Key Points
-
Autophagy is positively regulated by AMPK, ULK1 and beclin-1 complexes, and negatively regulated by AKT activation, mTORC1 and mTORC2 complexes
-
Autophagy drives cell-fate decisions (including T-cell selection, cell differentiation and apoptosis) and is important in immune cell function and signalling
-
Autophagy is upregulated in rheumatoid arthritis fibroblast-like synoviocytes and systemic lupus erythematosus T cells and plasma cells, increasing their survival and activity
-
Autophagy is downregulated in osteoarthritis chondrocytes (increasing their apoptosis) and systemic sclerosis fibroblasts (increasing their myofibroblastic differentiation)
-
Many therapeutic agents either directly or indirectly target autophagy; however, these must be chosen carefully owing to the multicellular pathophysiology of rheumatic diseases
Abstract
Autophagy, an endogenous process necessary for the turnover of organelles, maintains cellular homeostasis and directs cell fate. Alterations to the regulation of autophagy contribute to the progression of various rheumatic diseases, including systemic lupus erythematosus (SLE), rheumatoid arthritis (RA), osteoarthritis (OA) and systemic sclerosis (SSc). Implicit in the progression of these diseases are cell-type-specific responses to surrounding factors that alter autophagy: chondrocytes within articular cartilage show decreased autophagy in OA, leading to rapid cell death and cartilage degeneration; fibroblasts from patients with SSc have restricted autophagy, similar to that seen in aged dermal fibroblasts; fibroblast-like synoviocytes from RA joints show altered autophagy, which contributes to synovial hyperplasia; and dysregulation of autophagy in haematopoietic lineage cells alters their function and maturation in SLE. Various upstream mechanisms also contribute to these diseases by regulating autophagy as part of their signalling cascades. In this Review, we discuss the links between autophagy, immune responses, fibrosis and cellular fates as they relate to pathologies associated with rheumatic diseases. Therapies in clinical use, and in preclinical or clinical development, are also discussed in relation to their effects on autophagy in rheumatic diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
09 February 2017
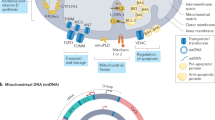
In the article, the sentence “The ubiquitin-like modifier-activating enzyme ATG7 ubiquitylates autophagy protein 5 (ATG5) so that it can form a functional complex with ubiquitin-like protein ATG12” should have appeared as “The ubiquitin-like modifier-activating enzyme ATG7 conjugates autophagy protein 5 (ATG5) with ubiquitin-like protein ATG12 so that it can form a functional complex”. In Figure 1 of the same article, ATG7 was incorrectly shown to be ubiquitylating ATG5, instead of conjugating ATG5 to ATG12. These errors have been corrected in the online version of the article.
References
Mizushima, N. Autophagy: process and function. Genes Dev. 21, 2861–2873 (2007).
Cuervo, A. M. & Wong, E. Chaperone-mediated autophagy: roles in disease and aging. Cell Res. 24, 92–104 (2014).
Li, W. W., Li, J. & Bao, J. K. Microautophagy: lesser-known self-eating. Cell. Mol. Life Sci. 69, 1125–1136 (2012).
Baek, K. H., Park, J. & Shin, I. Autophagy-regulating small molecules and their therapeutic applications. Chem. Soc. Rev. 41, 3245–3263 (2012).
Fleming, A., Noda, T., Yoshimori, T. & Rubinsztein, D. C. Chemical modulators of autophagy as biological probes and potential therapeutics. Nat. Chem. Biol. 7, 9–17 (2011).
Gros, F. & Muller, S. Pharmacological regulators of autophagy and their link with modulators of lupus disease. Br. J. Pharmacol. 171, 4337–4359 (2014).
Renna, M., Jimenez-Sanchez, M., Sarkar, S. & Rubinsztein, D. C. Chemical inducers of autophagy that enhance the clearance of mutant proteins in neurodegenerative diseases. J. Biol. Chem. 285, 11061–11067 (2010).
Rubinsztein, D. C., Codogno, P. & Levine, B. Autophagy modulation as a potential therapeutic target for diverse diseases. Nat. Rev. Drug Discov. 11, 709–730 (2012).
Hosokawa, N. et al. Atg101, a novel mammalian autophagy protein interacting with Atg13. Autophagy 5, 973–979 (2009).
Jung, C. H. et al. ULK–Atg13–FIP200 complexes mediate mTOR signaling to the autophagy machinery. Mol. Biol. Cell 20, 1992–2003 (2009).
Zhao, Z. et al. Coronavirus replication does not require the autophagy gene ATG5. Autophagy 3, 581–585 (2007).
Mizushima, N. et al. A protein conjugation system essential for autophagy. Nature 395, 395–398 (1998).
Fujita, N. et al. The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol. Biol. Cell 19, 2092–2100 (2008).
Itakura, E., Kishi, C., Inoue, K. & Mizushima, N. Beclin 1 forms two distinct phosphatidylinositol 3-kinase complexes with mammalian Atg14 and UVRAG. Mol. Biol. Cell 19, 5360–5372 (2008).
Vanhaesebroeck, B., Guillermet-Guibert, J., Graupera, M. & Bilanges, B. The emerging mechanisms of isoform-specific PI3K signalling. Nat. Rev. Mol. Cell Biol. 11, 329–341 (2010).
Zhong, Y. et al. Distinct regulation of autophagic activity by Atg14L and Rubicon associated with Beclin 1–phosphatidylinositol-3-kinase complex. Nat. Cell Biol. 11, 468–476 (2009).
Nazio, F. et al. mTOR inhibits autophagy by controlling ULK1 ubiquitylation, self-association and function through AMBRA1 and TRAF6. Nat. Cell Biol. 15, 406–416 (2013).
Hoyer-Hansen, M. et al. Control of macroautophagy by calcium, calmodulin-dependent kinase kinase-β, and Bcl-2. Mol. Cell 25, 193–205 (2007).
Liang, X. H. et al. Protection against fatal Sindbis virus encephalitis by Beclin, a novel Bcl-2-interacting protein. J. Virol. 72, 8586–8596 (1998).
Pattingre, S. et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 122, 927–939 (2005).
Chen, D. et al. Parkin mono-ubiquitinates Bcl-2 and regulates autophagy. J. Biol. Chem. 285, 38214–38223 (2010).
Shi, C. S. & Kehrl, J. H. TRAF6 and A20 regulate lysine 63-linked ubiquitination of Beclin-1 to control TLR4-induced autophagy. Sci. Signal. 3, ra42 (2010).
Wei, Y., Pattingre, S., Sinha, S., Bassik, M. & Levine, B. JNK1-mediated phosphorylation of Bcl-2 regulates starvation-induced autophagy. Mol. Cell 30, 678–688 (2008).
Zalckvar, E. et al. DAP-kinase-mediated phosphorylation on the BH3 domain of beclin 1 promotes dissociation of beclin 1 from Bcl-XL and induction of autophagy. EMBO Rep. 10, 285–292 (2009).
Hardie, D. G. Sensing of energy and nutrients by AMP-activated protein kinase. Am. J. Clin. Nutr. 93, 891S–896S (2011).
Kim, J., Kundu, M., Viollet, B. & Guan, K. L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell Biol. 13, 132–141 (2011).
Gwinn, D. M. et al. AMPK phosphorylation of raptor mediates a metabolic checkpoint. Mol. Cell 30, 214–226 (2008).
Loffler, A. S. et al. Ulk1-mediated phosphorylation of AMPK constitutes a negative regulatory feedback loop. Autophagy 7, 696–706 (2011).
Hara, K. et al. Raptor, a binding partner of target of rapamycin (TOR), mediates TOR action. Cell 110, 177–189 (2002).
Nojima, H. et al. The mammalian target of rapamycin (mTOR) partner, raptor, binds the mTOR substrates p70 S6 kinase and 4E-BP1 through their TOR signaling (TOS) motif. J. Biol. Chem. 278, 15461–15464 (2003).
Peterson, T. R. et al. DEPTOR is an mTOR inhibitor frequently overexpressed in multiple myeloma cells and required for their survival. Cell 137, 873–886 (2009).
Wang, L., Harris, T. E., Roth, R. A. & Lawrence, J. C. Jr. PRAS40 regulates mTORC1 kinase activity by functioning as a direct inhibitor of substrate binding. J. Biol. Chem. 282, 20036–20044 (2007).
Oshiro, N. et al. The proline-rich Akt substrate of 40 kDa (PRAS40) is a physiological substrate of mammalian target of rapamycin complex 1. J. Biol. Chem. 282, 20329–20339 (2007).
Duan, S. et al. mTOR generates an auto-amplification loop by triggering the βTrCP- and CK1α-dependent degradation of DEPTOR. Mol. Cell 44, 317–324 (2011).
Yang, H. et al. mTOR kinase structure, mechanism and regulation. Nature 497, 217–223 (2013).
Kaizuka, T. et al. Tti1 and Tel2 are critical factors in mammalian target of rapamycin complex assembly. J. Biol. Chem. 285, 20109–20116 (2010).
Dibble, C. C. & Cantley, L. C. Regulation of mTORC1 by PI3K signaling. Trends Cell Biol. 25, 545–555 (2015).
Zhang, Y. et al. Rheb is a direct target of the tuberous sclerosis tumour suppressor proteins. Nat. Cell Biol. 5, 578–581 (2003).
Long, X., Lin, Y., Ortiz-Vega, S., Yonezawa, K. & Avruch, J. Rheb binds and regulates the mTOR kinase. Curr. Biol. 15, 702–713 (2005).
Sancak, Y. et al. Ragulator–Rag complex targets mTORC1 to the lysosomal surface and is necessary for its activation by amino acids. Cell 141, 290–303 (2010).
Pearce, L. R. et al. Identification of Protor as a novel Rictor-binding component of mTOR complex-2. Biochem. J. 405, 513–522 (2007).
Pearce, L. R., Sommer, E. M., Sakamoto, K., Wullschleger, S. & Alessi, D. R. Protor-1 is required for efficient mTORC2-mediated activation of SGK1 in the kidney. Biochem. J. 436, 169–179 (2011).
Frias, M. A. et al. mSin1 is necessary for Akt/PKB phosphorylation, and its isoforms define three distinct mTORC2s. Curr. Biol. 16, 1865–1870 (2006).
Arnold, J. et al. Autophagy is dispensable for B-cell development but essential for humoral autoimmune responses. Cell Death Differ. 23, 853–864 (2016).
Matsuzawa, Y. et al. TNFAIP3 promotes survival of CD4 T cells by restricting MTOR and promoting autophagy. Autophagy 11, 1052–1062 (2015).
Pengo, N. et al. Plasma cells require autophagy for sustainable immunoglobulin production. Nat. Immunol. 14, 298–305 (2013).
Pua, H. H., Dzhagalov, I., Chuck, M., Mizushima, N. & He, Y. W. A critical role for the autophagy gene Atg5 in T cell survival and proliferation. J. Exp. Med. 204, 25–31 (2007).
Zhang, Y., Morgan, M. J., Chen, K., Choksi, S. & Liu, Z. G. Induction of autophagy is essential for monocyte-macrophage differentiation. Blood 119, 2895–2905 (2012).
Nedjic, J., Aichinger, M., Emmerich, J., Mizushima, N. & Klein, L. Autophagy in thymic epithelium shapes the T-cell repertoire and is essential for tolerance. Nature 455, 396–400 (2008).
Schmid, D., Pypaert, M. & Munz, C. Antigen-loading compartments for major histocompatibility complex class II molecules continuously receive input from autophagosomes. Immunity 26, 79–92 (2007).
Travassos, L. H. et al. Nod1 and Nod2 direct autophagy by recruiting ATG16L1 to the plasma membrane at the site of bacterial entry. Nat. Immunol. 11, 55–62 (2010).
Kuma, A., Mizushima, N., Ishihara, N. & Ohsumi, Y. Formation of the ∼350-kDa Apg12–Apg5•Apg16 multimeric complex, mediated by Apg16 oligomerization, is essential for autophagy in yeast. J. Biol. Chem. 277, 18619–18625 (2002).
Mizushima, N., Noda, T. & Ohsumi, Y. Apg16p is required for the function of the Apg12p–Apg5p conjugate in the yeast autophagy pathway. EMBO J. 18, 3888–3896 (1999).
Lee, H. K., Lund, J. M., Ramanathan, B., Mizushima, N. & Iwasaki, A. Autophagy-dependent viral recognition by plasmacytoid dendritic cells. Science 315, 1398–1401 (2007).
Lunemann, J. D. & Munz, C. Autophagy in CD4+ T-cell immunity and tolerance. Cell Death Differ. 16, 79–86 (2009).
Harris, J. et al. Autophagy controls IL-1β secretion by targeting pro-IL-1β for degradation. J. Biol. Chem. 286, 9587–9597 (2011).
Jia, G., Cheng, G., Gangahar, D. M. & Agrawal, D. K. Insulin-like growth factor-1 and TNF-α regulate autophagy through c-jun N-terminal kinase and Akt pathways in human atherosclerotic vascular smooth cells. Immunol. Cell Biol. 84, 448–454 (2006).
Copetti, T., Demarchi, F. & Schneider, C. p65/RelA binds and activates the beclin 1 promoter. Autophagy 5, 858–859 (2009).
Nivon, M., Richet, E., Codogno, P., Arrigo, A. P. & Kretz-Remy, C. Autophagy activation by NFκB is essential for cell survival after heat shock. Autophagy 5, 766–783 (2009).
Djavaheri-Mergny, M. et al. NF-κB activation represses tumor necrosis factor-α-induced autophagy. J. Biol. Chem. 281, 30373–30382 (2006).
Trocoli, A. & Djavaheri-Mergny, M. The complex interplay between autophagy and NF-κB signaling pathways in cancer cells. Am. J. Cancer Res. 1, 629–649 (2011).
Criollo, A. et al. The IKK complex contributes to the induction of autophagy. EMBO J. 29, 619–631 (2010).
Mizgalska, D. et al. Interleukin-1-inducible MCPIP protein has structural and functional properties of RNase and participates in degradation of IL-1β mRNA. FEBS J. 276, 7386–7399 (2009).
Roy, A. & Kolattukudy, P. E. Monocyte chemotactic protein-induced protein (MCPIP) promotes inflammatory angiogenesis via sequential induction of oxidative stress, endoplasmic reticulum stress and autophagy. Cell. Signal. 24, 2123–2131 (2012).
Roy, A., Zhang, M., Saad, Y. & Kolattukudy, P. E. Antidicer RNAse activity of monocyte chemotactic protein-induced protein-1 is critical for inducing angiogenesis. Am. J. Physiol. Cell Physiol. 305, C1021–C1032 (2013).
Niu, J., Azfer, A., Zhelyabovska, O., Fatma, S. & Kolattukudy, P. E. Monocyte chemotactic protein (MCP)-1 promotes angiogenesis via a novel transcription factor, MCP-1-induced protein (MCPIP). J. Biol. Chem. 283, 14542–14551 (2008).
Niu, J., Wang, K., Zhelyabovska, O., Saad, Y. & Kolattukudy, P. E. MCP-1-induced protein promotes endothelial-like and angiogenic properties in human bone marrow monocytic cells. J. Pharmacol. Exp. Ther. 347, 288–297 (2013).
Wang, K., Niu, J., Kim, H. & Kolattukudy, P. E. Osteoclast precursor differentiation by MCPIP via oxidative stress, endoplasmic reticulum stress, and autophagy. J. Mol. Cell Biol. 3, 360–368 (2011).
Kruse, K. et al. Inefficient clearance of dying cells in patients with SLE: anti-dsDNA autoantibodies, MFG-E8, HMGB-1 and other players. Apoptosis 15, 1098–1113 (2010).
Avery, D. T. et al. BAFF selectively enhances the survival of plasmablasts generated from human memory B cells. J. Clin. Invest. 112, 286–297 (2003).
Thompson, J. S. et al. BAFF-R, a newly identified TNF receptor that specifically interacts with BAFF. Science 293, 2108–2111 (2001).
Pers, J. O. et al. BAFF overexpression is associated with autoantibody production in autoimmune diseases. Ann. NY Acad. Sci. 1050, 34–39 (2005).
Petri, M. A. et al. Baseline predictors of systemic lupus erythematosus flares: data from the combined placebo groups in the phase III belimumab trials. Arthritis Rheum. 65, 2143–2153 (2013).
Stohl, W. et al. B lymphocyte stimulator overexpression in patients with systemic lupus erythematosus: longitudinal observations. Arthritis Rheum. 48, 3475–3486 (2003).
Rickert, R. C., Jellusova, J. & Miletic, A. V. Signaling by the tumor necrosis factor receptor superfamily in B-cell biology and disease. Immunol. Rev. 244, 115–133 (2011).
Otipoby, K. L. et al. BAFF activates Akt and Erk through BAFF-R in an IKK1-dependent manner in primary mouse B cells. Proc. Natl. Acad. Sci. USA 105, 12435–12438 (2008).
Qian, Y. et al. Act1, a negative regulator in CD40- and BAFF-mediated B cell survival. Immunity 21, 575–587 (2004).
Kim, H. A., Seo, G. Y. & Kim, P. H. Macrophage-derived BAFF induces AID expression through the p38MAPK/CREB and JNK/AP-1 pathways. J. Leukoc. Biol. 89, 393–398 (2011).
Schweighoffer, E. et al. The BAFF receptor transduces survival signals by co-opting the B cell receptor signaling pathway. Immunity 38, 475–488 (2013).
Clarke, A. J. et al. Autophagy is activated in systemic lupus erythematosus and required for plasmablast development. Ann. Rheum. Dis. 74, 912–920 (2015).
Dong, G. et al. STS-1 promotes IFN-α induced autophagy by activating the JAK1–STAT1 signaling pathway in B cells. Eur. J. Immunol. 45, 2377–2388 (2015).
Li, B., Yue, Y., Dong, C., Shi, Y. & Xiong, S. Blockade of macrophage autophagy ameliorates activated lymphocytes-derived DNA induced murine lupus possibly via inhibition of proinflammatory cytokine production. Clin. Exp. Rheumatol. 32, 705–714 (2014).
Weindel, C. G. et al. B cell autophagy mediates TLR7-dependent autoimmunity and inflammation. Autophagy 11, 1010–1024 (2015).
Lopez, P., Alonso-Perez, E., Rodriguez-Carrio, J. & Suarez, A. Influence of Atg5 mutation in SLE depends on functional IL-10 genotype. PLoS ONE 8, e78756 (2013).
Jarvinen, T. M. et al. Replication of GWAS-identified systemic lupus erythematosus susceptibility genes affirms B-cell receptor pathway signalling and strengthens the role of IRF5 in disease susceptibility in a Northern European population. Rheumatology 51, 87–92 (2012).
Zhang, Y. M. et al. Rare variants of ATG5 are likely to be associated with chinese patients with systemic lupus erythematosus. Medicine 94, e939 (2015).
Wofsy, D. & Seaman, W. E. Reversal of advanced murine lupus in NZB/NZW F1 mice by treatment with monoclonal antibody to L3T4. J. Immunol. 138, 3247–3253 (1987).
Mihara, M. et al. Immunologic abnormality in NZB/NZW F1 mice. Thymus-independent occurrence of B cell abnormality and requirement for T cells in the development of autoimmune disease, as evidenced by an analysis of the athymic nude individuals. J. Immunol. 141, 85–90 (1988).
La Cava, A. Lupus and T cells. Lupus 18, 196–201 (2009).
Pierdominici, M. et al. Role of autophagy in immunity and autoimmunity, with a special focus on systemic lupus erythematosus. FASEB J. 26, 1400–1412 (2012).
Alessandri, C. et al. T lymphocytes from patients with systemic lupus erythematosus are resistant to induction of autophagy. FASEB J. 26, 4722–4732 (2012).
Gros, F. et al. Macroautophagy is deregulated in murine and human lupus T lymphocytes. Autophagy 8, 1113–1123 (2012).
Li, C. et al. Autophagy is induced in CD4+ T cells and important for the growth factor-withdrawal cell death. J. Immunol. 177, 5163–5168 (2006).
Martinez, J. et al. Molecular characterization of LC3-associated phagocytosis reveals distinct roles for Rubicon, NOX2 and autophagy proteins. Nat. Cell Biol. 17, 893–906 (2015).
Martinez, J. et al. Noncanonical autophagy inhibits the autoinflammatory, lupus-like response to dying cells. Nature 533, 115–119 (2016).
Pope, R. M. Apoptosis as a therapeutic tool in rheumatoid arthritis. Nat. Rev. Immunol. 2, 527–535 (2002).
Steenvoorden, M. M. et al. Transition of healthy to diseased synovial tissue in rheumatoid arthritis is associated with gain of mesenchymal/fibrotic characteristics. Arthritis Res. Ther. 8, R165 (2006).
Huber, L. C. et al. Synovial fibroblasts: key players in rheumatoid arthritis. Rheumatology 45, 669–675 (2006).
Xu, K. et al. Reduced apoptosis correlates with enhanced autophagy in synovial tissues of rheumatoid arthritis. Inflamm. Res. 62, 229–237 (2013).
Kato, M., Ospelt, C., Gay, R. E., Gay, S. & Klein, K. Dual role of autophagy in stress-induced cell death in rheumatoid arthritis synovial fibroblasts. Arthritis Rheumatol. 66, 40–48 (2014).
Shin, Y. J. et al. Autophagy induction and CHOP under-expression promotes survival of fibroblasts from rheumatoid arthritis patients under endoplasmic reticulum stress. Arthritis Res. Ther. 12, R19 (2010).
Xu, K. et al. Autophagy induction contributes to the resistance to methotrexate treatment in rheumatoid arthritis fibroblast-like synovial cells through high mobility group box chromosomal protein 1. Arthritis Res. Ther. 17, 374 (2015).
Wegner, N. et al. Autoimmunity to specific citrullinated proteins gives the first clues to the etiology of rheumatoid arthritis. Immunol. Rev. 233, 34–54 (2010).
Scally, S. W. et al. A molecular basis for the association of the HLA-DRB1 locus, citrullination, and rheumatoid arthritis. J. Exp. Med. 210, 2569–2582 (2013).
Ireland, J. M. & Unanue, E. R. Autophagy in antigen-presenting cells results in presentation of citrullinated peptides to CD4 T cells. J. Exp. Med. 208, 2625–2632 (2011).
Ireland, J. M. & Unanue, E. R. Processing of proteins in autophagy vesicles of antigen-presenting cells generates citrullinated peptides recognized by the immune system. Autophagy 8, 429–430 (2012).
Sorice, M. et al. Autophagy generates citrullinated peptides in human synoviocytes: a possible trigger for anti-citrullinated peptide antibodies. Rheumatology (Oxford) http://dx.doi.org/10.1093/rheumatology/kew178 (2016).
Schett, G. & Gravallese, E. Bone erosion in rheumatoid arthritis: mechanisms, diagnosis and treatment. Nat. Rev. Rheumatol. 8, 656–664 (2012).
Liu, F. et al. Suppression of autophagy by FIP200 deletion leads to osteopenia in mice through the inhibition of osteoblast terminal differentiation. J. Bone Miner. Res. 28, 2414–2430 (2013).
Nollet, M. et al. Autophagy in osteoblasts is involved in mineralization and bone homeostasis. Autophagy 10, 1965–1977 (2014).
Lin, N. Y. et al. Autophagy regulates TNFα-mediated joint destruction in experimental arthritis. Ann. Rheum. Dis. 72, 761–768 (2013).
Greene, M. A. & Loeser, R. F. Aging-related inflammation in osteoarthritis. Osteoarthritis Cartilage 23, 1966–1971 (2015).
Kapoor, M., Martel-Pelletier, J., Lajeunesse, D., Pelletier, J. P. & Fahmi, H. Role of proinflammatory cytokines in the pathophysiology of osteoarthritis. Nat. Rev. Rheumatol. 7, 33–42 (2011).
Sasaki, H. et al. Autophagy modulates osteoarthritis-related gene expression in human chondrocytes. Arthritis Rheum. 64, 1920–1928 (2012).
Zhang, Y. et al. Cartilage-specific deletion of mTOR upregulates autophagy and protects mice from osteoarthritis. Ann. Rheum. Dis. 74, 1432–1440 (2015).
Carames, B. et al. Mechanical injury suppresses autophagy regulators and pharmacologic activation of autophagy results in chondroprotection. Arthritis Rheum. 64, 1182–1192 (2012).
Cinque, L. et al. FGF signalling regulates bone growth through autophagy. Nature 528, 272–275 (2015).
Carames, B., Olmer, M., Kiosses, W. B. & Lotz, M. K. The relationship of autophagy defects to cartilage damage during joint aging in a mouse model. Arthritis Rheumatol. 67, 1568–1576 (2015).
Hui, W. et al. Oxidative changes and signalling pathways are pivotal in initiating age-related changes in articular cartilage. Ann. Rheum. Dis. 75, 449–458 (2016).
Vuppalapati, K. K. et al. Targeted deletion of autophagy genes Atg5 or Atg7 in the chondrocytes promotes caspase-dependent cell death and leads to mild growth retardation. J. Bone Miner. Res. 30, 2249–2261 (2015).
Bouderlique, T. et al. Targeted deletion of Atg5 in chondrocytes promotes age-related osteoarthritis. Ann. Rheum. Dis. 75, 627–631 (2016).
Marion-Letellier, R., Savoye, G. & Ghosh, S. Fatty acids, eicosanoids and PPAR gamma. Eur. J. Pharmacol. http://dx.doi.org/10.1016/j.ejphar.2015.11.004 (2015).
Meirhaeghe, A. & Amouyel, P. Impact of genetic variation of PPARγ in humans. Mol. Genet. Metab. 83, 93–102 (2004).
Vasheghani, F. et al. PPARγ deficiency results in severe, accelerated osteoarthritis associated with aberrant mTOR signalling in the articular cartilage. Ann. Rheum. Dis. 74, 569–578 (2015).
Jiang, C., Ting, A. T. & Seed, B. PPAR-γ agonists inhibit production of monocyte inflammatory cytokines. Nature 391, 82–86 (1998).
Afif, H. et al. Peroxisome proliferator-activated receptor γ1 expression is diminished in human osteoarthritic cartilage and is downregulated by interleukin-1β in articular chondrocytes. Arthritis Res. Ther. 9, R31 (2007).
Dell'Accio, F. & Sherwood, J. PPARγ/mTOR signalling: striking the right balance in cartilage homeostasis. Ann. Rheum. Dis. 74, 477–479 (2015).
Ho, Y. Y., Lagares, D., Tager, A. M. & Kapoor, M. Fibrosis — a lethal component of systemic sclerosis. Nat. Rev. Rheumatol. 10, 390–402 (2014).
Frech, T. et al. Autophagy is a key feature in the pathogenesis of systemic sclerosis. Rheumatol. Int. 34, 435–439 (2014).
Dumit, V. I. et al. Altered MCM protein levels and autophagic flux in aged and systemic sclerosis dermal fibroblasts. J. Invest. Dermatol. 134, 2321–2330 (2014).
Varga, J. & Pasche, B. Transforming growth factor β as a therapeutic target in systemic sclerosis. Nat. Rev. Rheumatol. 5, 200–206 (2009).
Kim, S. I. et al. Autophagy promotes intracellular degradation of type I collagen induced by transforming growth factor (TGF)-β1. J. Biol. Chem. 287, 11677–11688 (2012).
Ghavami, S. et al. Autophagy is a regulator of TGF-β1-induced fibrogenesis in primary human atrial myofibroblasts. Cell Death Dis. 6, e1696 (2015).
Ding, Y. et al. Autophagy regulates TGF-β expression and suppresses kidney fibrosis induced by unilateral ureteral obstruction. J. Am. Soc. Nephrol. 25, 2835–2846 (2014).
Razani, B. et al. Caveolin-1 regulates transforming growth factor (TGF)-β/SMAD signaling through an interaction with the TGF-β type I receptor. J. Biol. Chem. 276, 6727–6738 (2001).
Del Galdo, F. et al. Decreased expression of caveolin 1 in patients with systemic sclerosis: crucial role in the pathogenesis of tissue fibrosis. Arthritis Rheum. 58, 2854–2865 (2008).
Castello-Cros, R. et al. Scleroderma-like properties of skin from caveolin-1-deficient mice: implications for new treatment strategies in patients with fibrosis and systemic sclerosis. Cell Cycle 10, 2140–2150 (2011).
Shi, J. H. et al. Reduced expression of microtubule-associated protein 1 light chain 3 in hypertrophic scars. Arch. Dermatol. Res. 304, 209–215 (2012).
Quan, T., Shao, Y., He, T., Voorhees, J. J. & Fisher, G. J. Reduced expression of connective tissue growth factor (CTGF/CCN2) mediates collagen loss in chronologically aged human skin. J. Invest. Dermatol. 130, 415–424 (2010).
Dekker, P. et al. Microarray-based identification of age-dependent differences in gene expression of human dermal fibroblasts. Mech. Ageing Dev. 133, 498–507 (2012).
Beyer, C. et al. β-catenin is a central mediator of pro-fibrotic Wnt signaling in systemic sclerosis. Ann. Rheum. Dis. 71, 761–767 (2012).
Akhmetshina, A. et al. Activation of canonical Wnt signalling is required for TGF-β-mediated fibrosis. Nat. Commun. 3, 735 (2012).
Gao, C. et al. Autophagy negatively regulates Wnt signalling by promoting Dishevelled degradation. Nat. Cell Biol. 12, 781–790 (2010).
Inoki, K. et al. TSC2 integrates Wnt and energy signals via a coordinated phosphorylation by AMPK and GSK3 to regulate cell growth. Cell 126, 955–968 (2006).
Kapoor, M., Kojima, F., Yang, L. & Crofford, L. J. Sequential induction of pro- and anti-inflammatory prostaglandins and peroxisome proliferators-activated receptor-gamma during normal wound healing: a time course study. Prostaglandins Leukot. Essent. Fatty Acids 76, 103–112 (2007).
Kapoor, M. et al. Loss of peroxisome proliferator-activated receptor γ in mouse fibroblasts results in increased susceptibility to bleomycin-induced skin fibrosis. Arthritis Rheum. 60, 2822–2829 (2009).
Wei, J. et al. PPARγ downregulation by TGFβ in fibroblast and impaired expression and function in systemic sclerosis: a novel mechanism for progressive fibrogenesis. PLoS ONE 5, e13778 (2010).
Pan, H. W., Xu, J. T. & Chen, J. S. Pioglitazone inhibits TGFβ induced keratocyte transformation to myofibroblast and extracellular matrix production. Mol. Biol. Rep. 38, 4501–4508 (2011).
Brown, E. J. et al. A mammalian protein targeted by G1-arresting rapamycin–receptor complex. Nature 369, 756–758 (1994).
Pallet, N. & Legendre, C. Adverse events associated with mTOR inhibitors. Expert Opin. Drug Saf. 12, 177–186 (2013).
Bruyn, G. A. et al. Everolimus in patients with rheumatoid arthritis receiving concomitant methotrexate: a 3-month, double-blind, randomised, placebo-controlled, parallel-group, proof-of-concept study. Ann. Rheum. Dis. 67, 1090–1095 (2008).
Takayama, K. et al. Local intra-articular injection of rapamycin delays articular cartilage degeneration in a murine model of osteoarthritis. Arthritis Res. Ther. 16, 482 (2014).
Matsuzaki, T. et al. Intra-articular administration of gelatin hydrogels incorporating rapamycin-micelles reduces the development of experimental osteoarthritis in a murine model. Biomaterials 35, 9904–9911 (2014).
Carames, B. et al. Autophagy activation by rapamycin reduces severity of experimental osteoarthritis. Ann. Rheum. Dis. 71, 575–581 (2012).
Yoshizaki, A. et al. Treatment with rapamycin prevents fibrosis in tight-skin and bleomycin-induced mouse models of systemic sclerosis. Arthritis Rheum. 62, 2476–2487 (2010).
Su, T. I. et al. Rapamycin versus methotrexate in early diffuse systemic sclerosis: results from a randomized, single-blind pilot study. Arthritis Rheum. 60, 3821–3830 (2009).
Kalia, S. & Dutz, J. P. New concepts in antimalarial use and mode of action in dermatology. Dermatol. Ther. 20, 160–174 (2007).
Amaravadi, R. K. et al. Principles and current strategies for targeting autophagy for cancer treatment. Clin. Cancer Res. 17, 654–666 (2011).
Marceau, F., Bawolak, M. T., Bouthillier, J. & Morissette, G. Vacuolar ATPase-mediated cellular concentration and retention of quinacrine: a model for the distribution of lipophilic cationic drugs to autophagic vacuoles. Drug Metab. Dispos. 37, 2271–2274 (2009).
Liu, Q. et al. Hydroxychloroquine facilitates autophagosome formation but not degradation to suppress the proliferation of cervical cancer SiHa cells. Oncol. Lett. 7, 1057–1062 (2014).
Ding, H. J., Denniston, A. K., Rao, V. K. & Gordon, C. Hydroxychloroquine-related retinal toxicity. Rheumatology (Oxford) 55, 957–967 (2016).
Zhou, G. et al. Role of AMP-activated protein kinase in mechanism of metformin action. J. Clin. Invest. 108, 1167–1174 (2001).
Phielix, E., Szendroedi, J. & Roden, M. The role of metformin and thiazolidinediones in the regulation of hepatic glucose metabolism and its clinical impact. Trends Pharmacol. Sci. 32, 607–616 (2011).
Teranishi, T. et al. Effects of pioglitazone and metformin on intracellular lipid content in liver and skeletal muscle of individuals with type 2 diabetes mellitus. Metabolism 56, 1418–1424 (2007).
Egan, D. F. et al. Phosphorylation of ULK1 (hATG1) by AMP-activated protein kinase connects energy sensing to mitophagy. Science 331, 456–461 (2011).
Arai, M. et al. Metformin, an antidiabetic agent, suppresses the production of tumor necrosis factor and tissue factor by inhibiting early growth response factor-1 expression in human monocytes in vitro. J. Pharmacol. Exp. Ther. 334, 206–213 (2010).
Hyun, B. et al. Metformin down-regulates TNF-α secretion via suppression of scavenger receptors in macrophages. Immune Netw. 13, 123–132 (2013).
Yan, H., Zhou, H. F., Hu, Y. & Pham, C. T. Suppression of experimental arthritis through AMP-activated protein kinase activation and autophagy modulation. J. Rheum. Dis. Treat. 1, 5 (2015).
Bennett, W. L. et al. Oral Diabetes Medications for Adults With Type 2 Diabetes: An Update (Agency for Healthcare Research and Quality (US), 2011).
Ji, M. M. et al. Induction of autophagy by valproic acid enhanced lymphoma cell chemosensitivity through HDAC-independent and IP3-mediated PRKAA activation. Autophagy 11, 2160–2171 (2015).
Gaitatzis, A. & Sander, J. W. The long-term safety of antiepileptic drugs. CNS Drugs 27, 435–455 (2013).
Nissen, S. E. & Wolski, K. Effect of rosiglitazone on the risk of myocardial infarction and death from cardiovascular causes. N. Engl. J. Med. 356, 2457–2471 (2007).
Lehmann, J. M. et al. An antidiabetic thiazolidinedione is a high affinity ligand for peroxisome proliferator-activated receptor γ (PPARγ). J. Biol. Chem. 270, 12953–12956 (1995).
Oakes, N. D. et al. A new antidiabetic agent, BRL 49653, reduces lipid availability and improves insulin action and glucoregulation in the rat. Diabetes 43, 1203–1210 (1994).
Miyazaki, Y., Matsuda, M. & DeFronzo, R. A. Dose-response effect of pioglitazone on insulin sensitivity and insulin secretion in type 2 diabetes. Diabetes Care 25, 517–523 (2002).
Bortolini, M., Wright, M. B., Bopst, M. & Balas, B. Examining the safety of PPAR agonists — current trends and future prospects. Expert Opin. Drug Saf. 12, 65–79 (2013).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT00554853 (2013).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT00763139 (2014).
Ormseth, M. J. et al. Peroxisome proliferator-activated receptor γ agonist effect on rheumatoid arthritis: a randomized controlled trial. Arthritis Res. Ther. 15, R110 (2013).
Ormseth, M. J. et al. Reversing vascular dysfunction in rheumatoid arthritis: improved augmentation index but not endothelial function with peroxisome proliferator-activated receptor gamma agonist therapy. Arthritis Rheumatol. 66, 2331–2338 (2014).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT01322308 (2014).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT02338999 (2016).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT00379600 (2012).
Gierman, L. M. et al. Metabolic stress-induced inflammation plays a major role in the development of osteoarthritis in mice. Arthritis Rheum. 64, 1172–1181 (2012).
Antonelli, A. et al. Peroxisome proliferator-activated receptor γ agonists reduce cell proliferation and viability and increase apoptosis in systemic sclerosis fibroblasts. Br. J. Dermatol. 168, 129–135 (2013).
Wu, M. et al. Rosiglitazone abrogates bleomycin-induced scleroderma and blocks profibrotic responses through peroxisome proliferator-activated receptor-γ. Am. J. Pathol. 174, 519–533 (2009).
Macri, C. et al. Modulation of deregulated chaperone-mediated autophagy by a phosphopeptide. Autophagy 11, 472–486 (2015).
Zimmer, R., Scherbarth, H. R., Rillo, O. L., Gomez-Reino, J. J. & Muller, S. Lupuzor/P140 peptide in patients with systemic lupus erythematosus: a randomised, double-blind, placebo-controlled phase IIb clinical trial. Ann. Rheum. Dis. 72, 1830–1835 (2013).
Muller, S. et al. Spliceosomal peptide P140 for immunotherapy of systemic lupus erythematosus: results of an early phase II clinical trial. Arthritis Rheum. 58, 3873–3883 (2008).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT02504645 (2016).
Muller, S. & Wallace, D. J. The importance of implementing proper selection of excipients in lupus clinical trials. Lupus 23, 609–614 (2014).
DeBosch, B. J. et al. Trehalose inhibits solute carrier 2A (SLC2A) proteins to induce autophagy and prevent hepatic steatosis. Sci. Signal. 9, ra21 (2016).
Navarra, S. V. et al. Efficacy and safety of belimumab in patients with active systemic lupus erythematosus: a randomised, placebo-controlled, phase 3 trial. Lancet 377, 721–731 (2011).
Bruce, I. N. et al. Long-term organ damage accrual and safety in patients with SLE treated with belimumab plus standard of care. Lupus http://dx.doi.org/10.1177/0961203315625119 (2016).
Wallace, D. J. et al. Safety profile of belimumab: pooled data from placebo-controlled phase 2 and 3 studies in patients with systemic lupus erythematosus. Lupus 22, 144–154 (2013).
Taylor, P. C. & Feldmann, M. Anti-TNF biologic agents: still the therapy of choice for rheumatoid arthritis. Nat. Rev. Rheumatol. 5, 578–582 (2009).
Ramiro, S. et al. Safety of synthetic and biological DMARDs: a systematic literature review informing the 2013 update of the EULAR recommendations for management of rheumatoid arthritis. Ann. Rheum. Dis. 73, 529–535 (2014).
Connor, A. M., Mahomed, N., Gandhi, R., Keystone, E. C. & Berger, S. A. TNFα modulates protein degradation pathways in rheumatoid arthritis synovial fibroblasts. Arthritis Res. Ther. 14, R62 (2012).
Selmi, C., Ceribelli, A., Naguwa, S. M., Cantarini, L. & Shoenfeld, Y. Safety issues and concerns of new immunomodulators in rheumatology. Expert Opin. Drug Saf. 14, 389–399 (2015).
Rice, L. M. et al. Fresolimumab treatment decreases biomarkers and improves clinical symptoms in systemic sclerosis patients. J. Clin. Invest. 125, 2795–2807 (2015).
Lacouture, M. E. et al. Cutaneous keratoacanthomas/squamous cell carcinomas associated with neutralization of transforming growth factor β by the monoclonal antibody fresolimumab (GC1008). Cancer Immunol. Immunother. 64, 437–446 (2015).
Morris, J. C. et al. Phase I study of GC1008 (fresolimumab): a human anti-transforming growth factor-beta (TGFβ) monoclonal antibody in patients with advanced malignant melanoma or renal cell carcinoma. PLoS ONE 9, e90353 (2014).
Stevenson, J. P. et al. Immunological effects of the TGFβ-blocking antibody GC1008 in malignant pleural mesothelioma patients. Oncoimmunology 2, e26218 (2013).
Trachtman, H. et al. A phase 1, single-dose study of fresolimumab, an anti-TGF-β antibody, in treatment-resistant primary focal segmental glomerulosclerosis. Kidney Int. 79, 1236–1243 (2011).
Sosulski, M. L. et al. Deregulation of selective autophagy during aging and pulmonary fibrosis: the role of TGFβ1. Aging Cell 14, 774–783 (2015).
Vingtdeux, V. et al. AMP-activated protein kinase signaling activation by resveratrol modulates amyloid-β peptide metabolism. J. Biol. Chem. 285, 9100–9113 (2010).
Xuzhu, G. et al. Resveratrol modulates murine collagen-induced arthritis by inhibiting Th17 and B-cell function. Ann. Rheum. Dis. 71, 129–135 (2012).
Heraud, F., Heraud, A. & Harmand, M. F. Apoptosis in normal and osteoarthritic human articular cartilage. Ann. Rheum. Dis. 59, 959–965 (2000).
Chia, S. et al. Novel agents and associated toxicities of inhibitors of the pi3k/Akt/mtor pathway for the treatment of breast cancer. Curr. Oncol. 22, 33–48 (2015).
Asahina, H. et al. Safety and tolerability of AZD8055 in Japanese patients with advanced solid tumors; a dose-finding phase I study. Invest. New Drugs 31, 677–684 (2013).
Bendell, J. C. et al. A phase I dose-escalation study to assess safety, tolerability, pharmacokinetics, and preliminary efficacy of the dual mTORC1/mTORC2 kinase inhibitor CC-223 in patients with advanced solid tumors or multiple myeloma. Cancer 121, 3481–3490 (2015).
Naing, A. et al. Safety, tolerability, pharmacokinetics and pharmacodynamics of AZD8055 in advanced solid tumours and lymphoma. Br. J. Cancer 107, 1093–1099 (2012).
Britten, C. D. et al. Phase I study of PF-04691502, a small-molecule, oral, dual inhibitor of PI3K and mTOR, in patients with advanced cancer. Invest. New Drugs 32, 510–517 (2014).
Shapiro, G. I. et al. First-in-human study of PF-05212384 (PKI-587), a small-molecule, intravenous, dual inhibitor of PI3K and mTOR in patients with advanced cancer. Clin. Cancer Res. 21, 1888–1895 (2015).
Seront, E. et al. Phase II study of dual phosphoinositol-3-kinase (PI3K) and mammalian target of rapamycin (mTOR) inhibitor BEZ235 in patients with locally advanced or metastatic transitional cell carcinoma (TCC). BJU Int. http://dx.doi.org/10.1111/bju.13415 (2016).
Dolly, S. et al. Phase I study of apitolisib (GDC-0980), dual phosphatidylinositol-3-kinase and mammalian target of rapamycin kinase inhibitor, in patients with advanced solid tumors. Clin. Cancer Res. http://dx.doi.org/10.1158/1078-0432.CCR-15-2225 (2016).
Basu, B. et al. First-in-human pharmacokinetic and pharmacodynamic study of the dual m-TORC 1/2 inhibitor AZD2014. Clin. Cancer Res. 21, 3412–3419 (2015).
Chen, J., Crawford, R. & Xiao, Y. Vertical inhibition of the PI3K/Akt/mTOR pathway for the treatment of osteoarthritis. J. Cell. Biochem. 114, 245–249 (2013).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT00779194 (2015).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT01914770 (2013).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT00392951 (2016).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT01946880 (2016).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT02349061 (2016).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT02246257 (2015).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT02644499 (2016).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT01941940 (2015).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT01256736 (2013).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT01211834 (2010).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT00805519 (2010).
US National Library of Science. ClinicalTrials.govhttps://clinicaltrials.gov/show/NCT02166229 (2014).
Acknowledgements
The authors' work is supported by the Canadian Institutes of Health Research Operating Grant 126016 to M.K. and the Arthritis Program Grant to M.K. The authors would like to acknowledge the help of Brian Wu in designing figures for this Review.
Author information
Authors and Affiliations
Contributions
Both authors researched data for this article, made substantial contributions to discussions of the content, wrote the article and reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Rockel, J., Kapoor, M. Autophagy: controlling cell fate in rheumatic diseases. Nat Rev Rheumatol 12, 517–531 (2016). https://doi.org/10.1038/nrrheum.2016.92
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrrheum.2016.92
This article is cited by
-
The “Intermediate” CD14 + CD16 + monocyte subpopulation plays a role in IVIG responsiveness of children with Kawasaki disease
Pediatric Rheumatology (2021)
-
Focus on the Multimodal Role of Autophagy in Rheumatoid Arthritis
Inflammation (2021)
-
Intravenous immunoglobulin mediates anti-inflammatory effects in peripheral blood mononuclear cells by inducing autophagy
Cell Death & Disease (2020)
-
Deep sequencing reveals a DAP1 regulatory haplotype that potentiates autoimmunity in systemic lupus erythematosus
Genome Biology (2020)
-
The relationship between autophagy and the immune system and its applications for tumor immunotherapy
Molecular Cancer (2019)