Abstract
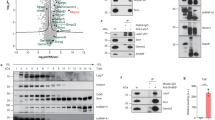
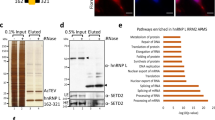
The human p100 protein is a vital transcription regulator that increases gene transcription by forming a physical bridge between promoter-specific activators and the basal transcription machinery. Here we demonstrate that the tudor and SN (TSN) domain of p100 interacts with U small nuclear ribonucleoprotein (snRNP) complexes, suggesting a role for p100 in the processing of precursor messenger RNA. We determined the crystal structure of the p100 TSN domain to delineate the molecular basis of p100's proposed functions. The interdigitated structure resembles a hook, with a hinge controlling the movement and orientation of the hook. Our studies suggest that a conserved aromatic cage hooks methyl groups of snRNPs and anchors p100 to the spliceosome. These structural insights partly explain the distinct roles of p100 in transcription and splicing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Yang, J. et al. Identification of p100 as a coactivator for STAT6 that bridges STAT6 with RNA polymerase II. EMBO J. 21, 4950–4958 (2002).
Valineva, T., Yang, J., Palovuori, R. & Silvennoinen, O. The transcriptional co-activator protein p100 recruits histone acetyltransferase activity to STAT6 and mediates interaction between the CREB-binding protein and STAT6. J. Biol. Chem. 280, 14989–14996 (2005).
Leverson, J.D. et al. Pim-1 kinase and p100 cooperate to enhance c-Myb activity. Mol. Cell 2, 417–425 (1998).
Valineva, T., Yang, J. & Silvennoinen, O. Characterization of RNA helicase A as component of STAT6-dependent enhanceosome. Nucleic Acids Res. 34, 3938–3946 (2006).
Tong, X., Drapkin, R., Yalamanchili, R., Mosialos, G. & Kieff, E. The Epstein-Barr virus nuclear protein 2 acidic domain forms a complex with a novel cellular coactivator that can interact with TFIIE. Mol. Cell. Biol. 15, 4735–4744 (1995).
Paukku, K., Yang, J. & Silvennoinen, O. TSN and nuclease-like domains containing protein p100 function as coactivators for signal transducer and activator of transcription 5. Mol. Endocrinol. 17, 1805–1814 (2003).
Low, S.H. et al. Polycystin-1, STAT6, and P100 function in a pathway that transduces ciliary mechanosensation and is activated in polycystic kidney disease. Dev. Cell 10, 57–69 (2006).
Caudy, A.A. et al. A micrococcal nuclease homologue in RNAi effector complexes. Nature 425, 411–414 (2003).
Callebaut, I. & Mornon, J.P. The human EBNA-2 coactivator p100: multidomain organization and relationship to the staphylococcal nuclease fold and to the TSN protein involved in Drosophila melanogaster development. Biochem. J. 321, 125–132 (1997).
Ponting, C.P. P100, a transcriptional coactivator, is a human homologue of staphylococcal nuclease. Protein Sci. 6, 459–463 (1997).
Selenko, P. et al. SMN tudor domain structure and its interaction with the Sm proteins. Nat. Struct. Biol. 8, 27–31 (2001).
Sprangers, R., Groves, M.R., Sinning, I. & Sattler, M. High-resolution X-ray and NMR structures of the SMN TSN domain: conformational variation in the binding site for symmetrically dimethylated arginine residues. J. Mol. Biol. 327, 507–520 (2003).
Murzin, A.G. OB (oligonucleotide/oligosaccharide binding)-fold: common structural and functional solution for non-homologous sequences. EMBO J. 12, 861–867 (1993).
Hynes, T.R. & Fox, R.O. The crystal structure of Staphylococcal nuclease refined at 1.7 Å resolution. Proteins Struct. Funct. Genet. 10, 92–105 (1991).
Combet, C., Jambon, M., Deleage, G. & Geourjon, C. Geno3D: automatic comparative molecular modeling of protein. Bioinformatics 18, 213–214 (2002).
Eissenberg, J.C. & Elgin, C.R. Antagonizing the neighbours. Nature 438, 1090–1091 (2005).
Brahms, H., Meheus, L., Brabandere, V., Fischer, U. & Luhrmann, R. Symmetrical dimethylation of arginine residues in spliceosomal Sm protein B/B0 and the Sm-like protein LSm4, and their interaction with the SMN protein. RNA 7, 1531–1542 (2001).
Friesen, W.J., Massenet, S., Paushkin, S., Wyce, A. & Dreyfuss, G. SMN, the product of the spinal muscular atrophy gene, binds preferentially to dimethylarginine-containing protein targets. Mol. Cell 7, 1111–1117 (2001).
Nielsen, P.R. et al. Structure of the HP1 chromodomain bound to histone H3 methylated at lysine 9. Nature 416, 103–107 (2002).
Jacobs, S.A. & Khorasanizadeh, S. Structure of the HP1 chromodomain bound to a lysine 9-methylated histone H3 tail. Science 295, 2080–2083 (2002).
Huang, Y., Fang, J., Bedford, M.T., Zhang, Y. & Xu, R-M. Recognition of histone H3 lysine-4 methylation by the double TSN domain of JMJD2A. Science 312, 748–751 (2006).
Huyen, Y. et al. Methylated lysine 79 of histone H3 targets 53BP1 to DNA double-strand breaks. Nature 432, 406–411 (2004).
Min, J., Zhang, Y. & Xu, R.M. Structural basis for specific binding of Polycomb chromodomain to histone H3 methylated at Lys 27. Genes Dev. 17, 1823–1828 (2003).
Botuyan, M.V. et al. Structural basis for the methylation state-specific recognition of histone H4–K20 by 53BP1 and Crb2 in DNA repair. Cell 127, 1361–1373 (2006).
Kiss, T. Biogenesis of small nuclear RNPs. J. Cell Sci. 117, 5949–5951 (2004).
Reed, R. Coupling transcription, splicing and mRNA export. Curr. Opin. Cell Biol. 15, 326–331 (2003).
Rayment, I. Reductive alkylation of lysine residues to alter crystallization properties of proteins. Methods Enzymol. 276, 171–179 (1997).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
de La Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–494 (1997).
Perrakis, A., Morris, R. & Lamzin, V.S. Automated protein model building combined with iterative structure refinement. Nat. Struct. Biol. 6, 458–463 (1999).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr D Biol. Crystallogr. 53, 240–255 (1997).
McRee, D.E. XtalView/Xfit — a versatile program for manipulating atomic coordinates and electron density. J. Struct. Biol. 125, 156–165 (1999).
Davis, I.W., Murray, L.W., Richardson, J.S. & Richardson, D.C. MOLPROBITY: structure validation and all-atom contact analysis for nucleic acids and their complexes. Nucleic Acids Res. 32, W615–W619 (2004).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Cryst. 26, 283–291 (1993).
Berman, H.M. et al. The Protein Data Bank and the challenge of structural genomics. Nat. Struct. Biol. 7, 957–959 (2000).
Frilander, M.J. & Steitz, J.A. Initial recognition of U12-dependent introns requires both U11/5′ splice-site and U12/branchpoint interactions. Genes Dev. 13, 851–863 (1999).
Baker, N.A., Sept, D., Joseph, S., Holst, M.J. & McCammon, J.A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 98, 10037–10041 (2001).
Acknowledgements
This work was funded by the 863 (grant 2006AA02A316) and 973 (grant 2006CB910901) projects of the Ministry of Science and Technology of China, the National Natural Science Foundation of China (grants 30670427, 30670441 and 30300070), the US National Institutes of Health (grant 1P50 GM62407), the University of Georgia Research Foundation, the Georgia Research Alliance, Program for New Century Excellent Talents in University (grant NCET-04-0245), Tianjin Municipal Science and Technology Commission (grant 07JCZDJC07300) and the Institute of Biophysics, Chinese Academy of Sciences. Supporting institutions for the SER-CAT 22-ID beamline at the Advanced Photon Source may be found at http://www.ser-cat.org/members.html. Use of the Advanced Photon Source was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences, under contract number W-31-109-Eng-38.
Author information
Authors and Affiliations
Contributions
N.S., M.Z., C.C. and H.X. contributed to the structural studies. J.Y., J.S., Y.D. and O.S. contributed to the mutagenesis and functional characterization of the p100 TSN domain. Z.J.-L., Y.L. and Z.Y. contributed to data collection and analysis. Z.-J.L., J.Y., Z. R. and B.-C.W. conceived the study and participated in its design and coordination. N.S., Z.-J.L., O.S. and J.Y. drafted the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2 (PDF 2030 kb)
Rights and permissions
About this article
Cite this article
Shaw, N., Zhao, M., Cheng, C. et al. The multifunctional human p100 protein 'hooks' methylated ligands. Nat Struct Mol Biol 14, 779–784 (2007). https://doi.org/10.1038/nsmb1269
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1269
This article is cited by
-
SND1 acts as a novel gene transcription activator recognizing the conserved Motif domains of Smad promoters, inducing TGFβ1 response and breast cancer metastasis
Oncogene (2017)
-
Tudor staphylococcal nuclease: biochemistry and functions
Cell Death & Differentiation (2016)
-
Structure and domain organization of Drosophila Tudor
Cell Research (2014)
-
Organization of Plasmodium falciparum spliceosomal core complex and role of arginine methylation in its assembly
Malaria Journal (2013)
-
Molecular cloning of BmTUDOR-SN and analysis of its role in the RNAi pathway in the silkworm, Bombyx mori (Lepidoptera: Bombycidae)
Applied Entomology and Zoology (2012)