Key Points
-
The response to DNA double-strand breaks (DSBs) is a highly dynamic signalling pathway that needs constant modulation by positive and negative control points. The negative regulation of DSB signalling is crucial to ensure the correct sequence and magnitude of signalling and repair reactions, to set the boundaries of the DNA damage-induced chromatin compartment and to reverse DNA damage-induced signalling events once DNA repair is completed.
-
Negative regulation of the DSB response occurs through various mechanisms, including the enzymatic removal of post-translational modifications and the modulation of protein stability.
-
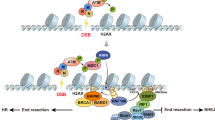
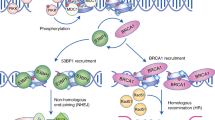
Chromatin-based DSB signalling is orchestrated by ATM (ataxia-telangiectasia mutated)-dependent phosphorylation events and RING finger 8 (RNF8)–RNF168-mediated chromatin ubiquitylation. As a consequence, cells use numerous phosphatases and deubiquitylating enzymes to balance the activities of ATM and RNF8–RNF168. In addition, RNF168 is a limiting component of DSB signalling and determines the boundaries of the DNA damage domain on chromatin.
-
Negative regulatory mechanisms are also crucial for homologous recombination-mediated DSB repair. Key nodes of negative regulation are the commitment to DNA end resection via CtBP-interacting protein (CtIP) and the ordered assembly of first replication protein A (RPA) and then RAD51 on end-resected DNA.
-
Cells use negative regulatory mechanisms to suppress DNA damage responses in physiological settings in which DNA repair reactions are not desirable, for example at telomeres and during mitosis. In addition, to evade detection and promote the transmission of their genome, DNA viruses have developed strategies to subvert and manipulate the cellular DSB response machinery.
Abstract
Single DNA lesions such as DNA double-strand breaks (DSBs) can cause cell death or trigger genome rearrangements that have oncogenic potential, and so the pathways that mend and signal DNA damage must be highly sensitive but, at the same time, selective and reversible. When initiated, boundaries must be set to restrict the DSB response to the site of the lesion. The integration of positive and, crucially, negative control points involving post-translational modifications such as phosphorylation, ubiquitylation and acetylation is key for building fast, effective responses to DNA damage and for mitigating the impact of DNA lesions on genome integrity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Jacob, F. & Monod, J. Genetic regulatory mechanisms in the synthesis of proteins. J. Mol. Biol. 3, 318–356 (1961).
Lim, W. A., Lee, C. M. & Tang, C. Design principles of regulatory networks: searching for the molecular algorithms of the cell. Mol. Cell 49, 202–212 (2013).
Chowdhury, D. et al. γ-H2AX dephosphorylation by protein phosphatase 2A facilitates DNA double-strand break repair. Mol. Cell 20, 801–809 (2005).
Gudjonsson, T. et al. TRIP12 and UBR5 suppress spreading of chromatin ubiquitylation at damaged chromosomes. Cell 150, 697–709 (2012). Demonstrates that RNF168 is a limiting component of the DSB response and that the pool of RNF168 molecules is kept in check by the E3 ubiquitin ligases TRIP12 and UBR5.
Nakada, S., Chen, G. I., Gingras, A. C. & Durocher, D. PP4 is a γH2AX phosphatase required for recovery from the DNA damage checkpoint. EMBO Rep. 9, 1019–1026 (2008).
Shao, G. et al. The Rap80–BRCC36 de-ubiquitinating enzyme complex antagonizes RNF8–Ubc13-dependent ubiquitination events at DNA double strand breaks. Proc. Natl Acad. Sci. USA 106, 3166–3171 (2009).
Acs, K. et al. The AAA-ATPase VCP/p97 promotes 53BP1 recruitment by removing L3MBTL1 from DNA double-strand breaks. Nature Struct. Mol. Biol. 18, 1345–1350 (2011).
Steger, M. et al. Prolyl isomerase PIN1 regulates DNA double-strand break repair by counteracting DNA end resection. Mol. Cell 50, 333–343 (2013).
Polo, S. E. & Jackson, S. P. Dynamics of DNA damage response proteins at DNA breaks: a focus on protein modifications. Genes Dev. 25, 409–433 (2011).
Kaidi, A. & Jackson, S. P. KAT5 tyrosine phosphorylation couples chromatin sensing to ATM signalling. Nature 498, 70–74 (2013).
Stucki, M. et al. MDC1 directly binds phosphorylated histone H2AX to regulate cellular responses to DNA double-strand breaks. Cell 123, 1213–1226 (2005).
Murga, M. et al. Global chromatin compaction limits the strength of the DNA damage response. J. Cell Biol. 178, 1101–1108 (2007).
Floyd, S. R. et al. The bromodomain protein Brd4 insulates chromatin from DNA damage signalling. Nature 498, 246–250 (2013).
Keogh, M. C. et al. A phosphatase complex that dephosphorylates γH2AX regulates DNA damage checkpoint recovery. Nature 439, 497–501 (2006).
Chowdhury, D. et al. A PP4-phosphatase complex dephosphorylates γ-H2AX generated during DNA replication. Mol. Cell 31, 33–46 (2008).
Cha, H. et al. Wip1 directly dephosphorylates γ-H2AX and attenuates the DNA damage response. Cancer Res. 70, 4112–4122 (2010).
Douglas, P. et al. Protein phosphatase 6 interacts with the DNA-dependent protein kinase catalytic subunit and dephosphorylates γ-H2AX. Mol. Cell. Biol. 30, 1368–1381 (2010).
Macurek, L. et al. Wip1 phosphatase is associated with chromatin and dephosphorylates γH2AX to promote checkpoint inhibition. Oncogene 29, 2281–2291 (2010).
Moon, S. H. et al. Wild-type p53-induced phosphatase 1 dephosphorylates histone variant γ-H2AX and suppresses DNA double strand break repair. J. Biol. Chem. 285, 12935–12947 (2010).
Xiao, A. et al. WSTF regulates the H2A.X DNA damage response via a novel tyrosine kinase activity. Nature 457, 57–64 (2008).
Cook, P. J. et al. Tyrosine dephosphorylation of H2AX modulates apoptosis and survival decisions. Nature 458, 591–596 (2009). Shows that the phosphatases EYA1 and EYA3 promote forward DNA damage signalling and repair by dephosphorylating Tyr142 of histone H2A.X.
Singh, N. et al. Dual recognition of phosphoserine and phosphotyrosine in histone variant H2A.X by DNA damage response protein MCPH1. Proc. Natl Acad. Sci. USA 109, 14381–14386 (2012).
Peng, G. et al. BRIT1/MCPH1 links chromatin remodelling to DNA damage response. Nature Cell Biol. 11, 865–872 (2009).
Wood, J. L., Singh, N., Mer, G. & Chen, J. MCPH1 functions in an H2AX-dependent but MDC1-independent pathway in response to DNA damage. J. Biol. Chem. 282, 35416–35423 (2007).
Krishnan, N. et al. Dephosphorylation of the C-terminal tyrosyl residue of the DNA damage-related histone H2A. X is mediated by the protein phosphatase eyes absent. J. Biol. Chem. 284, 16066–16070 (2009).
Galanty, Y., Belotserkovskaya, R., Coates, J. & Jackson, S. P. RNF4, a SUMO-targeted ubiquitin E3 ligase, promotes DNA double-strand break repair. Genes Dev. 26, 1179–1195 (2012).
Luo, K., Zhang, H., Wang, L., Yuan, J. & Lou, Z. Sumoylation of MDC1 is important for proper DNA damage response. EMBO J. 31, 3008–30019 (2012).
Shi, W. et al. Disassembly of MDC1 foci is controlled by ubiquitin-proteasome-dependent degradation. J. Biol. Chem. 283, 31608–31616 (2008).
Vyas, R. et al. RNF4 is required for DNA double-strand break repair in vivo. Cell Death Differ. 20, 490–502 (2013).
Yin, Y. et al. SUMO-targeted ubiquitin E3 ligase RNF4 is required for the response of human cells to DNA damage. Genes Dev. 26, 1196–1208 (2012). References 26, 27 and 30 identify the STUbL RNF4 as a DSB repair factor that controls MDC1 and RPA turnover.
Perry, J. J. P., Tainer, J. A. & Boddy, M. N. A. SIM-ultaneous role for SUMO and ubiquitin. Trends Biochem. Sci. 33, 201–208 (2008).
Zhang, D., Zaugg, K., Mak, T. W. & Elledge, S. J. A role for the deubiquitinating enzyme USP28 in control of the DNA-damage response. Cell 126, 529–542 (2006).
Huen, M. S. Y. et al. RNF8 transduces the DNA-damage signal via histone ubiquitylation and checkpoint protein assembly. Cell 131, 901–914 (2007).
Kolas, N. K. et al. Orchestration of the DNA-damage response by the RNF8 ubiquitin ligase. Science 318, 1637–1640 (2007).
Mailand, N. et al. RNF8 ubiquitylates histones at DNA double-strand breaks and promotes assembly of repair proteins. Cell 131, 887–900 (2007).
Wang, B. & Elledge, S. J. Ubc13/Rnf8 ubiquitin ligases control foci formation of the Rap80/Abraxas/Brca1/Brcc36 complex in response to DNA damage. Proc. Natl Acad. Sci. USA 104, 20759–20763 (2007).
Doil, C. et al. RNF168 binds and amplifies ubiquitin conjugates on damaged chromosomes to allow accumulation of repair proteins. Cell 136, 435–446 (2009).
Panier, S. et al. Tandem protein interaction modules organize the ubiquitin-dependent response to DNA double-strand breaks. Mol. Cell 47, 383–395 (2012).
Stewart, G. S. et al. The RIDDLE syndrome protein mediates a ubiquitin-dependent signaling cascade at sites of DNA damage. Cell 136, 420–434 (2009).
Panier, S. & Durocher, D. Regulatory ubiquitylation in response to DNA double-strand breaks. DNA Repair 8, 436–443 (2009).
Jackson, S. P. & Durocher, D. Regulation of DNA damage responses by ubiquitin and SUMO. Mol. Cell 49, 795–807 (2013).
Huang, J. et al. RAD18 transmits DNA damage signalling to elicit homologous recombination repair. Nature Cell Biol. 11, 592–603 (2009).
Shanbhag, N. M., Rafalska-Metcalf, I. U., Balane-Bolivar, C., Janicki, S. M. & Greenberg, R. A. ATM-dependent chromatin changes silence transcription in cis to DNA double-strand breaks. Cell 141, 970–981 (2010).
Bekker-Jensen, S. et al. HERC2 coordinates ubiquitin-dependent assembly of DNA repair factors on damaged chromosomes. Nature Cell Biol. 12, 80–86 (2009).
Danielsen, J. R. et al. DNA damage-inducible SUMOylation of HERC2 promotes RNF8 binding via a novel SUMO-binding zinc finger. J. Cell Biol. 197, 179–187 (2012).
Sy, S. M. et al. The ubiquitin specific protease USP34 promotes ubiquitin signaling at DNA double-strand breaks. Nucleic Acids Res. http://dx.doi.org/10.1093/nar/gkt622 (2013).
Nakada, S. et al. Non-canonical inhibition of DNA damage-dependent ubiquitination by OTUB1. Nature 466, 941–946 (2010). Demonstrates that OTUB1 inhibits RNF168-dependent chromatin ubiquitylation independently of its catalytic activity.
Edelmann, M. J. et al. Structural basis and specificity of human otubain 1-mediated deubiquitination. Biochem. J. 418, 379–390 (2009).
Mattiroli, F. et al. RNF168 ubiquitinates K13-15 on H2A/H2AX to drive DNA damage signaling. Cell 150, 1182–1195 (2012).
Juang, Y.-C. et al. OTUB1 co-opts Lys48-linked ubiquitin recognition to suppress E2 enzyme function. Mol. Cell 45, 384–397 (2012).
Sato, Y. et al. Molecular basis of K63-linked polyubiquitination inhibition by the interaction between human deubiquitinating enzyme OTUB1 and ubiquitin-conjugating enzyme UBC13. J. Biol. Chem. 287, 25860–25868 (2012).
Wiener, R., Zhang, X., Wang, T. & Wolberger, C. The mechanism of OTUB1-mediated inhibition of ubiquitination. Nature 483, 618–622 (2012).
Chen, J., Feng, W., Jiang, J., Deng, Y. & Huen, M. S. Y. Ring finger protein RNF169 antagonises the ubiquitin-dependent signaling cascade at sites of DNA damage. J. Biol. Chem. 287, 27715–2722 (2012).
Poulsen, M., Lukas, C., Lukas, J., Bekker-Jensen, S. & Mailand, N. Human RNF169 is a negative regulator of the ubiquitin-dependent response to DNA double-strand breaks. J. Cell Biol. 197, 189–199 (2012).
Cooper, E. M. et al. K63-specific deubiquitination by two JAMM/MPN+ complexes: BRISC-associated Brcc36 and proteasomal Poh1. EMBO J. 28, 621–631 (2009).
Cooper, E. M., Boeke, J. D. & Cohen, R. E. Specificity of the BRISC deubiquitinating enzyme is not due to selective binding to Lys63-linked polyubiquitin. J. Biol. Chem. 285, 10344–10352 (2010).
Feng, L., Wang, J. & Chen, J. The Lys63-specific deubiquitinating enzyme BRCC36 is regulated by two scaffold proteins localizing in different subcellular compartments. J. Biol. Chem. 285, 30982–30988 (2010).
Chen, X. BRCC36 is essential for ionizing radiation-induced BRCA1 phosphorylation and nuclear foci formation. Cancer Res. 66, 5039–5046 (2006).
Sobhian, B. et al. RAP80 targets BRCA1 to specific ubiquitin structures at DNA damage sites. Science 316, 1198–1202 (2007).
Kim, H., Chen, J. & Yu, X. Ubiquitin-binding protein RAP80 mediates BRCA1-dependent DNA damage response. Science 316, 1202–1205 (2007).
Wang, B. et al. Abraxas and RAP80 form a BRCA1 protein complex required for the DNA damage response. Science 316, 1194–1198 (2007).
Coleman, K. A. & Greenberg, R. A. The BRCA1–RAP80 complex regulates DNA repair mechanism utilization by restricting end resection. J. Biol. Chem. 286, 13669–13680 (2011).
Gatti, M. et al. A novel ubiquitin mark at the N-terminal tail of histone H2As targeted by RNF168 ubiquitin ligase. Cell Cycle 11, 2538–2544 (2012).
Komander, D., Clague, M. J. & Urbé, S. Breaking the chains: structure and function of the deubiquitinases. Nature Rev. Mol. Cell Biol. 10, 550–563 (2009).
Blickwedehl, J. et al. Proteasomes and proteasome activator 200 kDa (PA200) accumulate on chromatin in response to ionizing radiation. Radi. Res. 167, 663–674 (2007).
Butler, L. R. et al. The proteasomal de-ubiquitinating enzyme POH1 promotes the double-strand DNA break response. EMBO J. 31, 3918–3934 (2012).
Mosbech, A., Lukas, C., Bekker-Jensen, S. & Mailand, N. The deubiquitylating enzyme USP44 counteracts the DNA double-strand break response mediated by the RNF8 and RNF168 ubiquitin ligases. J. Biol. Chem. 288, 16579–16587 (2013).
Joo, H.-Y. et al. Regulation of cell cycle progression and gene expression by H2A deubiquitination. Nature 449, 1068–1072 (2007).
Botuyan, M. V. et al. Structural basis for the methylation state-specific recognition of histone H4-K20 by 53BP1 and Crb2 in DNA repair. Cell 127, 1361–1373 (2006).
Fradet-Turcotte, A. et al. 53BP1 is a reader of the DNA-damage-induced H2A Lys 15 ubiquitin mark. Nature 499, 50–54 (2013).
Zgheib, O., Pataky, K., Brugger, J. & Halazonetis, T. D. An oligomerized 53BP1 Tudor domain suffices for recognition of DNA double-strand breaks. Mol. Cell. Biol. 29, 1050–1058 (2009).
Mallette, F. A. et al. RNF8- and RNF168-dependent degradation of KDM4A/JMJD2A triggers 53BP1 recruitment to DNA damage sites. EMBO J. 31, 1865–1878 (2012).
Vaz, B., Halder, S. & Ramadan, K. Role of p97/VCP (Cdc48) in genome stability. Front. Genet. 4, 60 (2013).
Meerang, M. et al. The ubiquitin-selective segregase VCP/p97 orchestrates the response to DNA double-strand breaks. Nature Cell Biol. 13, 1376–1382 (2011).
Hsiao, K. Y. & Mizzen, C. A. Histone H4 deacetylation facilitates 53BP1 DNA damage signaling and double-strand break repair. J. Mol. Cell Biol. 5, 157–165 (2013).
Tang, J. et al. Acetylation limits 53BP1 association with damaged chromatin to promote homologous recombination. Nature Struct. Mol. Biol. 20, 317–325 (2013).
Sfeir, A. & de Lange, T. Removal of shelterin reveals the telomere end-protection problem. Science 336, 593–597 (2012).
de Lange, T. How shelterin solves the telomere end-protection problem. Cold Spring Harb. Symp. Quant. Biol. 75, 167–177 (2010).
d'Adda di Fagagna, F. et al. A DNA damage checkpoint response in telomere-initiated senescence. Nature 426, 194–198 (2003).
Denchi, E. L. & de Lange, T. Protection of telomeres through independent control of ATM and ATR by TRF2 and POT1. Nature 448, 1068–1071 (2007).
Takai, H., Smogorzewska, A. & de Lange, T. DNA damage foci at dysfunctional telomeres. Curr. Biol. 13, 1549–1556 (2003).
van Steensel, B., Smogorzewska, A. & de Lange, T. TRF2 protects human telomeres from end-to-end fusions. Cell 92, 401–413 (1998).
Okamoto, K. et al. A two-step mechanism for TRF2-mediated chromosome-end protection. Nature 494, 502–505 (2013). Shows that the shelterin component TRF2 inhibits RNF168-dependent chromatin ubiquitylation at telomeres by recruiting BRCC3 and UBR5.
Karlseder, J. et al. The telomeric protein TRF2 binds the ATM kinase and can inhibit the ATM-dependent DNA damage response. PLoS Biol. 2, E240 (2004).
Giunta, S., Belotserkovskaya, R. & Jackson, S. P. DNA damage signaling in response to double-strand breaks during mitosis. J. Cell Biol. 190, 197–207 (2010).
van Vugt, M. A. et al. A mitotic phosphorylation feedback network connects Cdk1, Plk1, 53BP1, and Chk2 to inactivate the G2/M DNA damage checkpoint. PLoS Biol. 8, e1000287 (2010).
Zhang, W., Peng, G., Lin, S. Y. & Zhang, P. DNA damage response is suppressed by the high cyclin-dependent kinase 1 activity in mitotic mammalian cells. J. Biol. Chem. 286, 35899–35905 (2011).
Lilley, C. E., Chaurushiya, M. S., Boutell, C., Everett, R. D. & Weitzman, M. D. The intrinsic antiviral defense to incoming HSV-1 genomes includes specific DNA repair proteins and is counteracted by the viral protein ICP0. PLoS Pathog. 7, e1002084 (2011).
Lilley, C. E. et al. A viral E3 ligase targets RNF8 and RNF168 to control histone ubiquitination and DNA damage responses. EMBO J. 29, 943–955 (2010).
Chaurushiya, M. S. et al. Viral E3 ubiquitin ligase-mediated degradation of a cellular E3: viral mimicry of a cellular phosphorylation mark targets the RNF8 FHA domain. Mol. Cell 46, 79–90 (2012).
Boutell, C. et al. A viral ubiquitin ligase has substrate preferential SUMO targeted ubiquitin ligase activity that counteracts intrinsic antiviral defence. PLoS Pathog. 7, e1002245 (2011).
Weitzman, M. D., Lilley, C. E. & Chaurushiya, M. S. Changing the ubiquitin landscape during viral manipulation of the DNA damage response. FEBS Lett. 585, 2897–2906 (2011).
Symington, L. S. & Gautier, J. Double-strand break end resection and repair pathway choice. Annu. Rev. Genet. 45, 247–271 (2011).
Chapman, J. R., Taylor, M. R. G. & Boulton, S. J. Playing the end game: DNA double-strand break repair pathway choice. Mol. Cell 47, 497–510 (2012).
Lieber, M. R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu. Rev. Biochem. 79, 181–211 (2010).
Zimmermann, M., Lottersberger, F., Buonomo, S. B., Sfeir, A. & de Lange, T. 53BP1 regulates DSB repair using Rif1 to control 5′ end resection. Science 339, 700–704 (2013).
Feng, L., Fong, K.-W., Wang, J., Wang, W. & Chen, J. RIF1 counteracts BRCA1-mediated end resection during DNA repair. J. Biol. Chem. 288, 11135–11143 (2013).
Escribano-Diaz, C. et al. A cell cycle-dependent regulatory circuit composed of 53BP1–RIF1 and BRCA1–CtIP controls DNA repair pathway choice. Mol. Cell 49, 872–883 (2013).
Di Virgilio, M. et al. Rif1 prevents resection of DNA breaks and promotes immunoglobulin class switching. Science 339, 711–715 (2013).
Chapman, J. R. et al. RIF1 is essential for 53BP1-dependent nonhomologous end joining and suppression of DNA double-strand break resection. Mol. Cell 49, 858–871 (2013). References 96–100 identify RIF1 as a 53BP1 effector protein during DSB repair pathway choice.
Callen, E. et al. 53BP1 mediates productive and mutagenic DNA repair through distinct phosphoprotein interactions. Cell 153, 1266–1280 (2013).
Langerak, P., Mejia-Ramirez, E., Limbo, O. & Russell, P. Release of Ku and MRN from DNA ends by Mre11 nuclease activity and Ctp1 is required for homologous recombination repair of double-strand breaks. PLoS Genet. 7, e1002271 (2011).
Kaidi, A., Weinert, B. T., Choudhary, C. & Jackson, S. P. Human SIRT6 promotes DNA end resection through CtIP deacetylation. Science 329, 1348–1353 (2010). Discovery that deacetylation of CtIP is a prerequisite for efficient DNA end resection and homologous recombination.
Dou, H., Huang, C., Singh, M., Carpenter, P. B. & Yeh, E. T. Regulation of DNA repair through deSUMOylation and SUMOylation of replication protein A complex. Mol. Cell 39, 333–345 (2010).
Antony, E. et al. Srs2 disassembles Rad51 filaments by a protein–protein interaction triggering ATP turnover and dissociation of Rad51 from DNA. Mol. Cell 35, 105–115 (2009).
Krejci, L. et al. DNA helicase Srs2 disrupts the Rad51 presynaptic filament. Nature 423, 305–309 (2003).
Veaute, X. et al. The Srs2 helicase prevents recombination by disrupting Rad51 nucleoprotein filaments. Nature 423, 309–312 (2003).
Moldovan, G.-L. et al. Inhibition of homologous recombination by the PCNA-interacting protein PARI. Mol. Cell 45, 75–86 (2012).
Roy, R., Chun, J. & Powell, S. N. BRCA1 and BRCA2: different roles in a common pathway of genome protection. Nature Rev. Cancer 12, 68–78 (2012).
Esashi, F. et al. CDK-dependent phosphorylation of BRCA2 as a regulatory mechanism for recombinational repair. Nature 434, 598–604 (2005).
Kasparek, T. R. & Humphrey, T. C. DNA double-strand break repair pathways, chromosomal rearrangements and cancer. Seminars Cell Dev. Biol. 22, 886–897 (2011).
Silverman, J., Takai, H., Buonomo, S. B. C., Eisenhaber, F. & de Lange, T. Human Rif1, ortholog of a yeast telomeric protein, is regulated by ATM and 53BP1 and functions in the S-phase checkpoint. Genes Dev. 18, 2108–2119 (2004).
Xu, D. et al. Rif1 provides a new DNA-binding interface for the Bloom syndrome complex to maintain normal replication. EMBO J. 29, 3140–3155 (2010).
Kim, J. M. et al. Inactivation of murine Usp1 results in genomic instability and a Fanconi anemia phenotype. Dev. Cell 16, 314–320 (2009). Demonstrates that USP1-mediated deubiquitylation of FANCD2 is a crucial step in the Fanconi anaemia pathway in vivo.
Harreman, M. et al. Distinct ubiquitin ligases act sequentially for RNA polymerase II polyubiquitylation. Proc. Natl Acad. Sci. USA 106, 20705–20710 (2009).
Kruse, J.-P. & Gu, W. Modes of p53 regulation. Cell 137, 609–622 (2009).
Purvis, J. E. et al. p53 dynamics control cell fate. Science 336, 1440–1444 (2012). Elegantly demonstrates that p53 dynamics are directly translated into the cell's decision to promote DNA repair or induce senescence.
Altmeyer, M. & Lukas, J. Guarding against collateral damage during chromatin transactions. Cell 153, 1431–1434 (2013).
Heyer, W.-D., Ehmsen, K. T. & Liu, J. Regulation of homologous recombination in eukaryotes. Annu. Rev. Genet. 44, 113–139 (2010).
Acknowledgements
The authors apologize to those whose important findings could not be mentioned as primary literature and/or cited owing to space constraints. The authors thank R. Szilard for critically reading the manuscript and S. Boulton for comments and for supporting S.P.'s participation in this Review. S.P. is supported by an European Molecular Biology Organization (EMBO) long-term fellowship. D.D. is the Thomas Kierans Chair in Mechanisms of Cancer Development and a Canada Research Chair (Tier 1) in Molecular Mechanisms of Genome Integrity. Work in the laboratory of D.D. is supported by a grant-in-aid from the Krembil Foundation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- DNA damage checkpoint
-
Signalling pathways that delay or arrest cell cycle progression in response to DNA damage.
- Linker histone
-
Histone that is structurally and functionally distinct from nucleosomal core histones. It provides an interaction platform for numerous chromatin components and participates in the formation of the 30 nm chromatin fibre by linking nucleosome core particles.
- Bromodomain
-
Protein–protein interaction domain that binds to acetylated Lys residues.
- RING
-
Protein domain that is present in many E3 ubiquitin ligases. Contains conserved His and Cys residues and coordinates two Zn2+ ions.
- E3 ubiquitin ligases
-
Key enzymes in the ubiquitylation reaction that are required for the attachment of ubiquitin moieties to a substrate protein. The E3 provides substrate specificity to a multistep reaction that first involves an E1 activating enzyme followed by an E2 ubiquitin-conjugating enzyme and then an E3 ubiquitin ligase.
- 26S proteasome
-
A large multisubunit protease complex that degrades polyubiquitylated proteins with certain ubiquitin chain topology. In mammals, it consists of a 20S proteolytic core particle and one or two 19S regulatory particles.
- Degradative ubiquitin conjugates
-
Polyubiquitin chains that target proteins for degradation by the 26S proteasome. The best-studied examples are Lys48-linked ubiquitin chains.
- Regulatory ubiquitylation
-
The addition of ubiquitin conjugates that regulate protein function instead of acting as a signal for proteasomal degradation. An example for regulatory ubiquitin conjugates are Lys63-linked ubiquitin chains.
- E2 ubiquitin-conjugating enzymes
-
Enzymes that are required for the second step during ubiquitin conjugation. They receive activated ubiquitin from the E1 activating enzyme and then interact with an E3 ubiquitin ligase to attach the ubiquitin moiety to the substrate protein.
- LRM
-
Small, linear motif that can be found adjacent to some ubiquitin-binding domains. Provides ligand specificity to ubiquitin-dependent protein–protein interactions.
- JAMM–MPN+
-
A Zn2+-coordinating protein domain that confers isopeptidase activity and is present in a small subset of deubiquitylating enzymes. It is also known as the JAB1–MPN–MOV34 metalloenzyme domain.
- Polycomb group
-
A group of proteins that have important roles in the maintenance of homeotic gene repression during development and stem cell renewal.
- Tudor domains
-
Protein–protein interaction domains that were first found in the Drosophila melanogaster protein Tudor. These domains bind to methylated Arg or Lys residues.
- Prolyl isomerase
-
Catalyses the cis–trans isomerization of peptide bonds that are amino-terminal to Pro residues in polypeptide chains.
- Fanconi anaemia pathway
-
A DNA damage signalling pathway that coordinates the repair of DNA interstrand crosslinks. Germline mutations in genes encoding key components of this pathway are the underlying cause of Fanconi anaemia, which is characterized by congenital defects, cancer susceptibility and cellular hypersensitivity to DNA crosslinking agents.
Rights and permissions
About this article
Cite this article
Panier, S., Durocher, D. Push back to respond better: regulatory inhibition of the DNA double-strand break response. Nat Rev Mol Cell Biol 14, 661–672 (2013). https://doi.org/10.1038/nrm3659
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm3659
This article is cited by
-
DEtail-seq is an ultra-efficient and convenient method for meiotic DNA break profiling in multiple organisms
Science China Life Sciences (2023)
-
VCP maintains nuclear size by regulating the DNA damage-associated MDC1–p53–autophagy axis in Drosophila
Nature Communications (2021)
-
DNA repair pathways and their roles in drug resistance for lung adenocarcinoma
Molecular Biology Reports (2021)
-
ULK1 inhibition overcomes compromised antigen presentation and restores antitumor immunity in LKB1-mutant lung cancer
Nature Cancer (2021)
-
ATM-associated signalling triggers the unfolded protein response and cell death in response to stress
Communications Biology (2020)