Key Points
-
Darwinian natural selection is a gradual process of change produced by random mutations followed by competitive selection. However, extant species indicate that some evolutionary changes have been more rapid than could occur by Darwinian evolution, and the theory of symbiogenesis, or evolution by the merging of dissimilar symbiotic entities, has gained support in recent years.
-
Plant RNA viruses were shown to compete in early studies on cross-protection, in which mild strains of a virus could protect plants from subsequent infection by more virulent strains. The requirement of this type of competition is that the strains are similar. Symbiosis involves a different paradigm from competition.
-
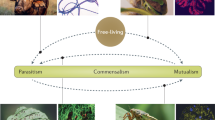
In mixed infections of plant viruses, there is a potential for symbiotic associations. The definition of symbiosis requires that the entities are dissimilar. Some well described examples of plant viruses that are symbiotic include potyviruses, which are often synergistic with other viruses; luteoviruses, which often require other viruses for dissemination; and satellites, which establish parasitic symbiosis with their helper viruses.
-
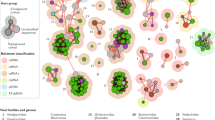
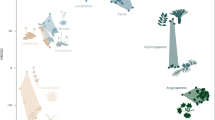
The effect of symbiosis on plant virus evolution can be seen in the modular nature of plant virus genes, which show evidence of recombination and reassortment in their evolutionary history when phylogenetic trees for different genes or genome components are analysed.
-
Symbiogenesis is probably a major component of plant virus evolution, allowing evolutionary leaps that could accommodate changing environments and host ranges. However, the gradual process of natural selection undoubtedly fine-tunes the resulting new species through random mutation and selection.
Abstract
Darwin's theory of evolution by natural selection has been supported by molecular evidence and by experimental evolution of viruses. However, it might not account for the evolution of all life, and an alternative model of evolution through symbiotic relationships also has gained support. In this review, the evolution of plant viruses has been reinterpreted in light of these two seemingly opposing theories by using evidence from the earliest days of plant virology to the present. Both models of evolution probably apply in different circumstances, but evolution by symbiotic association (symbiogenesis) is the most likely model for many evolutionary events that have resulted in rapid changes or the formation of new species. In viruses, symbiogenesis results in genomic reassortment or recombination events among disparate species. These are most noticeable by phylogenetic comparisons of extant viruses from different taxonomic groups.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Darwin, C. On the Origin of Species by Means of Natural Selection (John Murray, London, 1859).
Ryan, F. Darwin's Blind Spot (Houghton Mifflin Company, Boston, 2002). This excellent, clearly written book covers all aspects of the controversy between Darwinian evolution and symbiogenesis, and explains the role of symbiogenesis in evolution.
deBary, H. A. Die Erscheinung der Symbiose (Strasburg, 1879).
Mayo, M. A., Taliansky, M. E. & Fritsch, C. in Satellites and Defective RNAs (eds Vogt, P. K. & Jackson, O.) 65–79 (Springer, 1999).
Sapp, J. Evolution by Association, A History of Symbiosis (Oxford University Press, New York, 1994).
Drake, J. W., Charlesworth, B., Charlesworth, D. & Crow, J. F. Rates of spontaneous mutation. Genetics 148, 1667–1686 (1998).
Turnbull, M. & Webb, B. Perspectives on polydnavirus origins and evolution. Adv. Virus Res. 58, 203–254 (2002).
Margulis, L. Symbiosis in Cell Evolution (W. H. Freeman, New York, 1981).
Filée, J., Forterre, P. & Laurent, J. The role played by viruses in the evolution of their hosts: a view based on informational protein phylogenies. Res. Microbiol. 154, 237–243 (2003).
Villarreal, L. P. Viruses and the Evolution of Life (ASM Press, Washington DC, 2005).
Gorinsek, B., Bubensek, F. & Kordis, D. Evolutionary genomics of chromoviruses in eukaryotes. Mol. Biol. Evol. 21, 781–798 (2004).
Kazazian, H. H. Jr. Mobile elements: drivers of genome evolution. Science 303, 1626–1632 (2004).
Peterson-Burch, B. D., Wright, D. A., Laten, H. M. & Voytas, D. F. Retroviruses in plants? Trends Genet. 16, 151–152 (2000).
Shackelton, L. A. & Holmes, E. C. The evolution of large DNA viruses: combining genomic information of viruses and their hosts. Trends Microbiol. 12, 458–465 (2004).
Simon, A. E. & Bujarski, J. J. RNA–RNA recombination and evolution in virus-infected plants. Annu. Rev. Phytopathol. 32, 337–362 (1994).
Nagy, P. D. & Simon, A. E. New insights into the mechanisms of RNA recombination. Virology 235, 1–9 (1997).
Lai, M. M. C. RNA recombination in animal and plant viruses. Microbiol. Rev. 56, 61–79 (1992).
Roossinck, M. J. Cucumber mosaic virus, a model for RNA virus evolution. Mol. Plant Pathol. 2, 59–63 (2001).
Roossinck, M. J. Plant RNA virus evolution. Curr. Opin. Microbiol. 6, 406–409 (2003).
McKinney, H. H. Mosaic diseases in the Canary Islands, west of Africa and Gibraltar. J. Agric. Res. 39, 557–578 (1929).
Kunkel, L. O. Studies on acquired immunity with tobacco and acuba mosaics. Phytopathology 24, 437–466 (1934).
Fraser, R. S. S. in Plant Virology Protocols (eds Foster, G. D. & Taylor, S. C.) 13–24 (Humana Press, Totowa, 1998).
Bawden, F. C. Plant Viruses and Virus Diseases (The Ronald Press Company, New York, 1964).
Dietrich, C. & Maiss, E. Fluorescent labelling reveals spatial separation of potyvirus populations in mixed infected Nicotiana benthamiana plants. J. Gen. Virol. 84, 2871–2876 (2003).
Hall, J. S., French, R., Hein, G. L., Morris, T. J. & Stenger, D. C. Three distinct mechanisms facilitate genetic isolation of sympatric wheat streak mosaic virus lineages. Virology 282, 230–236 (2001).
Takeshita, M., Shigemune, N., Kikuhara, K., Furuya, N. & Takanami, Y. Spatial analysis for exclusive interactions between subgroups I and II of Cucumber mosaic virus in cowpea. Virology 328, 45–51 (2004).
Baulcombe, D. C. Mechanisms of pathogen-derived resistance to viruses in transgenic plants. Plant Cell 8, 1833–1844 (1996).
Bendahmane, M. & Beachy, R. N. Control of tobamovirus infections via pathogen-derived resistance. Adv. Virus Res. 53, 360–386 (1999).
Novella, I. S. Contributions of vesicular stomatitis virus to the understanding of RNA virus evolution. Curr. Opin. Microbiol. 6, 392–398 (2003).
Fraile, A. et al. A century of tobamovirus evolution in an Australian population of Nicotiana glauca. J. Virol. 71, 8316–8320 (1997).
Roossinck, M. J. & Palukaitis, P. Genetic analysis of helper virus-specific selective amplification of cucumber mosaic virus satellite RNAs. J. Mol. Evol. 40, 25–29 (1995).
Palukaitis, P. & Roossinck, M. J. Spontaneous change of a benign satellite RNA of cucumber mosaic virus to a pathogenic variant. Nature Biotechnol. 14, 1264–1268 (1996).
Rochow, W. F. & Ross, A. F. Virus multiplication in plants doubly infected with potato virus X and Y. Virology 1, 10–27 (1955).
Pruss, G., Ge, X., Shi, X. M., Carrington, J. C. & Vance, V. B. Plant viral synergism: the potyviral genome encodes a broad-range pathogenicity enhancer that transactivates replication of heterologous viruses. Plant Cell 9, 859–868 (1997).
Messenger, S. L., Molineux, I. J. & Bull, J. J. Virulence evolution in a virus obeys a trade-off. Proc. Biol. Sci. 266, 397–404 (1999).
Hull, R. Matthews' Plant Virology (Academic Press, San Diego, 2002).
Scheets, K. Maize chlorotic mottle machlomovirus and wheat streak mosaic rymovirus concentrations increase in the synergistic disease corn lethal necrosis. Virology 242, 28–38 (1998).
Cabrera, O. & Scholthof, K.-B. G. The complex viral etiology of St. Augustine decline. Plant Dis. 83, 902–904 (1999).
Simon, A. E., Roossinck, M. J. & Havelda, Z. Plant virus satellite and defective interfering RNAs: new paradigms for a new century. Annu. Rev. Phytopathol. 42, 415–437 (2004).
Xu, P. & Roossinck, M. J. in Encyclopedia of Life Sciences 1–6 (Macmillan Reference Ltd, London, 2001).
Roossinck, M. J., Sleat, D. & Palukaitis, P. Satellite RNAs of plant viruses: structures and biological effects. Microbiol. Rev. 56, 265–279 (1992).
Gibbs, M. in Molecular Basis of Virus Evolution (eds Gibbs, A. J., Calisher, C. H. & García-Arenal, F.) 351–368 (Cambridge University Press, Cambridge, 1995).
Murant, A. F. Dependence of groundnut rosette virus on its satellite RNA as well as on groundnut rosette assistor luteovirus for transmission by Aphis craccivora. J. Gen. Virol. 71, 2163–2166 (1990).
Robinson, D. J., Ryabov, E. V., Raj, S. K., Roberts, I. M. & Taliansky, M. E. Satellite RNA is essential for encapsidation of groundnut rosette umbravirus RNA by groundnut rosette assistor luteovirus coat protein. Virology 254, 105–114 (1999). This paper describes a fascinating and complex example of viral symbiotic mutualism, involving three separate entities.
Ryabov, E. V., Fraser, G., Mayo, M. A., Barker, H. & Talinasky, M. Umbravirus gene expression helps Potato leafroll virus to invade mesophyll tissues and to be transmitted mechanically between plants. Virology 286, 363–372 (2001).
Marchoux, G., Delecolle, B. & Selassie, K. G. Systemic infection to tobacco by pepper veinal mottle potyvirus (PVMV) depends on the presence of potato virus Y (PVY). J. Phytopathol. 137, 283–292 (1993).
Hacker, D. L. & Fowler, B. C. Complementation of the host range restriction of southern cowpea mosaic virus in bean by southern bean mosaic virus. Virology 266, 140–149 (2000).
Guerini, M. N. & Murphy, J. F. Resistance of Capsicum annuum 'Avelar' to pepper mottle potyvirus and alleviation of this resistance by co-infection with cucumber mosaic cucumovirus are associated with virus movement. J. Gen. Virol. 80, 2785–2792 (1999).
Gorbalenya, A. E. in Molecular Basis of Virus Evolution (eds Gibbs, A. J., Calisher, C. H. & García-Arenal, F.) 49–65 (Cambridge University Press, Cambridge, 1995).
White, P. S., Morales, F. J. & Roossinck, M. J. Interspecific reassortment in the evolution of a cucumovirus. Virology 207, 334–337 (1995).
Roossinck, M. J. Evolutionary history of Cucumber mosaic virus deduced by phylogenetic analyses. J. Virol. 76, 3382–3387 (2002). This paper shows incongruent trees for individual RNAs of this virus species, and interprets them to mean that genomic reassortment was important in the evolutionary history of the virus.
Morozov, S. Y., Dolja, V. V. & Atabekov, J. G. Probable reassortment of genomic elements among elongated RNA-containing plant viruses. J. Mol. Evol. 29, 52–62 (1989).
Botstein, D. A theory of modular evolution for bacteriophages. Ann. N. Y. Acad. Sci. 354, 484–490 (1980).
Nee, S. Mutualism, parasitism and competition in the evolution of coviruses. Phil. Trans. R. Soc. Lond. B Biol. Sci. 355, 1607–1613 (2000).
Robinson, D. J., Hamilton, W. D. O., Harrison, B. D. & Baulcombe, D. C. Two anomalous tobravirus isolates: evidence for RNA recombination in nature. J. Gen. Virol. 68, 2551–2561 (1987).
MacFarlane, S. A. Natural recombination among plant virus genomes: evidence from tobraviruses. Semin. Virol. 8, 25–31 (1997).
Masuta, C., Ueda, S., Suzuki, M. & Uyeda, I. Evolution of a quadripartite hybrid virus by interspecific exchange and recombination between replicase components of two related tripartite RNA viruses. Proc. Natl Acad. Sci. USA 95, 104787–10492 (1998). This paper shows symbiogenesis in action (although the term is not used in the paper).
Demler, S. A., Rucher, D. G. & deZoeten, G. A. The chimeric nature of the genome of pea enation mosaic virus: the independent replication of RNA 2. J. Gen. Virol. 74, 1–14 (1993).
Moonan, F., Molina, J. & Mirkov, T. E. Sugarcane yellow leaf virus: an emerging virus that has evolved by recombination between luteoviral and poleroviral ancestors. Virology 269, 156–171 (2000). This paper provides another excellent example of the results of symbiogenesis in the evolution of this virus.
Smith, G. R., Borg, Z., Lockhart, B. E. L., Braithwaite, K. S. & Gibbs, M. J. Sugarcane yellow leaf virus: a novel member of the Luteoviridae that probably arose by inter-species recombination. J. Gen. Virol. 81, 1865–1869 (2000).
Aaziz, R. & Tepfer, M. Recombination between genomic RNAs of two cucumoviruses under conditions of minimal selection pressure. Virology 263, 282–289 (1999).
Villarreal, L. P. & DeFilippis, V. R. A hypothesis for DNA viruses as the origin of eukaryotic replication proteins. J. Virol. 74, 7079–7084 (2000).
Takemura, M. Poxviruses and the origin of the eukaryotic nucleus. J. Mol. Evol. 52, 419–425 (2001).
Bell, P. J. L. Viral eukaryogenesis: was the ancestor of the nucleus a complex DNA virus? J. Mol. Evol. 53, 251–256 (2001).
Acknowledgements
The author wishes to thank E. Blancaflor, D. Brooks, A. Elmer, L. Marquez, G. May, J. Pita, M. Taylor and P. Xu for thoughtful discussions and editorial comments, B. Falk for unpublished data and L. Villarreal and F. Ryan for inspiration for this line of thinking. The author is also grateful to the S.R. Noble Foundation for financial support.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Related links
Glossary
- SALTATION
-
A rapid evolutionary event that changes a species into another species.
- SYMBIOGENESIS
-
The fusion of symbiotic organisms to form a new species. According to Ryan's definition, this can result from endosymbiosis, which involves genes and/or genomes, and exosymbiosis, in which the interaction involves behaviour or shared metabolites.
- POSITIVE-SENSE RNA VIRUS
-
A virus with a single-stranded RNA genome that can serve as a messenger RNA without further transcription.
- ISOGENIC
-
Genetically identical; derived from the same inbred line.
- SUPERINFECTION
-
Infection of a cell by a virus when it is already infected by another virus.
- SATELLITE RNA
-
A parasitic RNA associated with some plant RNA viruses. Satellite RNAs are completely dependent on their helper virus for replication and dissemination, but do not provide any essential function to the helper virus.
- PARARETROVIRUS
-
A virus that packages an RNA pre-genome that is converted to DNA in the virion. Retroviruses and pararetroviruses behave like RNA viruses in their evolutionary capacity because their replication machinery is highly error prone.
- CO-EVOLUTION
-
The evolution of two symbiotic species in which each change by one partner is compensated by a change in the other partner.
- CONGRUENT TREES
-
A set of phylogenetic trees using different genes that show the same patterns of evolution for each gene. Incongruent trees show markedly different topologies when different genes from the same set of taxa are analysed.
- CLADISTIC TREE
-
A tree depicting the phylogenetic relationships of a group of organisms. In a cladogram, only the branching order is significant, the branch lengths are not significant.
- SPECIATION
-
The formation of a new species. By classic definition, a new species is sexually incompatible with its parental species, although the definition for viruses, and many other organisms such as plants, is less clear.
Rights and permissions
About this article
Cite this article
Roossinck, M. Symbiosis versus competition in plant virus evolution. Nat Rev Microbiol 3, 917–924 (2005). https://doi.org/10.1038/nrmicro1285
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro1285
This article is cited by
-
Expanding known viral diversity in plants: virome of 161 species alongside an ancient canal
Environmental Microbiome (2022)
-
Interaction between ‘Candidatus Phytoplasma australasiae’ and Tomato yellow leaf curl virus in tomato plants
European Journal of Plant Pathology (2020)
-
Multipartite viruses: adaptive trick or evolutionary treat?
npj Systems Biology and Applications (2017)
-
In silico approach to reveal viral populations in grapevine cultivar Tannat using transcriptome data
Scientific Reports (2015)
-
Large-scale transcriptome comparison reveals distinct gene activations in wheat responding to stripe rust and powdery mildew
BMC Genomics (2014)