Abstract
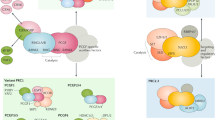
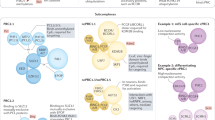
Polycomb group proteins are transcriptional repressors that are essential for normal gene regulation during development. Recent studies suggest that Polycomb repressive complexes (PRCs) recognize and are recruited to their genomic target sites through a range of different mechanisms, which involve transcription factors, CpG island elements and non-coding RNAs. Together with the realization that the interplay between PRC1 and PRC2 is more intricate than was previously appreciated, this has increased our understanding of the vertebrate Polycomb system at the molecular level.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Lewis, E. B. A gene complex controlling segmentation in Drosophila. Nature 276, 565–570 (1978).
Scelfo, A., Piunti, A. & Pasini, D. The controversial role of the Polycomb group proteins in transcription and cancer: how much do we not understand Polycomb proteins? FEBS J. 282, 1703–1722 (2015).
Simon, J. A. & Kingston, R. E. Occupying chromatin: Polycomb mechanisms for getting to genomic targets, stopping transcriptional traffic, and staying put. Mol. Cell 49, 808–824 (2013).
Steffen, P. A. & Ringrose, L. What are memories made of? How Polycomb and Trithorax proteins mediate epigenetic memory. Nat. Rev. Mol. Cell Biol. 15, 340–356 (2014).
Schwartz, Y. B. & Pirrotta, V. A new world of Polycombs: unexpected partnerships and emerging functions. Nat. Rev. Genet. 14, 853–864 (2013).
Di Croce, L. & Helin, K. Transcriptional regulation by Polycomb group proteins. Nat. Struct. Mol. Biol. 20, 1147–1155 (2013).
Grossniklaus, U. & Paro, R. Transcriptional silencing by Polycomb-group proteins. Cold Spring Harb. Perspect. Biol. 6, a019331 (2014).
Gao, Z. et al. PCGF homologs, CBX proteins, and RYBP define functionally distinct PRC1 family complexes. Mol. Cell 45, 344–356 (2012).
Ogawa, H., Ishiguro, K., Gaubatz, S., Livingston, D. M. & Nakatani, Y. A complex with chromatin modifiers that occupies E2F- and Myc-responsive genes in G0 cells. Science 296, 1132–1136 (2002).
Herranz, N. et al. Polycomb complex 2 is required for E-cadherin repression by the Snail1 transcription factor. Mol. Cell. Biol. 28, 4772–4781 (2008).
Dietrich, N. et al. REST-mediated recruitment of Polycomb repressor complexes in mammalian cells. PLoS Genet. 8, e1002494 (2012).
Arnold, P. et al. Modeling of epigenome dynamics identifies transcription factors that mediate Polycomb targeting. Genome Res. 23, 60–73 (2013).
Ren, X. & Kerppola, T. K. REST interacts with Cbx proteins and regulates Polycomb repressive complex 1 occupancy at RE1 elements. Mol. Cell. Biol. 31, 2100–2110 (2011).
Yu, M. et al. Direct recruitment of Polycomb repressive complex 1 to chromatin by core binding transcription factors. Mol. Cell 45, 330–343 (2012).
Maier, V. K. et al. Functional proteomic analysis of repressive histone methyltransferase complexes PRC2 and G9A reveals ZNF518B as a G9A regulator. Mol. Cell. Proteomics 14, 1435–1446 (2015).
Brockdorff, N. Noncoding RNA and Polycomb recruitment. RNA 19, 429–442 (2013).
Kohlmaier, A. et al. A chromosomal memory triggered by Xist regulates histone methylation in X inactivation. PLoS Biol. 2, E171 (2004).
Silva, J. et al. Establishment of histone H3 methylation on the inactive X chromosome requires transient recruitment of Eed–Enx1 Polycomb group complexes. Dev. Cell 4, 481–495 (2003).
Plath, K. et al. Role of histone H3 lysine 27 methylation in X inactivation. Science 300, 131–135 (2003).
Okamoto, I., Otte, A. P., Allis, C. D., Reinberg, D. & Heard, E. Epigenetic dynamics of imprinted X inactivation during early mouse development. Science 303, 644–649 (2004).
da Rocha, S. T. et al. Jarid2 is implicated in the initial Xist-induced targeting of PRC2 to the inactive X chromosome. Mol. Cell 53, 301–316 (2014).
Sarma, K. et al. ATRX directs binding of PRC2 to Xist RNA and Polycomb targets. Cell 159, 869–883 (2014).
Cerase, A. et al. Spatial separation of Xist RNA and Polycomb proteins revealed by superresolution microscopy. Proc. Natl Acad. Sci. USA 111, 2235–2240 (2014).
McHugh, C. A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015).
Pandey, R. R. et al. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol. Cell 32, 232–246 (2008).
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Deaton, A. M. & Bird, A. CpG islands and the regulation of transcription. Genes Dev. 25, 1010–1022 (2011).
Blackledge, N. P. & Klose, R. CpG island chromatin: a platform for gene regulation. Epigenetics 6, 147–152 (2011).
Mendenhall, E. M. et al. GC-rich sequence elements recruit PRC2 in mammalian ES cells. PLoS Genet. 6, e1001244 (2010).
Farcas, A. M. et al. KDM2B links the Polycomb repressive complex 1 (PRC1) to recognition of CpG islands. eLife 1, e00205 (2012).
He, J. et al. Kdm2b maintains murine embryonic stem cell status by recruiting PRC1 complex to CpG islands of developmental genes. Nat. Cell Biol. 15, 373–384 (2013).
Wu, X., Johansen, J. V. & Helin, K. Fbxl10/Kdm2b recruits Polycomb repressive complex 1 to CpG islands and regulates H2A ubiquitylation. Mol. Cell (2013).
Boulard, M., Edwards, J. R. & Bestor, T. H. FBXL10 protects Polycomb-bound genes from hypermethylation. Nat. Genet. 47, 479–485 (2015).
Li, G. et al. Jarid2 and PRC2, partners in regulating gene expression. Genes Dev. 24, 368–380 (2010).
Davidovich, C. et al. Toward a consensus on the binding specificity and promiscuity of PRC2 for RNA. Mol. Cell 57, 552–558 (2015).
Davidovich, C., Zheng, L., Goodrich, K. J. & Cech, T. R. Promiscuous RNA binding by Polycomb repressive complex 2. Nat. Struct. Mol. Biol. 20, 1250–1257 (2013).
Kanhere, A. et al. Short RNAs are transcribed from repressed Polycomb target genes and interact with Polycomb repressive complex-2. Mol. Cell 38, 675–688 (2010).
Kaneko, S., Son, J., Shen, S. S., Reinberg, D. & Bonasio, R. PRC2 binds active promoters and contacts nascent RNAs in embryonic stem cells. Nat. Struct. Mol. Biol. 20, 1258–1264 (2013).
Cifuentes-Rojas, C., Hernandez, A. J., Sarma, K. & Lee, J. T. Regulatory interactions between RNA and Polycomb repressive complex 2. Mol. Cell 55, 171–185 (2014).
Kaneko, S., Son, J., Bonasio, R., Shen, S. S. & Reinberg, D. Nascent RNA interaction keeps PRC2 activity poised and in check. Genes Dev. 28, 1983–1988 (2014).
Schmitges, F. W. et al. Histone methylation by PRC2 is inhibited by active chromatin marks. Mol. Cell 42, 330–341 (2011).
Herzog, V. A. et al. A strand-specific switch in noncoding transcription switches the function of a Polycomb/Trithorax response element. Nat. Genet. 46, 973–981 (2014).
Cai, L. et al. An H3K36 methylation-engaging Tudor motif of Polycomb-like proteins mediates PRC2 complex targeting. Mol. Cell 49, 571–582 (2013).
Qin, S. et al. Tudor domains of the PRC2 components PHF1 and PHF19 selectively bind to histone H3K36me3. Biochem. Biophys. Res. Commun. 430, 547–553 (2013).
Brien, G. L. et al. Polycomb PHF19 binds H3K36me3 and recruits PRC2 and demethylase NO66 to embryonic stem cell genes during differentiation. Nat. Struct. Mol. Biol. 19, 1273–1281 (2012).
Musselman, C. A. et al. Molecular basis for H3K36me3 recognition by the Tudor domain of PHF1. Nat. Struct. Mol. Biol. 19, 1266–1272 (2012).
Mozzetta, C. et al. The histone H3 lysine 9 methyltransferases G9a and GLP regulate Polycomb repressive complex 2-mediated gene silencing. Mol. Cell 53, 277–289 (2014).
Min, J., Zhang, Y. & Xu, R. M. Structural basis for specific binding of Polycomb chromodomain to histone H3 methylated at Lys 27. Genes Dev. 17, 1823–1828 (2003).
Wang, L. et al. Hierarchical recruitment of Polycomb group silencing complexes. Mol. Cell 14, 637–646 (2004).
Boyer, L. A. et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 441, 349–353 (2006).
Tavares, L. et al. RYBP–PRC1 complexes mediate H2A ubiquitylation at Polycomb target sites independently of PRC2 and H3K27me3. Cell 148, 664–678 (2012).
Blackledge, N. P. et al. Variant PRC1 complex-dependent H2A ubiquitylation drives PRC2 recruitment and Polycomb domain formation. Cell 157, 1445–1459 (2014).
Cooper, S. et al. Targeting Polycomb to pericentric heterochromatin in embryonic stem cells reveals a role for H2AK119u1 in PRC2 recruitment. Cell Rep. 7, 1456–1470 (2014).
Kalb, R. et al. Histone H2A monoubiquitination promotes histone H3 methylation in Polycomb repression. Nat. Struct. Mol. Biol. 21, 569–571 (2014).
Bhatnagar, S. et al. TRIM37 is a new histone H2A ubiquitin ligase and breast cancer oncoprotein. Nature 516, 116–120 (2014).
Cao, Q. et al. The central role of EED in the orchestration of Polycomb group complexes. Nat. Commun. 5, 3127 (2014).
Tardat, M. et al. Cbx2 targets PRC1 to constitutive heterochromatin in mouse zygotes in a parent-of-origin-dependent manner. Mol. Cell 58, 157–171 (2015).
Gambetta, M. C. & Müller, J. O-GlcNAcylation prevents aggregation of the Polycomb group repressor Polyhomeotic. Dev. Cell 31, 629–639 (2014).
Isono, K. et al. SAM domain polymerization links subnuclear clustering of PRC1 to gene silencing. Dev. Cell 26, 565–577 (2013).
Kim, C. A., Gingery, M., Pilpa, R. M. & Bowie, J. U. The SAM domain of Polyhomeotic forms a helical polymer. Nat. Struct. Biol. 9, 453–457 (2002).
Eskeland, R. et al. Ring1B compacts chromatin structure and represses gene expression independent of histone ubiquitination. Mol. Cell 38, 452–464 (2010).
Hansen, K. H. et al. A model for transmission of the H3K27me3 epigenetic mark. Nat. Cell Biol. 10, 1291–1300 (2008).
Margueron, R. et al. Role of the Polycomb protein EED in the propagation of repressive histone marks. Nature 461, 762–767 (2009).
Hosogane, M., Funayama, R., Nishida, Y., Nagashima, T. & Nakayama, K. Ras-induced changes in H3K27me3 occur after those in transcriptional activity. PLoS Genet. 9, e1003698 (2013).
Riising, E. M. et al. Gene silencing triggers Polycomb repressive complex 2 recruitment to CpG islands genome wide. Mol. Cell 55, 347–360 (2014).
Klose, R. J., Cooper, S., Farcas, A. M., Blackledge, N. P. & Brockdorff, N. Chromatin sampling — an emerging perspective on targeting Polycomb repressor proteins. PLoS Genet. 9, e1003717 (2013).
Ringrose, L. & Paro, R. Epigenetic regulation of cellular memory by the Polycomb and Trithorax group proteins. Annu. Rev. Genet. 38, 413–443 (2004).
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
Azuara, V. et al. Chromatin signatures of pluripotent cell lines. Nat. Cell Biol. 8, 532–538 (2006).
Voigt, P., Tee, W. W. & Reinberg, D. A double take on bivalent promoters. Genes Dev. 27, 1318–1338 (2013).
Gao, Z. et al. An AUTS2–Polycomb complex activates gene expression in the CNS. Nature 516, 349–354 (2014).
Xu, J. et al. Developmental control of Polycomb subunit composition by GATA factors mediates a switch to non-canonical functions. Mol. Cell 57, 304–316 (2015).
Ferrari, K. J. et al. Polycomb-dependent H3K27me1 and H3K27me2 regulate active transcription and enhancer fidelity. Mol. Cell 53, 49–62 (2014).
Ciferri, C. et al. Molecular architecture of human Polycomb repressive complex 2. eLife 1, e00005 (2012).
McGinty, R. K., Henrici, R. C. & Tan, S. Crystal structure of the PRC1 ubiquitylation module bound to the nucleosome. Nature 514, 591–596 (2014).
Acknowledgements
Work in the Klose laboratory is supported by the Wellcome Trust, the Lister Institute of Preventive Medicine and Exeter College, University of Oxford, UK. N.R.R. is supported by a Junior Research Fellowship at St John's College, University of Oxford. The authors would like to thank Dr Emilia Dimitrova and Dr Sarah Cooper for constructive comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Blackledge, N., Rose, N. & Klose, R. Targeting Polycomb systems to regulate gene expression: modifications to a complex story. Nat Rev Mol Cell Biol 16, 643–649 (2015). https://doi.org/10.1038/nrm4067
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm4067
This article is cited by
-
Functional dissection of PRC1 subunits RYBP and YAF2 during neural differentiation of embryonic stem cells
Nature Communications (2023)
-
Cell differentiation modifies the p53 transcriptional program through a combination of gene silencing and constitutive transactivation
Cell Death & Differentiation (2023)
-
Signaling pathways in cancer metabolism: mechanisms and therapeutic targets
Signal Transduction and Targeted Therapy (2023)
-
Recent advances in chromosome capture techniques unraveling 3D genome architecture in germ cells, health, and disease
Functional & Integrative Genomics (2023)
-
MiR-124-3p inhibits tumor progression in prostate cancer by targeting EZH2
Functional & Integrative Genomics (2023)