat. Cell Biol. 8, 188–194 (2006).
Epsztejn-Litman, S. et al. De novo DNA methylation promoted by G9a prevents reprogramming of embryonically silenced genes. Nat. Struct. Mol. Biol. 15, 1176–1183 (2008).
Imamura, M. et al. Transcriptional repression and DNA hypermethylation of a small set of ES cell marker genes in male germline stem cells. BMC Dev. Biol. 6, 34 (2006).
Ma, D.K., Chiang, C.H., Ponnusamy, K., Ming, G.L. & Song, H. G9a and Jhdm2a regulate embryonic stem cell fusion-induced reprogramming of adult neural stem cells. Stem Cells 26, 2131–2141 (2008).
Mikkelsen, T.S. et al. Dissecting direct reprogramming through integrative genomic analysis. Nature 454, 49–55 (2008).
Wutz, A. et al. Imprinted expression of the Igf2r gene depends on an intronic CpG island. Nature 389, 745–749 (1997).
Yu, W. et al. Epigenetic silencing of tumour suppressor gene p15 by its antisense RNA. Nature 451, 202–206 (2008).
Watanabe, T. et al. Endogenous siRNAs from naturally formed dsRNAs regulate transcripts in mouse oocytes. Nature 453, 539–543 (2008).
He, Y., Vogelstein, B., Velculescu, V.E., Papadopoulos, N. & Kinzler, K.W. The antisense transcriptomes of human cells. Science 322, 1855–1857 (2008).
Jones, P.A. The DNA methylation paradox. Trends Genet. 15, 34–37 (1999).
Lehnertz, B. et al. Suv39h-mediated histone H3 lysine 9 methylation directs DNA methylation to major satellite repeats at pericentric heterochromatin. Curr. Biol. 13, 1192–1200 (2003).
Vire, E. et al. The Polycomb group protein EZH2 directly controls DNA methylation. Nature 439, 871–874 (2006).
Schlesinger, Y. et al. Polycomb-mediated methylation on Lys27 of histone H3 methylation pre-marks genes for de novo methylation in cancer. Nat. Genet. 39, 232–236 (2007).
Mohn, F. et al. Lineage-specific Polycomb targets and de novo DNA methylation define restriction and potential of neuronal progenitors. Mol. Cell 30, 755–766 (2008).
Ohm, J.E. et al. A stem cell-like chromatin pattern may predispose tumor suppressor genes to DNA hypermethylation and heritable silencing. Nat. Genet. 39, 237–242 (2007).
Widschwendter, M. et al. Epigenetic stem cell signature in cancer. Nat. Genet. 39, 157–158 (2007).
Reynaud, C. et al. Monitoring of urinary excretion of modified nucleosides in cancer patients using a set of six monoclonal antibodies. Cancer Lett. 61, 255–262 (1992).
Gardiner-Garden, M. & Frommer, M. CpG island in vertebrate genomes. J. Mol. Biol. 196, 261–282 (1987).
Matys, V. et al. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34, D108–D110 (2006).
Witten, I.H. & Frank, E. Data Mining: Practical Machine Learning Tools and Techniques (Morgan Kaufmann, San Francisco, 2005).
Farthing, C.R. et al. Global mapping of DNA methylation in mouse promoters reveals epigenetic reprogramming of pluripotency genes. PLoS Genet. 4, e1000116 (2008).
Cowan, C.A., Atienza, J., Melton, D.A. & Eggan, K. Nuclear reprogramming of somatic cells after fusion with human embryonic stem cells. Science 309, 1369–1373 (2005).
Acknowledgements
This work was supported by the Israel Cancer Research Fund (H.C.), the Rosetrees Trust (H.C.), Lewis Sanders (H.C.), both Philip Morris USA Inc. and Philip Morris International (R.S.) and an Agilent University Relations grant (H.C.).
Author information
Authors and Affiliations
Contributions
R.S. and D.N. carried out all mDIP, bisulfite and transfection experiments and did most of the data analysis; D.R. and R.S. performed all of the labeling and microarray hybridizations; I.St., Z.Y. and I.Si. were responsible for all the computational biology including the development of algorithms; B.B. and N.B. prepared the DNA from human ES cells; R.S. managed and organized all of the experimental work and generated the main concepts; H.C. wrote the manuscript and contributed to the design and interpretation of all experiments.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Methods (PDF 2430 kb)
Supplementary Table 1
CpG island methylation data (XLS 14692 kb)
Supplementary Table 2
CpG island methylation data – BED file for UCSC upload (TXT 3949 kb)
Supplementary Table 3
Sequence motifs enriched in unmethylated islands (PDF 35 kb)
Supplementary Table 4
Prediction of unmethylated CpG islands (PDF 32 kb)
Supplementary Table 5
UMR methylation data (XLS 1710 kb)
Supplementary Table 6
Non-CpG island housekeeping gene promoters (PDF 81 kb)
Supplementary Table 7
Intragenic methylation and GO analysis of Refseq genes with internal islands (PDF 66 kb)
Supplementary Table 8
Comparison of published results estimating the percentage of methylated CpG islands in the Human genome (PDF 51 kb)
Rights and permissions
About this article
Cite this article
Straussman, R., Nejman, D., Roberts, D. et al. Developmental programming of CpG island methylation profiles in the human genome. Nat Struct Mol Biol 16, 564–571 (2009). https://doi.org/10.1038/nsmb.1594
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1594
This article is cited by
-
Selection and validation of a novel set of specific differential methylation markers and construction of a random forest prediction model for the accurate tissue origin identifications of body fluids involving young and middle-aged group of Chinese Han population
International Journal of Legal Medicine (2023)
-
Novel blood-based FUT7 DNA methylation is associated with lung cancer: especially for lung squamous cell carcinoma
Clinical Epigenetics (2022)
-
DNA hypermethylation of NOTCH2NLC in neuronal intranuclear inclusion disease: a case–control study
Journal of Neurology (2022)
-
What impact does oocyte vitrification have on epigenetics and gene expression?
Clinical Epigenetics (2020)
-
Ethanol extract of Ligustrum lucidum Ait. leaves suppressed hepatocellular carcinoma in vitro and in vivo
Cancer Cell International (2019)