Abstract
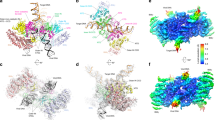
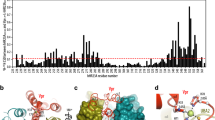
Simian virus 40 (SV40) provides a model system for the study of eukaryotic DNA replication, in which the viral protein, large T antigen (Tag), marshals human proteins to replicate the viral minichromosome. SV40 replication requires interaction of Tag with the host single-stranded DNA-binding protein, replication protein A (hRPA). The C-terminal domain of the hRPA32 subunit (RPA32C) facilitates initiation of replication, but whether it interacts with Tag is not known. Affinity chromatography and NMR revealed physical interaction between hRPA32C and the Tag origin DNA–binding domain, and a structural model of the complex was determined. Point mutations were then designed to reverse charges in the binding sites, resulting in substantially reduced binding affinity. Corresponding mutations introduced into intact hRPA impaired initiation of replication and primosome activity, implying that this interaction has a critical role in assembly and progression of the SV40 replisome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Fanning, E. & Knippers, R. Structure and function of simian virus 40 large tumor antigen. Annu. Rev. Biochem. 61, 55–85 (1992).
Bullock, P.A. The initiation of simian virus 40 DNA replication in vitro. Crit. Rev. Biochem. Mol. Biol. 32, 503–568 (1997).
Simmons, D.T. SV40 large T antigen functions in DNA replication and transformation. Adv. Virus Res. 55, 75–134 (2000).
Stenlund, A. Initiation of DNA replication: lessons from viral initiator proteins. Nat. Rev. Mol. Cell Biol. 4, 777–785 (2003).
Stauffer, M.E. & Chazin, W.J. Structural mechanisms of DNA replication, repair, and recombination. J. Biol. Chem. 279, 30915–30918 (2004).
Wold, M.S. Replication protein A: a heterotrimeric, single-stranded DNA-binding protein required for eukaryotic DNA metabolism. Annu. Rev. Biochem. 66, 61–92 (1997).
Iftode, C., Daniely, Y. & Borowiec, J.A. Replication protein A (RPA): the eukaryotic SSB. Crit. Rev. Biochem. Mol. Biol. 34, 141–180 (1999).
Bochkarev, A. & Bochkareva, E. From RPA to BRCA2: lessons from single-stranded DNA binding by the OB-fold. Curr. Opin. Struct. Biol. 14, 36–42 (2004).
Mer, G., Bochkarev, A., Chazin, W.J. & Edwards, A.M. Three-dimensional structure and function of replication protein A. Cold Spring Harb. Symp. Quant. Biol. 65, 193–200 (2000).
Weisshart, K. et al. Partial proteolysis of SV40 T antigen reveals intramolecular contacts between domains and conformation changes upon hexamer assembly. J. Biol. Chem. 279, 38943–38951 (2004).
Gai, D. et al. Insights into the oligomeric states, conformational changes and helicase activities of SV40 large tumor antigen. J. Biol. Chem. 279, 38952–38959 (2004).
Dornreiter, I. et al. Interaction of DNA polymerase α-primase with cellular replication protein A and SV40 T antigen. EMBO J. 11, 769–776 (1992).
Melendy, T. & Stillman, B. An interaction between replication protein A and SV40 T antigen appears essential for primosome assembly during SV40 DNA replication. J. Biol. Chem. 268, 3389–3395 (1993).
Collins, K.L. & Kelly, T.J. Effects of T antigen and replication protein A on the initiation of DNA synthesis by DNA polymerase α-primase. Mol. Cell. Biol. 11, 2108–2115 (1991).
Weisshart, K., Taneja, P. & Fanning, E. The replication protein A binding site in simian virus 40 (SV40) T antigen and its role in the initial steps of SV40 DNA replication. J. Virol. 72, 9771–9781 (1998).
Braun, K.A., Lao, Y., He, Z., Ingles, C.J. & Wold, M.S. Role of protein-protein interactions in the function of replication protein A (RPA): RPA modulates the activity of DNA polymerase α by multiple mechanisms. Biochemistry 36, 8443–8454 (1997).
Mer, G. et al. Structural basis for the recognition of DNA repair proteins UNG2, XPA, and RAD52 by replication factor A. Cell 103, 449–456 (2000).
Lee, S.H. & Kim, D.K. The role of the 34-kDa subunit of human replication protein A in simian virus 40 DNA replication in vitro. J. Biol. Chem. 270, 12801–12807 (1995).
Kenny, M.K., Schlegel, U., Furneaux, H. & Hurwitz, J. The role of human single-stranded DNA binding protein and its individual subunits in simian virus 40 DNA replication. J. Biol. Chem. 265, 7693–7700 (1990)
Matsumoto, T., Eki, T. & Hurwitz, J. Studies on the initiation and elongation reactions in the simian virus 40 DNA replication system. Proc. Natl. Acad. Sci. USA 87, 9712–9716 (1990).
Arunkumar, A.I., Stauffer, M.E., Bochkareva, E., Bochkarev, A. & Chazin, W.J. Independent and coordinated functions of replication protein A tandem high affinity single-stranded DNA binding domains. J. Biol. Chem. 278, 41077–41082 (2003).
Luo, X., Sanford, D.G., Bullock, P.A. & Bachovchin, W.W. Solution structure of the origin DNA–binding domain of SV40 T-antigen. Nat. Struct. Biol. 3, 1034–1039 (1996).
Wun-Kim, K. et al. The DNA-binding domain of simian virus 40 tumor antigen has multiple functions. J. Virol. 67, 7608–7611 (1993).
Simmons, D.T., Loeber, G. & Tegtmeyer, P. Four major sequence elements of simian virus 40 large T antigen coordinate its specific and nonspecific DNA binding. J. Virol. 64, 1973–1983 (1990).
Bradshaw, E.M. et al. T antigen origin-binding domain of simian virus 40: determinants of specific DNA binding. Biochemistry 43, 6928–6936 (2004).
Titolo, S., Welchner, E., White, P.W. & Archambault, J. Characterization of the DNA-binding properties of the origin-binding domain of simian virus 40 large T antigen by fluorescence anisotropy. J. Virol. 77, 5512–5518 (2003).
Yuzhakov, A., Kelman, Z., Hurwitz, J. & O'Donnell, M. Multiple competition reactions for RPA order the assembly of the DNA polymerase δ holoenzyme. EMBO J. 18, 6189–6199 (1999).
Santocanale, C., Neecke, H., Longhese, M.P., Lucchini, G. & Plevani, P. Mutations in the gene encoding the 34 kDa subunit of yeast replication protein A cause defective S phase progression. J. Mol. Biol. 254, 595–607 (1995).
Daughdrill, G.W. et al. Chemical shift changes provide evidence for overlapping single-stranded DNA- and XPA-binding sites on the 70 kDa subunit of human replication protein A. Nucleic Acids Res. 31, 4176–4183 (2003).
Jackson, D., Dhar, K., Wahl, J.K., Wold, M.S. & Borgstahl, G.E. Analysis of the human replication protein A:Rad52 complex: evidence for crosstalk between RPA32, RPA70, Rad52 and DNA. J. Mol. Biol. 321, 133–148 (2002).
Kowalczykowski, S.C. Some assembly required. Nat. Struct. Biol. 7, 1087–1089 (2000).
Blackwell, L.J. & Borowiec, J.A. Human replication protein A binds single-stranded DNA in two distinct complexes. Mol. Cell. Biol. 14, 3993–4001 (1994).
Blackwell, L.J., Borowiec, J.A. & Mastrangelo, I.A. Single-stranded-DNA binding alters human replication protein A structure and facilitates interaction with DNA-dependent protein kinase. Mol. Cell. Biol. 16, 4798–4807 (1996).
Iftode, C. & Borowiec, J.A. 5′→3′ molecular polarity of human replication protein A (RPA) binding to pseudo-origin DNA substrates. Biochemistry 39, 11970–11981 (2000).
Bastin-Shanower, S.A. & Brill, S.J. Functional analysis of the four DNA binding domains of replication protein A. The role of RPA2 in ssDNA binding. J. Biol. Chem. 276, 36446–36453 (2001).
de Laat, W.L. et al. DNA-binding polarity of human replication protein A positions nucleases in nucleotide excision repair. Genes Dev. 12, 2598–2609 (1998).
Ott, R.D., Wang, Y. & Fanning, E. Mutational analysis of simian virus 40 T-antigen primosome activities in viral DNA replication. J. Virol. 76, 5121–5130 (2002).
Henricksen, L.A., Umbricht, C.B. & Wold, M.S. Recombinant replication protein A: expression, complex formation, and functional characterization. J. Biol. Chem. 269, 11121–11132 (1994).
Pervushin, K.V., Riek, R., Wider, G. & Wuthrich, K. Attenuated T2 relaxation by mutual cancellation of dipole-dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc. Natl. Acad. Sci. USA 94, 12366–12371 (1997).
Sass, H.J., Musco, G., Stahl, S.J., Wingfield, P.T. & Grzesiek, S. Solution NMR of proteins within polyacrylamide gels: diffusional properties and residual alignment by mechanical stress or embedding of oriented purple membranes. J. Biomol. NMR 18, 303–309 (2000).
Tycko, R., Blanco, F.J. & Ishii, Y. Alignment of biopolymers in strained gels: a new way to create detectable dipole-dipole couplings in high-resolution biomolecular NMR. J. Am. Chem. Soc. 122, 9340–9349 (2000).
Bax, A. Weak alignment offers new NMR opportunities to study protein structure and dynamics. Protein Sci. 12, 1–16 (2003).
Zweckstetter, M. & Bax, A. Prediction of sterically induced alignment in a dilute liquid crystalline phase: aid to protein structure determination by NMR. J. Am. Chem. Soc. 122, 3791–3792 (2000)
Dominguez, C., Boelens, R. & Bonvin, A.M. HADDOCK: a protein-protein docking approach based on biochemical or biophysical information. J. Am. Chem. Soc. 125, 1731–1737 (2003).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Koradi, R., Billeter, M. & Wuthrich, K. MOLMOL: a program for display and analysis of macromolecular structures. J. Mol. Graph. 14, 51–55, 29–32 (1996).
Laskowski, R.A., Rullman, J.A., MacArthur, M.W., Kaptein, R. & Thornton, J.M. AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 8, 477–486 (1996).
Huang, S.G., Weisshart, K., Gilbert, I. & Fanning, E. Stoichiometry and mechanism of assembly of SV40 T antigen complexes with the viral origin of DNA replication and DNA polymerase α-primase. Biochemistry 37, 15345–15352 (1998).
Simmons, D.T., Gai, D., Parsons, R., Debes, A. & Roy, R. Assembly of the replication initiation complex on SV40 origin DNA. Nucleic Acids Res. 32, 1103–1112 (2004).
Waga, S. & Stillman, B. The DNA replication fork in eukaryotic cells. Annu. Rev. Biochem. 67, 721–751 (1998).
Acknowledgements
We thank S. Bhattacharya, B. Dattilo, L. Douthitt, G. Hubbell, J. Jacob, M. Karra, M. Kenny, V. Klymovych, S. Meyn, C.S. Newlon, C. Sanders, L. Schwertman, E.M. Warren, D.R. Williams and M.S. Wold for valuable advice and assistance. Accelrys provided a gift of NMR software. Financial support is gratefully acknowledged from the US National Institutes of Health for operating grants and support to the Vanderbilt-Ingram Cancer Center and the Vanderbilt Center in Molecular Toxicology, as well as from the Howard Hughes Medical Institute Professors Program (to E.F.) and Vanderbilt University.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Mapping the Tag-OBD-binding site of RPA32C. (PDF 553 kb)
Supplementary Fig. 2
Mapping the RPA32C-binding site of Tag-OBD. (PDF 610 kb)
Supplementary Fig. 3
NMR analysis of side chains at the intermolecular surface. (PDF 280 kb)
Supplementary Fig. 4
NMR chemical shift analysis of the interaction of Tag-OBD with RPA32C. (PDF 287 kb)
Supplementary Fig. 5
Sequence and surface comparison of yRP32C and hRPA32C. (PDF 795 kb)
Supplementary Fig. 6
Characterization of hRPAy32C and hRPAΔ223. (PDF 81 kb)
Supplementary Fig. 7
Quantitative comparison of wild-type and mutant RPA in replication assays. (PDF 65 kb)
Rights and permissions
About this article
Cite this article
Arunkumar, A., Klimovich, V., Jiang, X. et al. Insights into hRPA32 C-terminal domain–mediated assembly of the simian virus 40 replisome. Nat Struct Mol Biol 12, 332–339 (2005). https://doi.org/10.1038/nsmb916
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb916
This article is cited by
-
RPA-coated single-stranded DNA as a platform for post-translational modifications in the DNA damage response
Cell Research (2015)
-
Structural analysis reveals DNA binding properties of Rv2827c, a hypothetical protein from Mycobacterium tuberculosis
Journal of Structural and Functional Genomics (2009)
-
Structural mechanism of RPA loading on DNA during activation of a simple pre-replication complex
The EMBO Journal (2006)