Abstract
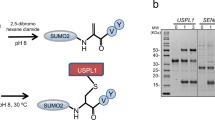
Post-translational modification with small ubiquitin-related modifier (SUMO) alters the function of many proteins, but the molecular mechanisms and consequences of this modification are still poorly defined. During a screen for novel SUMO1 targets, we identified the ubiquitin-conjugating enzyme E2-25K (Hip2). SUMO attachment severely impairs E2-25K ubiquitin thioester and unanchored ubiquitin chain formation in vitro. Crystal structures of E2-25K(1–155) and of the E2-25K(1–155)–SUMO conjugate (E2-25K*SUMO) indicate that SUMO attachment interferes with E1 interaction through its location on the N-terminal helix. The SUMO acceptor site in E2-25K, Lys14, does not conform to the consensus site found in most SUMO targets (ΨKXE), and functions only in the context of an α-helix. In contrast, adjacent SUMO consensus sites are modified only when in unstructured peptides. The demonstration that secondary structure elements are part of SUMO attachment signals could contribute to a better prediction of SUMO targets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Pickart, C.M. Mechanisms underlying ubiquitination. Annu. Rev. Biochem. 70, 503–533 (2001).
Passmore, L.A. & Barford, D. Getting into position: the catalytic mechanisms of protein ubiquitylation. Biochem. J. 379, 513–525 (2004).
Pichler, A. & Melchior, F. SUMO E3 ligases. In SUMOylation. Molecular Biology and Biochemistry (ed. Van Wilson, G.) (Horizon Press, Norwich, UK, 2004).
Johnson, E.S. Protein modification by SUMO. Annu. Rev. Biochem. 73, 355–382 (2004).
Macauley, M.S. et al. Structural and dynamic independence of isopeptide-linked RanGAP1 and SUMO-1. J. Biol. Chem. 279, 49131–49137 (2004).
Chen, Z. & Pickart, C.M. A 25-kilodalton ubiquitin carrier protein (E2) catalyzes multi-ubiquitin chain synthesis via lysine 48 of ubiquitin. J. Biol. Chem. 265, 21835–21842 (1990).
Song, S. et al. Essential role of E2-25K/Hip-2 in mediating amyloid-β neurotoxicity. Mol. Cell 12, 553–563 (2003).
Kalchman, M.A. et al. Huntingtin is ubiquitinated and interacts with a specific ubiquitin-conjugating enzyme. J. Biol. Chem. 271, 19385–19394 (1996).
Lelouard, H. et al. Dendritic cell aggresome-like induced structures are dedicated areas for ubiquitination and storage of newly synthesized defective proteins. J. Cell Biol. 164, 667–675 (2004).
Girdwood, D. et al. P300 transcriptional repression is mediated by SUMO modification. Mol. Cell 11, 1043–1054 (2003).
Hardeland, U., Steinacher, R., Jiricny, J. & Schar, P. Modification of the human thymine-DNA glycosylase by ubiquitin-like proteins facilitates enzymatic turnover. EMBO J. 21, 1456–1464 (2002).
Hoege, C., Pfander, B., Moldovan, G.L., Pyrowolakis, G. & Jentsch, S. RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature 419, 135–141 (2002).
Huang, T.T., Wuerzberger-Davis, S.M., Wu, Z.H. & Miyamoto, S. Sequential modification of NEMO/IKKγ by SUMO-1 and ubiquitin mediates NF-κB activation by genotoxic stress. Cell 115, 565–576 (2003).
Haldeman, M.T., Xia, G., Kasperek, E.M. & Pickart, C.M. Structure and function of ubiquitin conjugating enzyme E2-25K: the tail is a core-dependent activity element. Biochemistry 36, 10526–10537 (1997).
Hamilton, K.S. et al. Structure of a conjugating enzyme-ubiquitin thiolester intermediate reveals a novel role for the ubiquitin tail. Structure 9, 897–904 (2001).
Cook, W.J., Jeffrey, L.C., Kasperek, E. & Pickart, C.M. Structure of tetraubiquitin shows how multiubiquitin chains can be formed. J. Mol. Biol. 236, 601–609 (1994).
Phillips, C.L., Thrower, J., Pickart, C.M. & Hill, C.P. Structure of a new crystal form of tetraubiquitin. Acta Crystallogr. D 57, 341–344 (2001).
Hu, M. et al. Crystal structure of a UBP-family deubiquitinating enzyme in isolation and in complex with ubiquitin aldehyde. Cell 111, 1041–1054 (2002).
Mossessova, E. & Lima, C.D. Ulp1-SUMO crystal structure and genetic analysis reveal conserved interactions and a regulatory element essential for cell growth in yeast. Mol. Cell 5, 865–876 (2000).
Walden, H. et al. The structure of the APPBP1–UBA3–NEDD8–ATP complex reveals the basis for selective ubiquitin-like protein activation by an E1. Mol. Cell 12, 1427–1437 (2003).
Sullivan, M.L. & Vierstra, R.D. Cloning of a 16-kDa ubiquitin carrier protein from wheat and Arabidopsis thaliana. Identification of functional domains by in vitro mutagenesis. J. Biol. Chem. 266, 23878–23885 (1991).
Bencsath, K.P., Podgorski, M.S., Pagala, V.R., Slaughter, C.A. & Schulman, B.A. Identification of a multifunctional binding site on Ubc9p required for Smt3p conjugation. J. Biol. Chem. 277, 47938–47945 (2002).
Huang, D.T. et al. A unique E1-E2 interaction required for optimal conjugation of the ubiquitin-like protein NEDD8. Nat. Struct. Mol. Biol. 11, 927–935 (2004).
Sampson, D.A., Wang, M. & Matunis, M.J. The small ubiquitin-like modifier-1 (SUMO-1) consensus sequence mediates Ubc9 binding and is essential for SUMO-1 modification. J. Biol. Chem. 276, 21664–21669 (2001).
Bernier-Villamor, V., Sampson, D.A., Matunis, M.J. & Lima, C.D. Structural basis for E2-mediated SUMO conjugation revealed by a complex between ubiquitin-conjugating enzyme Ubc9 and RanGAP1. Cell 108, 345–356 (2002).
Lin, D. et al. Identification of a substrate recognition site on Ubc9. J. Biol. Chem. 277, 21740–21748 (2002).
Huang, L. et al. Structure of an E6AP–UbcH7 complex: insights into ubiquitination by the E2-E3 enzyme cascade. Science 286, 1321–1326 (1999).
Cope, G.A. & Deshaies, R.J. COP9 signalosome: a multifunctional regulator of SCF and other cullin-based ubiquitin ligases. Cell 114, 663–671 (2003).
Wolf, D.A., Zhou, C. & Wee, S. The COP9 signalosome: an assembly and maintenance platform for cullin ubiquitin ligases? Nat. Cell Biol. 5, 1029–1033 (2003).
Mahajan, R., Delphin, C., Guan, T., Gerace, L. & Melchior, F. A small ubiquitin-related polypeptide involved in targeting RanGAP1 to nuclear pore complex protein RanBP2. Cell 88, 97–107 (1997).
Pichler, A., Gast, A., Seeler, J.S., Dejean, A. & Melchior, F. The nucleoporin RanBP2 has SUMO1 E3 ligase activity. Cell 108, 109–120 (2002).
Haldeman, M.T., Finley, D. & Pickart, C.M. Dynamics of ubiquitin conjugation during erythroid differentiation in vitro. J. Biol. Chem. 270, 9507–9516 (1995).
Shevchenko, A., Wilm, M., Vorm, O. & Mann, M. Mass spectrometric sequencing of proteins silver-stained polyacrylamide gels. Anal. Chem. 68, 850–858 (1996).
Gobom, J., Nordhoff, E., Mirgorodskaya, E., Ekman, R. & Roepstorff, P. Sample purification and preparation technique based on nano-scale reversed-phase columns for the sensitive analysis of complex peptide mixtures by matrix-assisted laser desorption/ionization mass spectrometry. J. Mass Spectrom. 34, 105–116 (1999).
Collaborative Computational Project 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Perrakis, A., Morris, R. & Lamzin, V.S. Automated protein model building combined with iterative structure refinement. Nat. Struct. Biol. 6, 458–463 (1999).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Pickart, C.M. & Vella, A.T. Levels of active ubiquitin carrier proteins decline during erythroid maturation. J. Biol. Chem. 263, 12028–12035 (1988).
Acknowledgements
Our special thanks go to S. Swaminathan, C. Pickart, L. Hengst, A. Perrakis and members of the labs for stimulating discussions, T. Buesgen for her advice on GST-SUMO1 pull-downs, P. Celie for the CD experiments, H. Langedijk of Pepscan for the peptide synthesis, and A. Klanner, J. Vordemann and E. Stieger for excellent technical assistance. C. Pickart, N. Dantuma, A. Geerlof and R. Hay are gratefully acknowledged for providing reagents.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Conventional SUMO sites in E2-25K structure. (PDF 1777 kb)
Rights and permissions
About this article
Cite this article
Pichler, A., Knipscheer, P., Oberhofer, E. et al. SUMO modification of the ubiquitin-conjugating enzyme E2-25K. Nat Struct Mol Biol 12, 264–269 (2005). https://doi.org/10.1038/nsmb903
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb903
This article is cited by
-
Proteomic strategies for characterizing ubiquitin-like modifications
Nature Reviews Methods Primers (2021)
-
SET SUMOylation promotes its cytoplasmic retention and induces tau pathology and cognitive impairments
Acta Neuropathologica Communications (2019)
-
Hypoxia-induced Slug SUMOylation enhances lung cancer metastasis
Journal of Experimental & Clinical Cancer Research (2019)
-
Multilevel structure–activity profiling reveals multiple green tea compound families that each modulate ubiquitin-activating enzyme and ubiquitination by a distinct mechanism
Scientific Reports (2019)
-
Hip2 ubiquitin-conjugating enzyme has a role in UV-induced G1/S arrest and re-entry
Genes & Genomics (2019)