Abstract
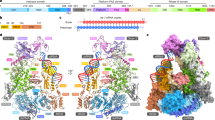
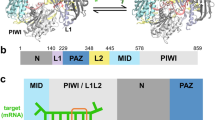
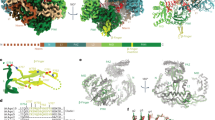
RISC, the RNA-induced silencing complex, uses short interfering RNAs (siRNAs) or micro RNAs (miRNAs) to select its targets in a sequence-dependent manner. Key RISC components are Argonaute proteins, which contain two characteristic domains, PAZ and PIWI. PAZ is highly conserved and is found only in Argonaute proteins and Dicer. We have solved the crystal structure of the PAZ domain of Drosophila Argonaute2. The PAZ domain contains a variant of the OB fold, a module that often binds single-stranded nucleic acids. PAZ domains show low-affinity nucleic acid binding, probably interacting with the 3′ ends of single-stranded regions of RNA. PAZ can bind the characteristic two-base 3′ overhangs of siRNAs, indicating that although PAZ may not be a primary nucleic acid binding site in Dicer or RISC, it may contribute to the specific and productive incorporation of siRNAs and miRNAs into the RNAi pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hannon, G.J. RNA interference. Nature 418, 244–251 (2002).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Carrington, J.C. & Ambros, V. Role of microRNAs in plant and animal development. Science 301, 336–338 (2003).
Volpe, T. et al. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297, 1833–1837 (2002).
Hall, I.M. et al. Establishment and maintenance of a heterochromatin domain. Science 297, 2232–2237 (2002).
Elbashir, S.M., Martinez, J., Patkaniowska, A., Lendeckel, W. & Tuschl, T. Functional anatomy of siRNAs for mediating efficient RNAi in Drosophila melanogaster embryo lysate. EMBO J. 20, 6877–6888 (2001).
Schwarz, D.S., Hutvagner, G., Haley, B. & Zamore, P.D. Evidence that siRNAs function as guides, not primers, in the Drosophila and human RNAi pathways. Mol. Cell 10, 537–548 (2002).
Bernstein, E., Caudy, A.A., Hammond, S.M. & Hannon, G.J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, 363–366 (2001).
Hammond, S.M., Boettcher, S., Caudy, A.A., Kobayashi, R. & Hannon, G.J. Argonaute2, a link between genetic and biochemical analyses of RNAi. Science 293, 1146–1150 (2001).
Martinez, J., Patkaniowska, A., Urlaub, H., Luhrmann, R. & Tuschl, T. Single-stranded antisense siRNAs guide target RNA cleavage in RNAi. Cell 110, 563–574 (2002).
Cerutti, L., Mian, N. & Bateman, A. Domains in gene silencing and cell differentiation proteins: the novel PAZ domain and redefinition of the Piwi domain. Trends Biochem. Sci. 25, 481–482 (2000).
Carmell, M.A., Xuan, Z., Zhang, M.Q. & Hannon, G.J. The Argonaute family: tentacles that reach into RNAi, developmental control, stem cell maintenance, and tumorigenesis. Genes Dev. 16, 2733–2742 (2002).
Murzin, A.G. OB(oligonucleotide/oligosaccharide binding)-fold: common structural and functional solution for non-homologous sequences. EMBO J. 12, 861–867 (1993).
Theobald, D.L., Mitton-Fry, R.M. & Wuttke, D.S. Nucleic acid recognition by OB-fold proteins. Ann. Rev. Biophys. Biomol. Struct. 32, 115–133 (2003).
Murzin, A.G., Brenner, S.E., Hubbard, T. & Chothia, C. SCOP: a structural classification of proteins database for the investigation of sequences and structures. J. Mol. Biol. 247, 536–540 (1995).
Anderson, E.M., Halsey, W.A. & Wuttke, D.S. Site-directed mutagenesis reveals the thermodynamic requirements for single-stranded DNA recognition by the telomere-binding protein Cdc13. Biochemistry 42, 3751–3758 (2003).
Hockensmith, J.W., Kubasek, W.L., Vorachek, W.R., Evertsz, E.M. & von Hippel, P.H. Laser cross-linking of protein–nucleic acid complexes. Methods Enzymol. 208, 211–236 (1991).
Anston, A. Single-stranded RNA binding proteins. Curr. Opin. Struct. Biol. 10, 87–94 (2000).
Theobald, D.L., Cervantes, R.B., Lundblad, V. & Wuttke, D.S. Homology among telomeric end-protection proteins. Structure 11, 1049–1050 (2003).
Nykanen, A., Haley, B. & Zamore, P.D. ATP requirements and small interfering RNA structure in the RNA interference pathway. Cell 107, 309–321 (2001).
Zhang, H., Kolb, F.A., Brondani, V., Billy, E. & Filipowicz, W. Human Dicer preferentially cleaves dsRNAs at their termini without a requirement for ATP. EMBO J. 21, 5875–5885 (2002).
Lee, Y. et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 425, 415–419 (2003).
Navaza, J. & Saludjian, P. AMoRe: an automated molecular replacement program package. Methods Enzymol. 276, 581–594 (1997).
Jones, T.A. & Kjeldgaard, M. Electron-density map interpretation. Methods Enzymol. 277, 173–208 (1997).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Terwilliger, T.C. Maximum likelihood density modification. Acta Crystallogr. D 56, 965–972 (2000).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Esnouf, R.M. An extensively modified version of MolScript that includes greatly enhanced coloring capabilities. J. Mol. Graph. 15, 132–134 (1997).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Cryst. 24, 946–950 (1991).
Bacon, D.J. & Anderson, W.F. A fast algorithm for rendering space-filling molecule pictures. J. Molec. Graph. 6, 219–220 (1988).
Merritt, E.A. & Murphy, M.E.P. Raster3D version 2.0—a program for photorealistic molecular graphics. Acta Crystallogr. D 50, 869–873 (1994).
Rees, B., Webster, G., Delarue, M., Boeglin, M.A. & Moras, D. Aspartyl tRNA-synthetase from Escherichia coli: flexibility and adaptability to the substrates. J. Mol. Biol. 299, 1157–1164 (2000).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association—insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Acknowledgements
We thank E. Enemark and M. Carmell for help with refinement and figures, A. Caudy for aid in characterizing the biochemical properties of GST-PAZ, M. Myers for mass spectrometry and A. Heroux (beamline X26C) for support with data collection at the National Synchrotron Light Source (NSLS) at Brookhaven National Laboratory. The NSLS is supported by the US Department of Energy, Division of Materials Sciences and Division of Chemical Sciences. J.J.S. is a Bristol-Myers Squibb Predoctoral Fellow, N.H.T. is a Leslie C. Quick Jr. Predoctoral Fellow. This work was supported by the Watson School of Biological Sciences (L.J.) and the US National Institutes of Health (G.J.H.). G.J.H. is a Rita Allen Foundation Scholar and is supported by an Innovator Award from the US Army Breast Cancer Research Fund.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Song, JJ., Liu, J., Tolia, N. et al. The crystal structure of the Argonaute2 PAZ domain reveals an RNA binding motif in RNAi effector complexes. Nat Struct Mol Biol 10, 1026–1032 (2003). https://doi.org/10.1038/nsb1016
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb1016