Abstract
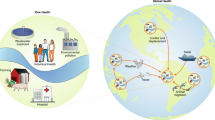
Bacterial resistance to antibiotics continues to pose a serious threat to human and animal health. Given the considerable spatial and temporal heterogeneity in the distribution of resistance and the factors that affect its evolution, dissemination and persistence, we argue that antibiotic resistance must be viewed as an ecological problem. A fundamental difficulty in assessing the causal relationship between antibiotic use and resistance is the confounding influence of geography: the co-localization of resistant bacterial species with antibiotic use does not necessarily imply causation and could represent the presence of environmental conditions and factors that have independently contributed to the occurrence of resistance. Here, we show how landscape ecology, which links the biotic and abiotic factors of an ecosystem, might help to untangle the complexity of antibiotic resistance and improve the interpretation of ecological studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Salyers, A. A. & Amabile-Cuevas, C. F. Why are antibiotic resistance genes so resistant to elimination? Antimicrob. Agents Chemother. 41, 2321–2325 (1997).
Salyers, A. A., Shoemaker, N. B. & Bonheyo, G. T. in Bacterial Resistance to Antimicrobials (eds Lewis, K., Salyers, A. A., Taber, H. W. & Wax, R. G.) 1–18 (Marcel Dekker, New York, 2002).
Summers, A. O. Generally overlooked fundamentals of bacterial genetics and ecology. Clin. Infect. Dis. 34, (Suppl. 3), S85–S92 (2002).
Aarestrup, F. M., Kruse, H., Tast, E., Hammerum, A. M. & Jensen, L. B. Associations between the use of antimicrobial agents for growth promotion and the occurrence of resistance among Enterococcus faecium from broilers and pigs in Denmark, Finland, and Norway. Microb. Drug Resist. 6, 63–70 (2000).
Aarestrup, F. M. et al. Effect of abolishment of the use of antimicrobial agents for growth promotion on occurrence of antimicrobial resistance in fecal enterococci from food animals in Denmark. Antimicrob. Agents Chemother. 45, 2054–2059 (2001).
Casewell, M., Friis, C., Marco, E., McMullin, P. & Phillips, I. The European ban on growth-promoting antibiotics and emerging consequences for human and animal health. J. Antimicrob. Chemother. 52, 159–161 (2003).
Heuer, O. E., Pedersen, K., Andersen, J. S. & Madsen, M. Vancomycin-resistant enterococci (VRE) in broiler flocks 5 years after the avoparcin ban. Microb. Drug Resist. 8, 133–138 (2002).
Heuer, O. E., Pedersen, K., Jensen, L. B., Madsen, M. & Olsen, J. E. Persistence of vancomycin-resistant enterococci (VRE) in broiler houses after the avoparcin ban. Microb. Drug Resist. 8, 355–361 (2002).
Phillips, I. et al. Does the use of antibiotics in food animals pose a risk to human health? A critical review of published data. J. Antimicrob. Chemother. 53, 28–52 (2004).
Kieser, T., Bibb, M. J., Buttner, M. J., Chater, K. F. & Hopwood, D. A. Practical Streptomyces Genetics (John Innes Centre, Norwich, UK, 2000).
Waksman, S. The role of antibiotics in nature. Perspect. Biol. Med. 4, 271–287 (1961).
Yim, G., Wang, H. H. & Davies, J. The truth about antibiotics. Int. J. Med. Microbiol. 296, 163–170 (2006).
D'Costa, V. M., McGrann, K. M., Hughes, D. W. & Wright, G. D. Sampling the antibiotic resistome. Science 311, 374–377 (2006).
Davies, J. Are antibiotics naturally antibiotics? J. Ind. Microbiol. Biotechnol. 33, 496–499 (2006).
Schmitt, H., Stoob, K., Hamscher, G., Smit, E. & Seinen, W. Tetracyclines and tetracycline resistance in agricultural soils: microcosm and field studies. Microb. Ecol. 51, 267–276 (2006).
Kummerer, K. Drugs in the environment: emission of drugs, diagnostic aids and disinfectants into wastewater by hospitals in relation to other sources — a review. Chemosphere 45, 957–969 (2001).
Kummerer, K. & Henninger, A. Promoting resistance by the emission of antibiotics from hospitals and households into effluent. Clin. Microbiol. Infect. 9, 1203–1214 (2003).
Kolpin, D. W. et al. Pharmaceuticals, hormones, and other organic wastewater contaminants in U.S. streams, 1999–2000: a national reconnaissance. Environ. Sci. Technol. 36, 1202–1211 (2002).
Schrag, S. J., Perrot, V. & Levin, B. R. Adaptation to the fitness costs of antibiotic resistance in Escherichia coli. Proc. Biol. Sci. 264, 1287–1291 (1997).
Andersson, D. I. & Levin, B. R. The biological cost of antibiotic resistance. Curr. Opin. Microbiol. 2, 489–493 (1999).
Lenski, R. E. Bacterial evolution and the cost of antibiotic resistance. Int. Microbiol. 1, 265–270 (1998).
Sorum, M. et al. Prevalence, persistence, and molecular characterization of glycopeptide-resistant enterococci in Norwegian poultry and poultry farmers 3 to 8 years after the ban on avoparcin. Appl. Environ. Microbiol. 72, 516–521 (2006).
Khachatryan, A. R., Hancock, D. D., Besser, T. E. & Call, D. R. Role of calf-adapted Escherichia coli in maintenance of antimicrobial drug resistance in dairy calves. Appl. Environ. Microbiol. 70, 752–757 (2004).
Khachatryan, A. R., Hancock, D. D., Besser, T. E. & Call, D. R. Antimicrobial drug resistance genes do not convey a secondary fitness advantage to calf-adapted Escherichia coli. Appl. Environ. Microbiol. 72, 443–448 (2006).
Luo, N., Sahin, O., Lin, J., Michel, L. O. & Zhang, Q. In vivo selection of Campylobacter isolates with high levels of fluoroquinolone resistance associated with gyrA mutations and the function of the CmeABC efflux pump. Antimicrob. Agents Chemother. 47, 390–394 (2003).
Sander, P. et al. Fitness cost of chromosomal drug resistance-conferring mutations. Antimicrob. Agents Chemother. 46, 1204–1211 (2002).
Levin, B. R., Perrot, V. & Walker, N. Compensatory mutations, antibiotic resistance and the population genetics of adaptive evolution in bacteria. Genetics 154, 985–997 (2000).
Forman, R. T. T. Land Mosaics: The Ecology of Landscapes and Regions (Cambridge University Press, Cambridge UK, 1995).
Martiny, J. B. et al. Microbial biogeography: putting microorganisms on the map. Nature Rev. Microbiol. 4, 102–112 (2006).
Pavlovsky, E. N. Natural Nidality of Transmissible Diseases: With Special Reference to the Landscape Ecology of Zooanthroponoses. (University of Illinois Press, Urbana Illinois, 1966).
Kitron, U. Landscape ecology and epidemiology of vector-borne diseases: tools for spatial analysis. J. Med. Entomol. 35, 435–445 (1998).
Reisen, W. K., Lothrop, H. D., Presser, S. B., Hardy, J. L. & Gordon, E. W. Landscape ecology of arboviruses in southeastern California: temporal and spatial patterns of enzootic activity in Imperial Valley, 1991–1994. J. Med. Entomol. 34, 179–188 (1997).
Guerra, M. et al. Predicting the risk of Lyme disease: habitat suitability for Ixodes scapularis in the north central United States. Emerg. Infect. Dis. 8, 289–297 (2002).
Pei, R., Kim, S. C., Carlson, K. H. & Pruden, A. Effect of river landscape on the sediment concentrations of antibiotics and corresponding antibiotic resistance genes (ARG). Water Res. 40, 2427–2435 (2006).
Unicomb, L. E. et al. Low-level fluoroquinolone resistance among Campylobacter jejuni isolates in Australia. Clin. Infect. Dis. 42, 1368–1374 (2006).
Davelos, A. L., Kinkel, L. L. & Samac, D. A. Spatial variation in frequency and intensity of antibiotic interactions among Streptomycetes from prairie soil. Appl. Environ. Microbiol. 70, 1051–1058 (2004).
Swaminathan, B., Barrett, T. J. & Fields, P. Surveillance for human Salmonella infections in the United States. J. AOAC Int. 89, 553–559 (2006).
Zhao, S. et al. Antimicrobial resistance and genetic relatedness among Salmonella from retail foods of animal origin: NARMS retail meat surveillance. Foodborne. Pathog. Dis. 3, 106–117 (2006).
Lipsitch, M. & Samore, M. H. Antimicrobial use and antimicrobial resistance: a population perspective. Emerg. Infect. Dis. 8, 347–354 (2002).
Harris, A. D. et al. Control-group selection importance in studies of antimicrobial resistance: examples applied to Pseudomonas aeruginosa, Enterococci, and Escherichia coli. Clin. Infect. Dis. 34, 1558–1563 (2002).
Lipsitch, M. Measuring and interpreting associations between antibiotic use and penicillin resistance in Streptococcus pneumoniae. Clin. Infect. Dis. 32, 1044–1054 (2001).
Rubin, D. B. Estimating causal effects of treatments in randomized and nonrandomized studies. J. Educ. Psychol. 66, 688–701 (1974).
Hofler, M. Causal inference based on counterfactuals. BMC. Med. Res. Methodol. 5, 28 (2005).
Maldonado, G. & Greenland, S. Estimating causal effects. Int. J. Epidemiol. 31, 422–429 (2002).
McGowan, J. E. Jr. Antimicrobial resistance in hospital organisms and its relation to antibiotic use. Rev. Infect. Dis. 5, 1033–1048 (1983).
Alonso, A., Sanchez, P. & Martinez, J. L. Environmental selection of antibiotic resistance genes. Environ. Microbiol. 3, 1–9 (2001).
Borgen, K., Sorum, M., Wasteson, Y., Kruse, H. & Oppegaard, H. Genetic linkage between erm(B) and vanA in Enterococcus hirae of poultry origin. Microb. Drug Resist. 8, 363–368 (2002).
Sander, J. E., Hofacre, C. L., Cheng, I. H. & Wyatt, R. D. Investigation of resistance of bacteria from commercial poultry sources to commercial disinfectants. Avian Dis. 46, 997–1000 (2002).
Sidhu, M. S., Sorum, H. & Holck, A. Resistance to quaternary ammonium compounds in food-related bacteria. Microb. Drug Resist. 8, 393–399 (2002).
Sidhu, M. S., Heir, E., Leegaard, T., Wiger, K. & Holck, A. Frequency of disinfectant resistance genes and genetic linkage with β-lactamase transposon Tn552 among clinical staphylococci. Antimicrob. Agents Chemother. 46, 2797–2803 (2002).
Sidhu, M. S., Heir, E., Sorum, H. & Holck, A. Genetic linkage between resistance to quaternary ammonium compounds and β-lactam antibiotics in food-related Staphylococcus spp. Microb. Drug Resist. 7, 363–371 (2001).
Guerra, B., Soto, S., Helmuth, R. & Mendoza, M. C. Characterization of a self-transferable plasmid from Salmonella enterica serotype typhimurium clinical isolates carrying two integron-borne gene cassettes together with virulence and drug resistance genes. Antimicrob. Agents Chemother. 46, 2977–2981 (2002).
Baker-Austin, C., Wright, M. S., Stepanauskas, R. & McArthur, J. V. Co-selection of antibiotic and metal resistance. Trends Microbiol. 14, 176–182 (2006).
Stepanauskas, R. et al. Coselection for microbial resistance to metals and antibiotics in freshwater microcosms. Environ. Microbiol. 8, 1510–1514 (2006).
Barkay, T., Miller, S. M. & Summers, A. O. Bacterial mercury resistance from atoms to ecosystems. FEMS Microbiol. Rev. 27, 355–384 (2003).
Wireman, J., Liebert, C. A., Smith, T. & Summers, A. O. Association of mercury resistance with antibiotic resistance in the Gram-negative fecal bacteria of primates. Appl. Environ. Microbiol. 63, 4494–4503 (1997).
Liebert, C. A., Hall, R. M. & Summers, A. O. Transposon Tn21, flagship of the floating genome. Microbiol. Mol. Biol. Rev. 63, 507–522 (1999).
Bass, L. et al. Incidence and characterization of integrons, genetic elements mediating multiple-drug resistance, in avian Escherichia coli. Antimicrob. Agents Chemother. 43, 2925–2929 (1999).
Berg, J., Tom-Petersen, A. & Nybroe, O. Copper amendment of agricultural soil selects for bacterial antibiotic resistance in the field. Lett. Appl. Microbiol. 40, 146–151 (2005).
Hasman, H. & Aarestrup, F. M. tcrB, a gene conferring transferable copper resistance in Enterococcus faecium: occurrence, transferability, and linkage to macrolide and glycopeptide resistance. Antimicrob. Agents Chemother. 46, 1410–1416 (2002).
Hasman, H. & Aarestrup, F. M. Relationship between copper, glycopeptide, and macrolide resistance among Enterococcus faecium strains isolated from pigs in Denmark between 1997 and 2003. Antimicrob. Agents Chemother. 49, 454–456 (2005).
Sarmah, A. K., Meyer, M. T. & Boxall, A. B. A global perspective on the use, sales, exposure pathways, occurrence, fate and effects of veterinary antibiotics (VAs) in the environment. Chemosphere 65, 725–759 (2006).
Rooklidge, S. J. Environmental antimicrobial contamination from terraccumulation and diffuse pollution pathways. Sci. Total Environ. 325, 1–13 (2004).
Chander, Y., Kumar, K., Goyal, S. M. & Gupta, S. C. Antibacterial activity of soil-bound antibiotics. J. Environ. Qual. 34, 1952–1957 (2005).
Clay, S. A., Liu, Z., Thaler, R. & Kennouche, H. Tylosin sorption to silty clay loam soils, swine manure, and sand. J. Environ. Sci. Health B 40, 841–850 (2005).
Aga, D. S. et al. Determination of the persistence of tetracycline antibiotics and their degradates in manure-amended soil using enzyme-linked immunosorbent assay and liquid chromatography-mass spectrometry. J. Agric. Food Chem. 53, 7165–7171 (2005).
Kumar, K., Gupta, S. C., Baidoo, S. K., Chander, Y. & Rosen, C. J. Antibiotic uptake by plants from soil fertilized with animal manure. J. Environ. Qual. 34, 2082–2085 (2005).
Kummerer, K. Resistance in the environment. J. Antimicrob. Chemother. 54, 311–320 (2004).
Novais, C., Coque, T. M., Ferreira, H., Sousa, J. C. & Peixe, L. Environmental contamination with vancomycin-resistant enterococci from hospital sewage in Portugal. Appl. Environ. Microbiol. 71, 3364–3368 (2005).
Lewis, D. J. et al. Linking on-farm dairy management practices to storm-flow fecal coliform loading for California coastal watersheds. Environ. Monit. Assess. 107, 407–425 (2005).
Rodgers, P., Soulsby, C., Hunter, C. & Petry, J. Spatial and temporal bacterial quality of a lowland agricultural stream in northeast Scotland. Sci. Total Environ. 314–316, 289–302 (2003).
Moore, J. A. Surface transport of microorganisms by water. Biotechnology 15, 41–55 (1991).
Gibbs, S. G., Green, C. F., Tarwater, P. M. & Scarpino, P. V. Airborne antibiotic resistant and nonresistant bacteria and fungi recovered from two swine herd confined animal feeding operations. J. Occup. Environ. Hyg. 1, 699–706 (2004).
Chapin, A., Rule, A., Gibson, K., Buckley, T. & Schwab, K. Airborne multidrug-resistant bacteria isolated from a concentrated swine feeding operation. Environ. Health Perspect. 113, 137–142 (2005).
Elliott, P., Wakefield, J., Best, N. & Briggs, D. Spatial epidemiology: Methods and Applications. (Oxford University Press, Oxford, 2000).
Lawson, A. B. Statistical Methods in Spatial Epidemiology. (John Wiley, Chichester 2001).
Hendrickx, G. et al. The spatial pattern of trypanosomosis prevalence predicted with the aid of satellite imagery. Parasitology 120, 121–134 (2000).
Kitron, U. & Kazmierczak, J. J. Spatial analysis of the distribution of Lyme disease in Wisconsin. Am. J. Epidemiol. 145, 558–566 (1997).
Chadee, D. D. & Kitron, U. Spatial and temporal patterns of imported malaria cases and local transmission in Trinidad. Am. J. Trop. Med. Hyg. 61, 513–517 (1999).
Kleinschmidt, I., Sharp, B., Mueller, I. & Vounatsou, P. Rise in malaria incidence rates in South Africa: a small-area spatial analysis of variation in time trends. Am. J. Epidemiol. 155, 257–264 (2002).
Ali, M., Emch, M., Yunus, M. & Sack, R. B. Are the environmental niches of Vibrio cholerae O139 different from those of Vibrio cholerae O1 El Tor? Int. J. Infect. Dis. 5, 214–219 (2001).
Ali, M., Emch, M., Donnay, J. P., Yunus, M. & Sack, R. B. The spatial epidemiology of cholera in an endemic area of Bangladesh. Soc. Sci. Med. 55, 1015–1024 (2002).
Myaux, J., Ali, M., Felsenstein, A., Chakraborty, J. & De Francisco, A. Spatial distribution of watery diarrhoea in children: identification of 'risk areas' in a rural community in Bangladesh. Health Place. 3, 181–186 (1997).
Cifuentes, E., Mazari-Hiriart, M., Carneiro, F., Bianchi, F. & Gonzalez, D. The risk of enteric diseases in young children and environmental indicators in sentinel areas of Mexico City. Int. J. Environ. Health Res. 12, 53–62 (2002).
Njemanze, P. C., Anozie, J., Ihenacho, J. O., Russell, M. J. & Uwaeziozi, A. B. Application of risk analysis and geographic information system technologies to the prevention of diarrheal diseases in Nigeria. Am. J. Trop. Med. Hyg. 61, 356–360 (1999).
Tobler, W. R. A computer movie simulating urban growth in the Detroit region. Econ. Geograph. 46, 234–240 (1970).
Singer, R. S., Case, J. T., Carpenter, T. E., Walker, R. L. & Hirsh, D. C. Assessment of spatial and temporal clustering of ampicillin- and tetracycline-resistant strains of Pasteurella multocida and P. haemolytica isolated from cattle in California. J. Am. Vet. Med. Assoc. 212, 1001–1005 (1998).
Metlay, J. P., Branas, C. C. & Fishman, N. O. Hospital-reported pneumococcal susceptibility to penicillin. Emerg. Infect. Dis. 10, 54–59 (2004).
Ward, M. P. & Carpenter, T. E. Techniques for analysis of disease clustering in space and in time in veterinary epidemiology. Prev. Vet. Med. 45, 257–284 (2000).
Singer, R. S., Reid-Smith, R. & Sischo, W. M. Stakeholder position paper: Epidemiological perspectives on antibiotic use in animals. Prev. Vet. Med. 73, 153–161 (2006).
Diez-Roux, A. V. Bringing context back into epidemiology: variables and fallacies in multilevel analysis. Am. J. Public Health 88, 216–222 (1998).
Dohoo, I. R., Martin, S. W. & Strynh, H. in Veterinary Epidemiologic Research. (Atlantic Veterinary College Inc., 2003).
Frost, L. S., Leplae, R., Summers, A. O. & Toussaint, A. Mobile genetic elements: the agents of open source evolution. Nature Rev. Microbiol. 3, 722–732 (2005).
Gogarten, J. P. & Townsend, J. P. Horizontal gene transfer, genome innovation and evolution. Nature Rev. Microbiol. 3, 679–687 (2005).
Thomas, C. M. & Nielsen, K. M. Mechanisms of, and barriers to, horizontal gene transfer between bacteria. Nature Rev. Microbiol. 3, 711–721 (2005).
Handelsman, J. Metagenomics: application of genomics to uncultured microorganisms. Microbiol. Mol. Biol. Rev. 68, 669–685 (2004).
Riesenfeld, C. S., Goodman, R. M. & Handelsman, J. Uncultured soil bacteria are a reservoir of new antibiotic resistance genes. Environ. Microbiol. 6, 981–989 (2004).
Franklin, R. B. & Mills, A. L. Multi-scale variation in spatial heterogeneity for microbial community structure in an eastern Virginia agricultural field. FEMS Microbiol. Ecol. 44, 335–346 (2003).
Yu, Z., Michel, F. C. Jr, Hansen, G., Wittum, T. & Morrison, M. Development and application of real-time PCR assays for quantification of genes encoding tetracycline resistance. Appl. Environ. Microbiol. 71, 6926–6933 (2005).
Smith, M. S. et al. Quantification of tetracycline resistance genes in feedlot lagoons by real-time PCR. Appl. Environ. Microbiol. 70, 7372–7377 (2004).
Hayes, J. R., Wagner, D. D., English, L. L., Carr, L. E. & Joseph, S. W. Distribution of streptogramin resistance determinants among Enterococcus faecium from a poultry production environment of the USA. J. Antimicrob. Chemother. 55, 123–126 (2005).
Acknowledgements
We thank Tim Boyer, Janet Anderson and the reviewers for many helpful suggestions in improving the manuscript. We also thank Jeffrey Bender for supplying the Minnesota STEC dataset that was used in this manuscript. R.S.S. was partially supported by a United States Department of Agriculture National Research Initiative Competitive Grant.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
Entrez Genome Project
FURTHER INFORMATION
Rights and permissions
About this article
Cite this article
Singer, R., Ward, M. & Maldonado, G. Can landscape ecology untangle the complexity of antibiotic resistance?. Nat Rev Microbiol 4, 943–952 (2006). https://doi.org/10.1038/nrmicro1553
Issue Date:
DOI: https://doi.org/10.1038/nrmicro1553
This article is cited by
-
The impact of public health interventions on the future prevalence of ESBL-producing Klebsiella pneumoniae: a population based mathematical modelling study
BMC Infectious Diseases (2022)
-
Quantifying and predicting antimicrobials and antimicrobial resistance genes in waterbodies through a holistic approach: a study in Minnesota, United States
Scientific Reports (2021)
-
Marine biofilms constitute a bank of hidden microbial diversity and functional potential
Nature Communications (2019)
-
Acquisition and dissemination of cephalosporin-resistant E. coli in migratory birds sampled at an Alaska landfill as inferred through genomic analysis
Scientific Reports (2018)
-
Suppressive drug combinations and their potential to combat antibiotic resistance
The Journal of Antibiotics (2017)