Key Points
-
Alphaproteobacteria are Gram-negative bacteria that have heterogeneous lifestyles, including aquatic and soil-dwelling lineages, endosymbionts, pathogens of plants, arthropods or mammals, as well as the ancestors of eukaryotic mitochondria.
-
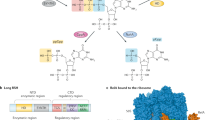
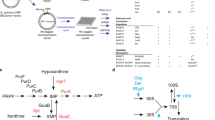
The phosphorylated guanosines cyclic di-GMP (c-di-GMP), and guanosine tetraphosphate and guanosine pentaphosphate (collectively referred to as (p)ppGpp) act as antagonistic second messengers that control the progression of the cell cycle in Alphaproteobacteria.
-
The intracellular concentration of c-di-GMP surges at the G1–S transition of the cell cycle in Caulobacter crescentus to stimulate the initiation of DNA replication, and c-di-GMP coordinates remodelling of the differentiating pole.
-
c-di-GMP regulates the proteolytic degradation of key regulators of the cell cycle.
-
The alarmone (p)ppGpp promotes the swarmer phase of C. crescentus cells by delaying the initiation of DNA replication and polar differentiation.
-
The alarmone is crucial for several Rhizobiales to sustain symbiotic or pathogenic relationships with their respective eukaryotic host cells.
Abstract
The class Alphaproteobacteria includes Gram-negative free-living, symbiotic and obligate intracellular bacteria, as well as important plant, animal and human pathogens. Recent work has established the key antagonistic roles that phosphorylated guanosines, cyclic-di-GMP (c-di-GMP) and the alarmones guanosine tetraphosphate and guanosine pentaphosphate (collectively referred to as (p)ppGpp), have in the regulation of the cell cycle in these bacteria. In this Review, we discuss the insights that have been gained into the regulation of the initiation of DNA replication and cytokinesis by these second messengers, with a particular focus on the cell cycle of Caulobacter crescentus. We explore how the fluctuating levels of c-di-GMP and (p)ppGpp during the progression of the cell cycle and under conditions of stress control the synthesis and proteolysis of key regulators of the cell cycle. As these signals also promote bacterial interactions with host cells, the enzymes that control (p)ppGpp and c-di-GMP are attractive antibacterial targets.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Andersson, S. G. et al. The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature 396, 133–140 (1998).
Batut, J., Andersson, S. G. & O'Callaghan, D. The evolution of chronic infection strategies in the α-proteobacteria. Nat. Rev. Microbiol. 2, 933–945 (2004).
Evinger, M. & Agabian, N. Envelope-associated nucleoid from Caulobacter crescentus stalked and swarmer cells. J. Bacteriol. 132, 294–301 (1977).
Skerker, J. M. & Laub, M. T. Cell-cycle progression and the generation of asymmetry in Caulobacter crescentus. Nat. Rev. Microbiol. 2, 325–337 (2004).
Schrader, J. M. & Shapiro, L. Synchronization of Caulobacter crescentus for investigation of the bacterial cell cycle. J. Vis. Exp. http://dx.doi.org/10.3791/52633 (2015).
Laub, M. T., Shapiro, L. & McAdams, H. H. Systems biology of Caulobacter. Annu. Rev. Genet. 41, 429–441 (2007).
Lasker, K. et al. CauloBrowser: a systems biology resource for Caulobacter crescentus. Nucleic Acids Res. 44, D640–D645 (2016).
Schrader, J. M. et al. The coding and noncoding architecture of the Caulobacter crescentus genome. PLoS Genet. 10, e1004463 (2014).
Zhou, B. et al. The global regulatory architecture of transcription during the Caulobacter cell cycle. PLoS Genet. 11, e1004831 (2015).
Kahng, L. S. & Shapiro, L. The CcrM DNA methyltransferase of Agrobacterium tumefaciens is essential, and its activity is cell cycle regulated. J. Bacteriol. 183, 3065–3075 (2001).
De Nisco, N. J., Abo, R. P., Wu, C. M., Penterman, J. & Walker, G. C. Global analysis of cell cycle gene expression of the legume symbiont Sinorhizobium meliloti. Proc. Natl Acad. Sci. USA 111, 3217–3224 (2014).
Hallez, R., Bellefontaine, A. F., Letesson, J. J. & De Bolle, X. Morphological and functional asymmetry in α-proteobacteria. Trends Microbiol. 12, 361–365 (2004).
Hallez, R. et al. The asymmetric distribution of the essential histidine kinase PdhS indicates a differentiation event in Brucella abortus. EMBO J. 26, 1444–1455 (2007).
Lam, H., Matroule, J. Y. & Jacobs-Wagner, C. The asymmetric spatial distribution of bacterial signal transduction proteins coordinates cell cycle events. Dev. Cell 5, 149–159 (2003).
Schaper, S. et al. Cyclic di-GMP regulates multiple cellular functions in the symbiotic α-proteobacterium Sinorhizobium meliloti. J. Bacteriol. 198, 521–535 (2016).
Grangeon, R., Zupan, J. R., Anderson-Furgeson, J. & Zambryski, P. C. PopZ identifies the new pole, and PodJ identifies the old pole during polar growth in Agrobacterium tumefaciens. Proc. Natl Acad. Sci. USA 112, 11666–11671 (2015).
Haglund, C. M., Choe, J. E., Skau, C. T., Kovar, D. R. & Welch, M. D. Rickettsia Sca2 is a bacterial formin-like mediator of actin-based motility. Nat. Cell Biol. 12, 1057–1063 (2010).
Kirkpatrick, C. L. & Viollier, P. H. Poles apart: prokaryotic polar organelles and their spatial regulation. Cold Spring Harb. Perspect. Biol. 3, a006809 (2011).
Kobayashi, H., De Nisco, N. J., Chien, P., Simmons, L. A. & Walker, G. C. Sinorhizobium meliloti CpdR1 is critical for co-ordinating cell cycle progression and the symbiotic chronic infection. Mol. Microbiol. 73, 586–600 (2009).
Laub, M. T., McAdams, H. H., Feldblyum, T., Fraser, C. M. & Shapiro, L. Global analysis of the genetic network controlling a bacterial cell cycle. Science 290, 2144–2148 (2000).
Collier, J. Regulation of chromosomal replication in Caulobacter crescentus. Plasmid 67, 76–87 (2012).
Marczynski, G. T. & Shapiro, L. Control of chromosome replication in Caulobacter crescentus. Annu. Rev. Microbiol. 56, 625–656 (2002).
Kahng, L. S. & Shapiro, L. Polar localization of replicon origins in the multipartite genomes of Agrobacterium tumefaciens and Sinorhizobium meliloti. J. Bacteriol. 185, 3384–3391 (2003).
Frage, B. et al. Spatiotemporal choreography of chromosome and megaplasmids in the Sinorhizobium meliloti cell cycle. Mol. Microbiol. 100, 808–823 (2016).
Katayama, T., Ozaki, S., Keyamura, K. & Fujimitsu, K. Regulation of the replication cycle: conserved and diverse regulatory systems for DnaA and oriC. Nat. Rev. Microbiol. 8, 163–170 (2010).
Domian, I. J., Quon, K. C. & Shapiro, L. Cell type-specific phosphorylation and proteolysis of a transcriptional regulator controls the G1-to-S transition in a bacterial cell cycle. Cell 90, 415–424 (1997).
Jacobs, C., Domian, I. J., Maddock, J. R. & Shapiro, L. Cell cycle-dependent polar localization of an essential bacterial histidine kinase that controls DNA replication and cell division. Cell 97, 111–120 (1999).
Biondi, E. G. et al. Regulation of the bacterial cell cycle by an integrated genetic circuit. Nature 444, 899–904 (2006).
Iniesta, A. A., McGrath, P. T., Reisenauer, A., McAdams, H. H. & Shapiro, L. A phospho-signaling pathway controls the localization and activity of a protease complex critical for bacterial cell cycle progression. Proc. Natl Acad. Sci. USA 103, 10935–10940 (2006).
Skerker, J. M., Prasol, M. S., Perchuk, B. S., Biondi, E. G. & Laub, M. T. Two-component signal transduction pathways regulating growth and cell cycle progression in a bacterium: a system-level analysis. PLoS Biol. 3, e334 (2005).
Lau, J., Hernandez-Alicea, L., Vass, R. H. & Chien, P. A. Phosphosignaling adaptor primes the AAA+ protease ClpXP to drive cell cycle-regulated proteolysis. Mol. Cell 59, 104–116 (2015).
Chen, Y. E., Tsokos, C. G., Biondi, E. G., Perchuk, B. S. & Laub, M. T. Dynamics of two phosphorelays controlling cell cycle progression in Caulobacter crescentus. J. Bacteriol. 191, 7417–7429 (2009).
Pini, F. et al. Cell cycle control by the master regulator CtrA in Sinorhizobium meliloti. PLoS Genet. 11, e1005232 (2015).
Pini, F. et al. The DivJ, CbrA and PleC system controls DivK phosphorylation and symbiosis in Sinorhizobium meliloti. Mol. Microbiol. 90, 54–71 (2013).
Bellefontaine, A. F. et al. Plasticity of a transcriptional regulation network among α-proteobacteria is supported by the identification of CtrA targets in Brucella abortus. Mol. Microbiol. 43, 945–960 (2002).
Cheng, Z., Miura, K., Popov, V. L., Kumagai, Y. & Rikihisa, Y. Insights into the CtrA regulon in development of stress resistance in obligatory intracellular pathogen Ehrlichia chaffeensis. Mol. Microbiol. 82, 1217–1234 (2011).
Schallies, K. B., Sadowski, C., Meng, J., Chien, P. & Gibson, K. E. Sinorhizobium meliloti CtrA stability is regulated in a CbrA-dependent manner that is influenced by CpdR1. J. Bacteriol. 197, 2139–2149 (2015).
Van der Henst, C. et al. The histidine kinase PdhS controls cell cycle progression of the pathogenic alphaproteobacterium Brucella abortus. J. Bacteriol. 194, 5305–5314 (2012).
Barnett, M. J., Hung, D. Y., Reisenauer, A., Shapiro, L. & Long, S. R. A homolog of the CtrA cell cycle regulator is present and essential in Sinorhizobium meliloti. J. Bacteriol. 183, 3204–3210 (2001).
Kim, J., Heindl, J. E. & Fuqua, C. Coordination of division and development influences complex multicellular behavior in Agrobacterium tumefaciens. PLoS ONE 8, e56682 (2013).
Sadowski, C. S., Wilson, D., Schallies, K. B., Walker, G. & Gibson, K. E. The Sinorhizobium meliloti sensor histidine kinase CbrA contributes to free-living cell cycle regulation. Microbiology 159, 1552–1563 (2013).
Cheng, Z., Lin, M. & Rikihisa, Y. Ehrlichia chaffeensis proliferation begins with NtrY/NtrX and PutA/GlnA upregulation and CtrA degradation induced by proline and glutamine uptake. mBio 5, e02141 (2014).
Paul, R. et al. Allosteric regulation of histidine kinases by their cognate response regulator determines cell fate. Cell 133, 452–461 (2008).
Abel, S. et al. Bi-modal distribution of the second messenger c-di-GMP controls cell fate and asymmetry during the Caulobacter cell cycle. PLoS Genet. 9, e1003744 (2013). This paper highlights the severe developmental and cell cycle defects of a c-di-GMP-deficient C. crescentus strain and shows that c-di-GMP concentration fluctuates during the course of the cell cycle in a synchronized population.
Christen, M. et al. Asymmetrical distribution of the second messenger c-di-GMP upon bacterial cell division. Science 328, 1295–1297 (2010). The first report to show the oscillations of c-di-GMP concentration during the cell cycle at the single-cell level using an in vivo c-di-GMP–FRET reporter system.
Paul, R. et al. Cell cycle-dependent dynamic localization of a bacterial response regulator with a novel di-guanylate cyclase output domain. Genes Dev. 18, 715–727 (2004). This study provides the first demonstration that GGDEF domains have diguanylate cyclase (c-di-GMP synthetase) activity.
Kumagai, Y., Matsuo, J., Hayakawa, Y. & Rikihisa, Y. Cyclic di-GMP signaling regulates invasion by Ehrlichia chaffeensis of human monocytes. J. Bacteriol. 192, 4122–4133 (2010).
Abel, S. et al. Regulatory cohesion of cell cycle and cell differentiation through interlinked phosphorylation and second messenger networks. Mol. Cell 43, 550–560 (2011). This study shows that the cell cycle proteolytic degradation of the phosphodiesterase PdeA enables c-di-GMP to surge at the G1–S transition and remodel the differentiating pole.
Joshi, K. K., Berge, M., Radhakrishnan, S. K., Viollier, P. H. & Chien, P. An adaptor hierarchy regulates proteolysis during a bacterial cell cycle. Cell 163, 419–431 (2015). This study shows that specialized adaptor hierarchy controls substrate degradation by a primed protease during the C. crescentus cell cycle.
Fumeaux, C. et al. Cell cycle transition from S-phase to G1 in Caulobacter is mediated by ancestral virulence regulators. Nat. Commun. 5, 4081 (2014).
Fiebig, A. et al. A cell cycle and nutritional checkpoint controlling bacterial surface adhesion. PLoS Genet. 10, e1004101 (2014).
Ardissone, S. et al. Cell cycle constraints on capsulation and bacteriophage susceptibility. eLife 3, e03587 (2014).
Mann, T. H., Seth Childers, W., Blair, J. A., Eckart, M. R. & Shapiro, L. A cell cycle kinase with tandem sensory PAS domains integrates cell fate cues. Nat. Commun. 7, 11454 (2016).
Lori, C. et al. Cyclic di-GMP acts as a cell cycle oscillator to drive chromosome replication. Nature 523, 236–239 (2015). This study reports that the binding of c-di-GMP to the cell cycle kinase CckA inhibits its kinase activity while stimulating its phosphatase activity, which leads to the dephosphorylation of CtrA and ultimately triggers the initiation of DNA replication.
Quon, K. C., Marczynski, G. T. & Shapiro, L. Cell cycle control by an essential bacterial two-component signal transduction protein. Cell 84, 83–93 (1996).
Janakiraman, B., Mignolet, J., Narayanan, S., Viollier, P. H. & Radhakrishnan, S. K. In-phase oscillation of global regulons is orchestrated by a pole-specific organizer. Proc. Natl Acad. Sci. USA 113, 12550–12555 (2016).
Radhakrishnan, S. K., Thanbichler, M. & Viollier, P. H. The dynamic interplay between a cell fate determinant and a lysozyme homolog drives the asymmetric division cycle of Caulobacter crescentus. Genes Dev. 22, 212–225 (2008).
Dubey, B. N. et al. Cyclic di-GMP mediates a histidine kinase/phosphatase switch by noncovalent domain cross-linking. Sci. Adv. 2, e1600823 (2016). A crystallographic analysis that shows that c-di-GMP binds to the catalytic domain of CckA.
Rikihisa, Y. Molecular pathogenesis of Ehrlichia chaffeensis infection. Annu. Rev. Microbiol. 69, 283–304 (2015).
Brilli, M. et al. The diversity and evolution of cell cycle regulation in α-proteobacteria: a comparative genomic analysis. BMC Syst. Biol. 4, 52 (2010).
Soding, J., Biegert, A. & Lupas, A. N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 33, W244–W248 (2005).
Duerig, A. et al. Second messenger-mediated spatiotemporal control of protein degradation regulates bacterial cell cycle progression. Genes Dev. 23, 93–104 (2009). The first report to show that a degenerated GGDEF domain can evolve into a c-di-GMP receptor.
Smith, S. C. et al. Cell cycle-dependent adaptor complex for ClpXP-mediated proteolysis directly integrates phosphorylation and second messenger signals. Proc. Natl Acad. Sci. USA 111, 14229–14234 (2014).
McGrath, P. T., Iniesta, A. A., Ryan, K. R., Shapiro, L. & McAdams, H. H. A dynamically localized protease complex and a polar specificity factor control a cell cycle master regulator. Cell 124, 535–547 (2006).
Ozaki, S. et al. Activation and polar sequestration of PopA, a c-di-GMP effector protein involved in Caulobacter crescentus cell cycle control. Mol. Microbiol. 94, 580–594 (2014).
Bowman, G. R. et al. A polymeric protein anchors the chromosomal origin/ParB complex at a bacterial cell pole. Cell 134, 945–955 (2008).
Ebersbach, G., Briegel, A., Jensen, G. J. & Jacobs-Wagner, C. A self-associating protein critical for chromosome attachment, division, and polar organization in Caulobacter. Cell 134, 956–968 (2008).
Wu, J., Ohta, N. & Newton, A. An essential, multicomponent signal transduction pathway required for cell cycle regulation in Caulobacter. Proc. Natl Acad. Sci. USA 95, 1443–1448 (1998).
Radhakrishnan, S. K., Pritchard, S. & Viollier, P. H. Coupling prokaryotic cell fate and division control with a bifunctional and oscillating oxidoreductase homolog. Dev. Cell 18, 90–101 (2010).
Beaufay, F. et al. A NAD-dependent glutamate dehydrogenase coordinates metabolism with cell division in Caulobacter crescentus. EMBO J. 34, 1786–1800 (2015).
Viollier, P. H., Sternheim, N. & Shapiro, L. Identification of a localization factor for the polar positioning of bacterial structural and regulatory proteins. Proc. Natl Acad. Sci. USA 99, 13831–13836 (2002).
Hinz, A. J., Larson, D. E., Smith, C. S. & Brun, Y. V. The Caulobacter crescentus polar organelle development protein PodJ is differentially localized and is required for polar targeting of the PleC development regulator. Mol. Microbiol. 47, 929–941 (2003).
Marques, M. V., Gomes, S. L. & Gober, J. W. A gene coding for a putative sigma 54 activator is developmentally regulated in Caulobacter crescentus. J. Bacteriol. 179, 5502–5510 (1997).
Biondi, E. G. et al. A phosphorelay system controls stalk biogenesis during cell cycle progression in Caulobacter crescentus. Mol. Microbiol. 59, 386–401 (2006).
Bhat, N. H., Vass, R. H., Stoddard, P. R., Shin, D. K. & Chien, P. Identification of ClpP substrates in Caulobacter crescentus reveals a role for regulated proteolysis in bacterial development. Mol. Microbiol. 88, 1083–1092 (2013).
Vass, R. H., Zeinert, R. D. & Chien, P. Protease regulation and capacity during Caulobacter growth. Curr. Opin. Microbiol. 34, 75–81 (2016).
Aldridge, P. & Jenal, U. Cell cycle-dependent degradation of a flagellar motor component requires a novel-type response regulator. Mol. Microbiol. 32, 379–391 (1999).
Grunenfelder, B. et al. Identification of the protease and the turnover signal responsible for cell cycle-dependent degradation of the Caulobacter FliF motor protein. J. Bacteriol. 186, 4960–4971 (2004).
Huitema, E., Pritchard, S., Matteson, D., Radhakrishnan, S. K. & Viollier, P. H. Bacterial birth scar proteins mark future flagellum assembly site. Cell 124, 1025–1037 (2006).
Davis, N. J. et al. De- and repolarization mechanism of flagellar morphogenesis during a bacterial cell cycle. Genes Dev. 27, 2049–2062 (2013). This study shows that C. crescentus uses TipF, a protein that contains a degenerated EAL domain that has lost its ability to function as a phosphodiesterase, as a c-di-GMP receptor and that the binding of c-di-GMP to TipF helps to determine where and when a new flagellum is assembled.
Christen, M. et al. DgrA is a member of a new family of cyclic diguanosine monophosphate receptors and controls flagellar motor function in Caulobacter crescentus. Proc. Natl Acad. Sci. USA 104, 4112–4117 (2007).
Gonzalez, D. & Collier, J. Effects of (p)ppGpp on the progression of the cell cycle of Caulobacter crescentus. J. Bacteriol. 196, 2514–2525 (2014).
Sanselicio, S. & Viollier, P. H. Convergence of alarmone and cell cycle signaling from trans-encoded sensory domains. mBio 6, e01415-15 (2015).
Ronneau, S., Petit, K., De Bolle, X. & Hallez, R. Phosphotransferase-dependent accumulation of (p)ppGpp in response to glutamine deprivation in Caulobacter crescentus. Nat. Commun. 7, 11423 (2016). This paper shows that C. crescentus uses the phosphotransferase system PTSNtr to evaluate the intracellular concentration of glutamine as a proxy for nitrogen availability and to ultimately modulate the activities of the bifunctional (p)ppGpp synthetase and hydrolase SpoT.
Vercruysse, M. et al. Stress response regulators identified through genome-wide transcriptome analysis of the (p)ppGpp-dependent response in Rhizobium etli. Genome Biol. 12, R17 (2011).
Potrykus, K. & Cashel, M. (p)ppGpp: still magical? Annu. Rev. Microbiol. 62, 35–51 (2008).
Boutte, C. C., Henry, J. T. & Crosson, S. ppGpp and polyphosphate modulate cell cycle progression in Caulobacter crescentus. J. Bacteriol. 194, 28–35 (2012).
Wang, J. D., Sanders, G. M. & Grossman, A. D. Nutritional control of elongation of DNA replication by (p)ppGpp. Cell 128, 865–875 (2007).
Chiaverotti, T. A., Parker, G., Gallant, J. & Agabian, N. Conditions that trigger guanosine tetraphosphate accumulation in Caulobacter crescentus. J. Bacteriol. 145, 1463–1465 (1981).
Lesley, J. A. & Shapiro, L. SpoT regulates DnaA stability and initiation of DNA replication in carbon-starved Caulobacter crescentus. J. Bacteriol. 190, 6867–6880 (2008).
Boutte, C. C. & Crosson, S. The complex logic of stringent response regulation in Caulobacter crescentus: starvation signalling in an oligotrophic environment. Mol. Microbiol. 80, 695–714 (2011).
Atkinson, G. C., Tenson, T. & Hauryliuk, V. The RelA/SpoT homolog (RSH) superfamily: distribution and functional evolution of ppGpp synthetases and hydrolases across the tree of life. PLoS ONE 6, e23479 (2011).
Stott, K. V. et al. (p)ppGpp modulates cell size and the initiation of DNA replication in Caulobacter crescentus in response to a block in lipid biosynthesis. Microbiology 161, 553–564 (2015).
Gorbatyuk, B. & Marczynski, G. T. Regulated degradation of chromosome replication proteins DnaA and CtrA in Caulobacter crescentus. Mol. Microbiol. 55, 1233–1245 (2005).
Leslie, D. J. et al. Nutritional control of DNA replication initiation through the proteolysis and regulated translation of DnaA. PLoS Genet. 11, e1005342 (2015).
Wargachuk, R. & Marczynski, G. T. The Caulobacter crescentus homolog of DnaA (HdaA) also regulates the proteolysis of the replication initiator protein DnaA. J. Bacteriol. 197, 3521–3532 (2015).
Jonas, K., Liu, J., Chien, P. & Laub, M. T. Proteotoxic stress induces a cell-cycle arrest by stimulating Lon to degrade the replication initiator DnaA. Cell 154, 623–636 (2013).
Liu, J., Francis, L. I., Jonas, K., Laub, M. T. & Chien, P. ClpAP is an auxiliary protease for DnaA degradation in Caulobacter crescentus. Mol. Microbiol. 102, 1075–1085 (2016).
Hauryliuk, V., Atkinson, G. C., Murakami, K. S., Tenson, T. & Gerdes, K. Recent functional insights into the role of (p)ppGpp in bacterial physiology. Nat. Rev. Microbiol. 13, 298–309 (2015).
Sanselicio, S., Berge, M., Theraulaz, L., Radhakrishnan, S. K. & Viollier, P. H. Topological control of the Caulobacter cell cycle circuitry by a polarized single-domain PAS protein. Nat. Commun. 6, 7005 (2015). This study describes the first link between the CtrA pathway and the phosphotransferase system PTSNtr through a polarized PAS domain protein.
Holtzendorff, J. et al. Oscillating global regulators control the genetic circuit driving a bacterial cell cycle. Science 304, 983–987 (2004).
Fioravanti, A. et al. DNA binding of the cell cycle transcriptional regulator GcrA depends on N6-adenosine methylation in Caulobacter crescentus and other Alphaproteobacteria. PLoS Genet. 9, e1003541 (2013).
Gonzalez, D. & Collier, J. Genomic adaptations to the loss of a conserved bacterial DNA methyltransferase. mBio 6, e00952 (2015).
Vinella, D., Joseleau-Petit, D., Thevenet, D., Bouloc, P. & D'Ari, R. Penicillin-binding protein 2 inactivation in Escherichia coli results in cell division inhibition, which is relieved by FtsZ overexpression. J. Bacteriol. 175, 6704–6710 (1993).
Haakonsen, D. L., Yuan, A. H. & Laub, M. T. The bacterial cell cycle regulator GcrA is a σ70 cofactor that drives gene expression from a subset of methylated promoters. Genes Dev. 29, 2272–2286 (2015).
Murray, S. M., Panis, G., Fumeaux, C., Viollier, P. H. & Howard, M. Computational and genetic reduction of a cell cycle to its simplest, primordial components. PLoS Biol. 11, e1001749 (2013).
da Silva, C. A., Balhesteros, H., Mazzon, R. R. & Marques, M. V. SpdR, a response regulator required for stationary-phase induction of Caulobacter Crescentus cspD. J. Bacteriol. 192, 5991–6000 (2010).
da Silva, C. A., Lourenco, R. F., Mazzon, R. R., Ribeiro, R. A. & Marques, M. V. Transcriptomic analysis of the stationary phase response regulator SpdR in Caulobacter crescentus. BMC Microbiol. 16, 66 (2016).
Dalebroux, Z. D., Svensson, S. L., Gaynor, E. C. & Swanson, M. S. ppGpp conjures bacterial virulence. Microbiol. Mol. Biol. Rev. 74, 171–199 (2010).
Dalebroux, Z. D. & Swanson, M. S. ppGpp: magic beyond RNA polymerase. Nat. Rev. Microbiol. 10, 203–212 (2012).
Gaca, A. O., Colomer-Winter, C. & Lemos, J. A. Many means to a common end: the intricacies of (p)ppGpp metabolism and its control of bacterial homeostasis. J. Bacteriol. 197, 1146–1156 (2015).
Krol, E. & Becker, A. ppGpp in Sinorhizobium meliloti: biosynthesis in response to sudden nutritional downshifts and modulation of the transcriptome. Mol. Microbiol. 81, 1233–1254 (2011).
Hanna, N. et al. Global Rsh-dependent transcription profile of Brucella suis during stringent response unravels adaptation to nutrient starvation and cross-talk with other stress responses. BMC Genom. 14, 459 (2013).
Masuda, S. & Bauer, C. E. Null mutation of HvrA compensates for loss of an essential relA/spoT-like gene in Rhodobacter capsulatus. J. Bacteriol. 186, 235–239 (2004).
Wells, D. H. & Long, S. R. The Sinorhizobium meliloti stringent response affects multiple aspects of symbiosis. Mol. Microbiol. 43, 1115–1127 (2002).
Moris, M. et al. Effective symbiosis between Rhizobium etli and Phaseolus vulgaris requires the alarmone ppGpp. J. Bacteriol. 187, 5460–5469 (2005).
Wells, D. H. & Long, S. R. Mutations in rpoBC suppress the defects of a Sinorhizobium meliloti relA mutant. J. Bacteriol. 185, 5602–5610 (2003).
Kim, S., Watanabe, K., Suzuki, H. & Watarai, M. Roles of Brucella abortus SpoT in morphological differentiation and intramacrophagic replication. Microbiology 151, 1607–1617 (2005).
Dozot, M. et al. The stringent response mediator Rsh is required for Brucella melitensis and Brucella suis virulence, and for expression of the type IV secretion system virB. Cell. Microbiol. 8, 1791–1802 (2006).
O'Callaghan, D. et al. A homologue of the Agrobacterium tumefaciens VirB and Bordetella pertussis Ptl type IV secretion systems is essential for intracellular survival of Brucella suis. Mol. Microbiol. 33, 1210–1220 (1999).
Comerci, D. J., Martinez-Lorenzo, M. J., Sieira, R., Gorvel, J. P. & Ugalde, R. A. Essential role of the VirB machinery in the maturation of the Brucella abortus-containing vacuole. Cell. Microbiol. 3, 159–168 (2001).
Deghelt, M. et al. G1-arrested newborn cells are the predominant infectious form of the pathogen Brucella abortus. Nat. Commun. 5, 4366 (2014). This paper reports that the infectious populationof B. abortus is predominantly constituted of newborn cells that reside in G1 phase.
Ross, W. et al. ppGpp binding to a site at the RNAP–DksA interface accounts for its dramatic effects on transcription initiation during the stringent response. Mol. Cell 62, 811–823 (2016). This study shows that RNAP has a second (p)ppGpp-binding site at the interface between the β′-subunitand the RNAP-associated transcription factor DksA.
Ross, W., Vrentas, C. E., Sanchez-Vazquez, P., Gaal, T. & Gourse, R. L. The magic spot: a ppGpp binding site on E. coli RNA polymerase responsible for regulation of transcription initiation. Mol. Cell 50, 420–429 (2013).
Adams, D. W. & Errington, J. Bacterial cell division: assembly, maintenance and disassembly of the Z ring. Nat. Rev. Microbiol. 7, 642–653 (2009).
Kuroda, A. & Ohtake, H. Molecular analysis of polyphosphate accumulation in bacteria. Biochemistry (Mosc) 65, 304–308 (2000).
Wexselblatt, E. et al. Relacin, a novel antibacterial agent targeting the stringent response. PLoS Pathog. 8, e1002925 (2012).
An, S. Q. & Ryan, R. P. Combating chronic bacterial infections by manipulating cyclic nucleotide-regulated biofilm formation. Future Med. Chem. 8, 949–961 (2016).
Deutscher, J. et al. The bacterial phosphoenolpyruvate:carbohydrate phosphotransferase system: regulation by protein phosphorylation and phosphorylation-dependent protein–protein interactions. Microbiol. Mol. Biol. Rev. 78, 231–256 (2014).
Goodwin, R. A. & Gage, D. J. Biochemical characterization of a nitrogen-type phosphotransferase system reveals that enzyme EINtr integrates carbon and nitrogen signaling in Sinorhizobium meliloti. J. Bacteriol. 196, 1901–1907 (2014).
Karstens, K., Zschiedrich, C. P., Bowien, B., Stulke, J. & Gorke, B. Phosphotransferase protein EIIANtr interacts with SpoT, a key enzyme of the stringent response, in Ralstonia eutropha H16. Microbiology 160, 711–722 (2014).
Chou, S. H. & Galperin, M. Y. Diversity of cyclic di-GMP-binding proteins and mechanisms. J. Bacteriol. 198, 32–46 (2016).
Schirmer, T. & Jenal, U. Structural and mechanistic determinants of c-di-GMP signalling. Nat. Rev. Microbiol. 7, 724–735 (2009).
Christen, B. et al. Allosteric control of cyclic di-GMP signaling. J. Biol. Chem. 281, 32015–32024 (2006).
Guzzo, C. R., Dunger, G., Salinas, R. K. & Farah, C. S. Structure of the PilZ–FimXEAL–c-di-GMP complex responsible for the regulation of bacterial type IV pilus biogenesis. J. Mol. Biol. 425, 2174–2197 (2013).
Acknowledgements
R.H. is a research associate of the Fond de la Recherche Scientifique-FNRS (F.R.S.-FNRS). P.H.V. is supported by the Swiss National Science Foundation (grants CRSII3_160703 and 31003A_162716) and the Département de l'Instruction Publique du Canton de Genève.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Endosymbionts
-
Organisms that live inside other organisms with which they sustain biological interactions.
- Magic spot
-
The first visualization of (p)ppGpp, which appeared as a spot on a thin-layer chromatogram of radiolabelled nucleotides extracted from Escherichia coli cells that were starved of amino acids.
- Holdfast
-
An adhesive material that is located at the tip of the stalk and is composed of proteins and polysaccharides.
- Stalk
-
An extension of the cell envelope that is constructed at the flagellated pole of Caulobacter crescentus following differentiation, when the flagellum is ejected and the polar pili are retracted.
- Exponential phase
-
A growth phase during which the nutrients are not limiting, which enables the number of microorganisms to increase exponentially and at a constant growth rate.
- Stationary phase
-
A growth phase during which a growth-limiting factor emerges that results in the number of microorganisms remaining roughly constant.
- Fluorescence resonance energy transfer
-
(FRET). A phenomenon by which the energy of a fluorophore (donor) after excitation is transferred to another fluorophore (acceptor), so that the second one will emit fluorescence without being directly excited. This technique is useful for measuring interactions between molecules in vivo.
- Phosphodiesterase
-
An enzyme that breaks phosphodiester bonds in cyclic nucleotides.
- Per–Arnt–Sim domains
-
(PAS domains). Domains that are commonly found at the amino-terminal extremity of signalling proteins and are involved in the binding of a range of small ligands, such as metabolites.
- FtsZ cytokinetic tubulin
-
FtsZ is a tubulin-like GTPase that is involved in cytokinesis in almost all bacteria. Following GTP binding, FtsZ monomers self-assemble to generate a ring-like structure at the division site, where it acts not only as a recruitment platform for cell division proteins but also as an coordinator of cell wall synthesis at the septum.
- Basal body
-
A protein complex that is found at the base of a bacterial flagellum and is composed of several ring-like structures, each of which is associated with, embedded or anchored in one layer of the cell envelope.
- Cyclic GMP phosphodiesterase–adenylyl cyclase–FhlA domain
-
(GAF domain). A domain that is commonly found in enzymes and is involved in the binding of a range of small ligands, such as metabolites.
- Symbiotic relationship
-
Biological interactions between two organisms, in which neither (amensalism), one (commensalism or parasitism) or both organisms (mutualism) benefit from the interactions.
Rights and permissions
About this article
Cite this article
Hallez, R., Delaby, M., Sanselicio, S. et al. Hit the right spots: cell cycle control by phosphorylated guanosines in alphaproteobacteria. Nat Rev Microbiol 15, 137–148 (2017). https://doi.org/10.1038/nrmicro.2016.183
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro.2016.183
This article is cited by
-
The stringent response and physiological roles of (pp)pGpp in bacteria
Nature Reviews Microbiology (2021)
-
Extraction and detection of guanosine 5′-diphosphate-3′-diphosphate in amino acid starvation cells of Clavibacter michiganensis
Brazilian Journal of Microbiology (2021)
-
Cell cycle control and environmental response by second messengers in Caulobacter crescentus
BMC Bioinformatics (2020)
-
Reciprocal growth control by competitive binding of nucleotide second messengers to a metabolic switch in Caulobacter crescentus
Nature Microbiology (2020)
-
Crosstalking second messengers
Nature Microbiology (2020)