Abstract
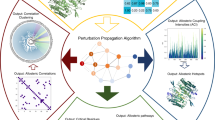
A fundamental goal in cellular signaling is to understand allosteric communication, the process by which signals originating at one site in a protein propagate reliably to affect distant functional sites. The general principles of protein structure that underlie this process remain unknown. Here, we describe a sequence-based statistical method for quantitatively mapping the global network of amino acid interactions in a protein. Application of this method for three structurally and functionally distinct protein families (G protein–coupled receptors, the chymotrypsin class of serine proteases and hemoglobins) reveals a surprisingly simple architecture for amino acid interactions in each protein family: a small subset of residues forms physically connected networks that link distant functional sites in the tertiary structure. Although small in number, residues comprising the network show excellent correlation with the large body of mechanistic data available for each family. The data suggest that evolutionarily conserved sparse networks of amino acid interactions represent structural motifs for allosteric communication in proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Gether, U. Uncovering molecular mechanisms involved in activation of G protein-coupled receptors. Endocr. Rev. 21, 90–113 (2000).
Menon, S.T., Han, M. & Sakmar, T.P. Rhodopsin: structural basis of molecular physiology. Physiol. Rev. 81, 1659–1688 (2001).
Hedstrom, L., Szilagyi, L. & Rutter, W.J. Converting trypsin to chymotrypsin: the role of surface loops. Science 255, 1249–1253 (1992).
Hedstrom, L. Trypsin: a case study in the structural determinants of enzyme specificity. Biol. Chem. 377, 465–470 (1996).
Patten, P.A. et al. The immunological evolution of catalysis. Science 271, 1086–1091 (1996).
Perutz, M.F., Wilkinson, A.J., Paoli, M. & Dodson, G.G. The stereochemical mechanism of the cooperative effects in hemoglobin revisited. Annu. Rev. Biophys. Biomol. Struct. 27, 1–34 (1998).
Perutz, M.F., Fermi, G., Luisi, B., Shaanan, B. & Liddington, R.C. Stereochemistry of cooperative mechanisms in hemoglobin. Cold Spring Harb. Symp. Quant. Biol. 52, 555–565 (1987).
Perutz, M.F. Stereochemistry of cooperative effects in haemoglobin. Nature 228, 726–739 (1970).
Paoli, M., Liddington, R., Tame, J., Wilkinson, A. & Dodson, G. Crystal structure of T state haemoglobin with oxygen bound at all four haems. J. Mol. Biol. 256, 775–792 (1996).
Perona, J.J., Hedstrom, L., Rutter, W.J. & Fletterick, R.J. Structural origins of substrate discrimination in trypsin and chymotrypsin. Biochemistry 34, 1489–1499 (1995).
Williams, D.C. Jr., Benjamin, D.C., Poljak, R.J. & Rule, G.S. Global changes in amide hydrogen exchange rates for a protein antigen in complex with three different antibodies. J. Mol. Biol. 257, 866–876 (1996).
Schreiber, G. & Fersht, A.R. Energetics of protein-protein interactions: analysis of the barnase-barstar interface by single mutations and double mutant cycles. J. Mol. Biol. 248, 478–486 (1995).
Hidalgo, P. & MacKinnon, R. Revealing the architecture of a K+ channel pore through mutant cycles with a peptide inhibitor. Science 268, 307–310 (1995).
Carter, P.J., Winter, G., Wilkinson, A.J. & Fersht, A.R. The use of double mutants to detect structural changes in the active site of the tyrosyl-tRNA synthetase (Bacillus stearothermophilus). Cell 38, 835–840 (1984).
Lockless, S.W. & Ranganathan, R. Evolutionarily conserved pathways of energetic connectivity in protein families. Science 286, 295–299 (1999).
Lichtarge, O., Bourne, H.R. & Cohen, F.E. An evolutionary trace method defines binding surfaces common to protein families. J. Mol. Biol. 257, 342–358 (1996).
Marcotte, E.M. et al. Detecting protein function and protein-protein interactions from genome sequences. Science 285, 751–753 (1999).
Pellegrini, M., Marcotte, E.M., Thompson, M.J., Eisenberg, D. & Yeates, T.O. Assigning protein functions by comparative genome analysis: protein phylogenetic profiles. Proc. Natl. Acad. Sci. USA 96, 4285–4288 (1999).
Ballesteros, J.A., Shi, L. & Javitch, J.A. Structural mimicry in G protein-coupled receptors: implications of the high-resolution structure of rhodopsin for structure-function analysis of rhodopsin-like receptors. Mol. Pharmacol. 60, 1–19 (2001).
Saraceni-Richards, C.A. & Levy, S.B. Second-site suppressor mutations of inactivating substitutions at Gly247 of the tetracycline efflux protein, Tet(B). J. Bacteriol. 182, 6514–6516 (2000).
Minor, D.L. Jr., Masseling, S.J., Jan, Y.N. & Jan, L.Y. Transmembrane structure of an inwardly rectifying potassium channel. Cell 96, 879–891 (1999).
Cain, S.M., Matzke, E.A. & Brooker, R.J. The conserved motif in hydrophilic loop 2/3 and loop 8/9 of the lactose permease of Escherichia coli. Analysis of suppressor mutations. J. Membr. Biol. 176, 159–168 (2000).
Zhang, H., Skinner, M.M., Sandberg, W.S., Wang, A.H. & Terwilliger, T.C. Context dependence of mutational effects in a protein: the crystal structures of the V35I, I47V and V35I/I47V gene V protein core mutants. J. Mol. Biol. 259, 148–159 (1996).
Baldwin, E., Xu, J., Hajiseyedjavadi, O., Baase, W.A. & Matthews, B.W. Thermodynamic and structural compensation in 'size-switch' core repacking variants of bacteriophage T4 lysozyme. J. Mol. Biol. 259, 542–559 (1996).
Neher, E. How frequent are correlated changes in families of protein sequences? Proc. Natl. Acad. Sci. USA 91, 98–102 (1994).
Nakayama, T.A. & Khorana, H.G. Orientation of retinal in bovine rhodopsin determined by cross-linking using a photoactivatable analog of 11-cis-retinal. J. Biol. Chem. 265, 15762–15769 (1990).
Palczewski, K. et al. Crystal structure of rhodopsin: a G protein-coupled receptor. Science 289, 739–745 (2000).
Robinson, P.R., Cohen, G.B., Zhukovsky, E.A. & Oprian, D.D. Constitutively active mutants of rhodopsin. Neuron 9, 719–725 (1992).
Porter, J.E., Hwa, J. & Perez, D.M. Activation of the α1b-adrenergic receptor is initiated by disruption of an interhelical salt bridge constraint. J. Biol. Chem. 271, 28318–28323 (1996).
Yano, K., Kohn, L.D., Saji, M., Okuno, A. & Cutler, G.B. Jr. Phe576 plays an important role in the secondary structure and intracellular signaling of the human luteinizing hormone/chorionic gonadotropin receptor. J. Clin. Endocrinol. Metab. 82, 2586–2591 (1997).
Andres, A., Kosoy, A., Garriga, P. & Manyosa, J. Mutations at position 125 in transmembrane helix III of rhodopsin affect the structure and signalling of the receptor. Eur. J. Biochem. 268, 5696–5704 (2001).
Garriga, P., Liu, X. & Khorana, H.G. Structure and function in rhodopsin: correct folding and misfolding in point mutants at and in proximity to the site of the retinitis pigmentosa mutation Leu-125→Arg in the transmembrane helix C. Proc. Natl. Acad. Sci. USA 93, 4560–4564 (1996).
Okada, T., Ernst, O.P., Palczewski, K. & Hofmann, K.P. Activation of rhodopsin: new insights from structural and biochemical studies. Trends Biochem. Sci. 26, 318–324 (2001).
Han, M., Smith, S.O. & Sakmar, T.P. Constitutive activation of opsin by mutation of methionine 257 on transmembrane helix 6. Biochemistry 37, 8253–8261 (1998).
Gripentrog, J.M., Jesaitis, A.J. & Miettinen, H.M. A single amino acid substitution (N297A) in the conserved NPXXY sequence of the human N-formyl peptide receptor results in inhibition of desensitization and endocytosis, and a dose-dependent shift in p42/44 mitogen-activated protein kinase activation and chemotaxis. Biochem. J. 352, 399–407 (2000).
Meng, E.C. & Bourne, H.R. Receptor activation: what does the rhodopsin structure tell us? Trends Pharmacol. Sci. 22, 587–593 (2001).
Farrens, D.L., Altenbach, C., Yang, K., Hubbell, W.L. & Khorana, H.G. Requirement of rigid-body motion of transmembrane helices for light activation of rhodopsin. Science 274, 768–770 (1996).
Altenbach, C., Klein-Seetharaman, J., Cai, K., Khorana, H.G. & Hubbell, W.L. Structure and function in rhodopsin: mapping light-dependent changes in distance between residue 316 in helix 8 and residues in the sequence 60–75, covering the cytoplasmic end of helices TM1 and TM2 and their connection loop CL1. Biochemistry 40, 15493–15500 (2001).
Altenbach, C., Cai, K., Klein-Seetharaman, J., Khorana, H.G. & Hubbell, W.L. Structure and function in rhodopsin: mapping light-dependent changes in distance between residue 65 in helix TM1 and residues in the sequence 306–319 at the cytoplasmic end of helix TM7 and in helix H8. Biochemistry 40, 15483–15492 (2001).
Dunham, T.D. & Farrens, D.L. Conformational changes in rhodopsin. Movement of helix f detected by site-specific chemical labeling and fluorescence spectroscopy. J. Biol. Chem. 274, 1683–1690 (1999).
Cai, K. et al. Single-cysteine substitution mutants at amino acid positions 306-321 in rhodopsin, the sequence between the cytoplasmic end of helix VII and the palmitoylation sites: sulfhydryl reactivity and transducin activation reveal a tertiary structure. Biochemistry 38, 7925–7930 (1999).
Altenbach, C., Cai, K., Khorana, H.G. & Hubbell, W.L. Structural features and light-dependent changes in the sequence 306–322 extending from helix VII to the palmitoylation sites in rhodopsin: a site-directed spin-labeling study. Biochemistry 38, 7931–7937 (1999).
Fahmy, K. & Sakmar, T.P. Regulation of the rhodopsin-transducin interaction by a highly conserved carboxylic acid group. Biochemistry 32, 7229–7236 (1993).
Franke, R.R., Konig, B., Sakmar, T.P., Khorana, H.G. & Hofmann, K.P. Rhodopsin mutants that bind but fail to activate transducin. Science 250, 123–125 (1990).
Franke, R.R., Sakmar, T.P., Graham, R.M. & Khorana, H.G. Structure and function in rhodopsin. Studies of the interaction between the rhodopsin cytoplasmic domain and transducin. J. Biol. Chem. 267, 14767–14774 (1992).
Kim, J.M., Altenbach, C., Thurmond, R.L., Khorana, H.G. & Hubbell, W.L. Structure and function in rhodopsin: rhodopsin mutants with a neutral amino acid at E134 have a partially activated conformation in the dark state. Proc. Natl. Acad. Sci. USA 94, 14273–14278 (1997).
Getz, G., Levine, E. & Domany, E. Coupled two-way clustering analysis of gene microarray data. Proc. Natl. Acad. Sci. USA 97, 12079–12084 (2000).
Luque, I., Leavitt, S.A. & Freire, E. The linkage between protein folding and functional cooperativity: two sides of the same coin? Annu. Rev. Biophys. Biomol. Struct. 31, 235–256 (2002).
Scheidig, A.J., Hynes, T.R., Pelletier, L.A., Wells, J.A. & Kossiakoff, A.A. Crystal structures of bovine chymotrypsin and trypsin complexed to the inhibitor domain of Alzheimer's amyloid β-protein precursos (APPI) and basic pancreatic trypsin inhibitor (BPTI): engineering of inhibitors with altered specificities. Protein Sci. 6, 1806 (1997).
Graf, L. et al. Electrostatic complementarity within the substrate-binding pocket of trypsin. Proc. Natl. Acad. Sc.i USA 85, 4961–4965 (1988).
Hedstrom, L., Perona, J.J. & Rutter, W.J. Converting trypsin to chymotrypsin: residue 172 is a substrate specificity determinant. Biochemistry 33, 8757–8763 (1994).
Szabo, E., Bocskei, Z., Naray-Szabo, G. & Graf, L. The three-dimensional structure of Asp189Ser trypsin provides evidence for an inherent structural plasticity of the protease. Eur. J. Biochem. 263, 20–26 (1999).
Mace, J.E. & Agard, D.A. Kinetic and structural characterization of mutations of glycine 216 in α-lytic protease: a new target for engineering substrate specificity. J. Mol. Biol. 254, 720–736 (1995).
Davis, J.H. & Agard, D.A. Relationship between enzyme specificity and the backbone dynamics of free and inhibited α-lytic protease. Biochemistry 37, 7696–7707 (1998).
Liddington, R., Derewenda, Z., Dodson, E., Hubbard, R. & Dodson, G. High resolution crystal structures and comparisons of T-state deoxyhaemoglobin and two liganded T-state haemoglobins: T(α-oxy)haemoglobin and T(met)haemoglobin. J. Mol. Biol. 228, 551–579 (1992).
Stevens, S.Y., Sanker, S., Kent, C. & Zuiderweg, E.R. Delineation of the allosteric mechanism of a cytidylyltransferase exhibiting negative cooperativity. Nat. Struct. Biol. 8, 947–952 (2001).
Nicholson, L.K. et al. Flexibility and function in HIV-1 protease. Nat. Struct. Biol. 2, 274–280 (1995).
Osborne, M.J., Schnell, J., Benkovic, S.J., Dyson, H.J. & Wright, P.E. Backbone dynamics in dihydrofolate reductase complexes: role of loop flexibility in the catalytic mechanism. Biochemistry 40, 9846–9859 (2001).
Eisenmesser, E.Z., Bosco, D.A., Akke, M. & Kern, D. Enzyme dynamics during catalysis. Science 295, 1520–1523 (2002).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Thompson, J.D., Higgins, D.G. & Gibson, T.J. CLUSTALW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Beukers, M.W., Kristiansen, I., IJzerman, A.P. & Edvardsen, I. TinyGRAP database: a bioinformatics tool to mine G-protein-coupled receptor mutant data. Trends Pharmacol. Sci. 20, 475–477 (1999).
Horn, F. et al. GPCRDB: an information system for G protein-coupled receptors. Nucleic Acids Res. 26, 275–279 (1998).
Doolittle, R., Abelson, J.N. & Simon, M.I. Computer Methods for Macromolecular Sequence Analysis (Academic Press, San Diego; 1996).
Nicholls, A., Sharp, K. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed andschematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Bacon, D. & Anderson, W.F. A fast algorithm for rendering space-filling molecule pictures. J. Mol. Graph. 6, 219–220 (1988).
Acknowledgements
We thank J. Albanesi, M. Brown, A. Gilman, and members of the Ranganathan lab for critical reading of the manuscript. This work was partially supported by a grant from the Robert A. Welch Foundation to R.R., who is also a recipient of the Burroughs-Wellcome Fund New Investigator Award in the Basic Pharmacological Sciences and the Mallinckrodt Scholar Award. M.A.W. is a Research Associate and R.R. is an Associate Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Süel, G., Lockless, S., Wall, M. et al. Evolutionarily conserved networks of residues mediate allosteric communication in proteins. Nat Struct Mol Biol 10, 59–69 (2003). https://doi.org/10.1038/nsb881
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb881
This article is cited by
-
Vanilloid-dependent TRPV1 opening trajectory from cryoEM ensemble analysis
Nature Communications (2022)
-
The two redox states of the human NEET proteins’ [2Fe–2S] clusters
JBIC Journal of Biological Inorganic Chemistry (2021)
-
Glutantβase: a database for improving the rational design of glucose-tolerant β-glucosidases
BMC Molecular and Cell Biology (2020)
-
Computational design of G Protein-Coupled Receptor allosteric signal transductions
Nature Chemical Biology (2020)
-
Mapping allosteric communications within individual proteins
Nature Communications (2020)