Key Points
-
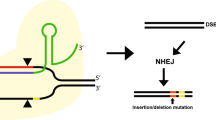
A new strategy for gene therapy that aims to correct genetic mutations directly in the chromosome is under development.
-
This approach uses single-stranded DNA to activate the DNA repair pathways of the cell and to direct the exchange of mutant bases at precise locations in the chromosome.
-
These single-stranded DNA molecules can be delivered into cells without the aid of viruses, which are the classical vector/vehicle that is used for gene therapy.
-
If successful, the 'gene-repair process' will probably reduce the alarming side effects that are encountered by the viral gene-therapy strategies.
-
The key to success lies in the capacity to correct or reverse mutations at a frequency that will bring therapeutic benefits.
-
The progress, feasibility and applicability of this strategy are reviewed, and approaches towards overcoming the barriers to success are also discussed.
Abstract
A technique that can direct the repair of a genetic mutation in a human chromosome using the DNA repair machinery of the cell is under development. Although this approach is not as mature as other forms of gene therapy and fundamental problems continue to arise, it promises to be the ultimate therapy for many inherited disorders. There is a continuing effort to understand the potential and the limitations of this controversial approach.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Thomas, C. E., Ehrhardt, A. & Kay, M. A. Progress and problems with the use of viral vectors for gene therapy. Nature Rev. Genet. 4, 346–358 (2003).
Andreani, M. et al. Long-term survival of ex-thalassemic patients with persistent mixed chimerism after bone marrow transplantation. Bone Marrow Transplant 25, 401–404 (2000).
Hayashi, S. et al. Mixed chimerism following in utero hematopoietic stem cell transplantation in murine models of hemoglobinopathy. Exp. Hematol. 31, 176–184 (2003).
Vasquez, K. M., Marburger, K., Intody, Z. & Wilson, J. H. Manipulating the mammalian genome by homologous recombination. Proc. Natl Acad. Sci. USA 98, 8403–8410 (2001).
Kotani, H., Sekiguchi, J. M., Dutta, S. & Kmiec, E. B. Genetic recombination of nucleosomal templates is mediated by transcription. Mol. Gen. Genet. 244, 410–419 (1994). This paper describes the initial observations that led to the design of the chimaera.
Kotani, H. & Kmiec, E. B. DNA cruciforms facilitate in vitro strand transfer on nucleosomal templates. Mol. Gen. Genet. 243, 681–690 (1994).
Paques, F. & Haber, J. E. Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 63, 349–404 (1999).
Moerschell, R. P., Tsunasawa, S. & Sherman, F. Transformation of yeast with synthetic oligonucleotides. Proc. Natl Acad. Sci. USA 85, 524–528 (1988). A pioneering paper that describes the use of oligonucleotides in yeast targeting.
Yamamoto, T., Moerschell, R. P., Wakem, L. P., Komar-Panicucci, S. & Sherman, F. Strand-specificity in the transformation of yeast with synthetic oligonucleotides. Genetics 131, 811–819 (1992).
Yamamoto, T., Moerschell, R. P., Wakem, L. P., Ferguson, D. & Sherman, F. Parameters affecting the frequencies of transformation and co-transformation with synthetic oligonucleotides in yeast. Yeast 8, 935–948 (1992).
Campbell, C. R., Keown, W., Lowe, L., Kirschling, D. & Kucherlapati, R. Homologous recombination involving small single-stranded oligonucleotides in human cells. New Biol. 1, 223–227 (1989).
Gamper, H. B. et al. The DNA strand of chimeric RNA/DNA oligonucleotides can direct gene repair/conversion activity in mammalian and plant cell-free extracts. Nucleic Acids Res. 28, 4332–4339 (2000). This article reports that gene repair using single-stranded oligonucleotides is as active as the chimaera and easier to synthesize.
Kmiec, E. B. Targeted gene repair. Gene Ther. 6, 1–3 (1999).
Kmiec, E. B., Kren, B. T. & Steer, C. J. in The Development of Human Gene Therapy (ed. Friedmann, T.) 101–110 (Cold Spring Harbor Laboratory Press, New York, 1998).
Kotani, H. & Kmiec, E. B. Transcription activates RecA-promoted homologous pairing of nucleosomal DNA. Mol. Cell Biol. 14, 1949–1955 (1994).
Kotani, H. et al. RNA facilitates RecA-mediated DNA pairing and strand transfer between molecules bearing limited regions of homology. Mol. Gen. Genet. 250, 626–634 (1996). This was one of the first studies to show that RNA can reduce the level of homology that is required for strand exchange.
Cole-Strauss, A. et al. Targeted gene repair directed by the chimeric RNA/DNA oligonucleotide in a mammalian cell-free extract. Nucleic Acids Res. 27, 1323–1330 (1999).
Igoucheva, O., Alexeev, V. & Yoon, K. Targeted gene correction by small single-stranded oligonucleotides in mammalian cells. Gene Ther. 8, 391–399 (2001). A careful study of the effects of single-stranded oligonucleotides in mammalian cells.
Igoucheva, O. & Yoon, K. Targeted single-base correction by RNA–DNA oligonucleotides. Hum. Gene Ther. 11, 2307–2312 (2000).
Igoucheva, O., Peritz, A. E., Levy, D. & Yoon, K. A sequence-specific gene correction by an RNA–DNA oligonucleotide in mammalian cells characterized by transfection and nuclear extract using a lacZ shuttle system. Gene Ther. 6, 1960–1971 (1999).
Gamper, H. B., Nulf, C. J., Corey, D. R. & Kmiec, E. B. The synaptic complex of RecA protein participates in hybridization and inverse strand exchange reactions. Biochemistry 42, 2643–2655 (2003).
Liu, L., Rice, M. C. & Kmiec, E. B. In vivo gene repair of point and frameshift mutations directed by chimeric RNA/DNA oligonucleotides and modified single-stranded oligonucleotides. Nucleic Acids Res. 29, 4238–4250 (2001).
Liu, L., Cheng, S., van Brabant, A. J. & Kmiec, E. B. Rad51p and Rad54p, but not Rad52p, elevate gene repair in Saccharomyces cerevisiae directed by modified single-stranded oligonucleotide vectors. Nucleic Acids Res. 31, 2742–2750 (2002). This study was the first to show that Rad51 stimulates gene repair in yeast.
Liu, L. et al. Strand bias in targeted gene repair is influenced by transcriptional activity. Mol. Cell Biol. 22, 3852–3863 (2002).
Brachman, E. E. & Kmiec, E. B. Targeted gene repair of cyc1 mutations in Saccharomyces cerevisiae directed by modified single-stranded DNA oligonucleotides. Genetics 163, 527–538 (2003).
Metz, R. et al. Mode of action of RNA/DNA oligonucleotides: progress in the development of gene repair as a therapy for α(1)-antitrypsin deficiency. Chest 121, 91–97 (2002).
Jayasena, V. K. & Johnston, B. H. Complement-stabilized D-loop. RecA-catalyzed stable pairing of linear DNA molecules at internal sites. J. Mol. Biol. 230, 1015–1024 (1993).
Belotserkovskii, B. P., Reddy, G. & Zarling, D. A. DNA hybrids stabilized by heterologies. Biochemistry 38, 10785–10792 (1999).
Belotserkovskii, B. P. & Zarling, D. A. Peptide nucleic acid (PNA) facilitates multistranded hybrid formation between linear double-stranded DNA targets and RecA protein-coated complementary single-stranded DNA probes. Biochemistry 41, 3686–3692 (2002).
Parekh-Olmedo, H., Drury, M. & Kmiec, E. B. Targeted nucleotide exchange in Saccharomyces cerevisiae directed by short oligonucleotides containing locked nucleic acids. Chem. Biol. 9, 1073–1084 (2002).
Drury, M. & Kmiec, E. B. DNA pairing is an important step in the process of targeted nucleotide exchange. Nucleic Acids Res. 31, 899–910 (2003).
Gruenert, D. C. Opportunities and challenges in targeting genes for therapy. Gene Ther. 6, 1347–1348 (1999).
Sangiuolo, F. et al. In vitro correction of cystic fibrosis epithelial cell lines by small fragment homologous replacement (SFHR) technique. BMC Med. Genet. 3, 8 (2002).
Kapsa, R. et al. In vivo and in vitro correction of the mdx dystrophin gene nonsense mutation by short-fragment homologous replacement. Hum. Gene Ther. 12, 629–642 (2001).
Liu, X. M. et al. Targeted correction of genomic single base-pair mutations using adeno-associated virus under non-selective conditions. Mol. Ther. 7, 159–160 (2003).
Check, E. Harmful potential of viral vectors fuels doubts over gene therapy. Nature 423, 573–574 (2003).
Brachman, E. E. & Kmiec, E. B. The 'biased' evolution of targeted gene repair. Curr. Opin. Mol. Ther. 4, 171–176 (2002).
Andersen, M. S., Sorensen, C. B., Bolund, L. & Jensen, T. G. Mechanisms underlying targeted gene correction using chimeric RNA/DNA and single-stranded DNA oligonucleotides. J. Mol. Med. 80, 770–781 (2002). This is a good review of the field of oligonucleotide targeting, with an emphasis on the mechanism.
Thorpe, P., Stevenson, B. J. & Porteous, D. J. Optimising gene repair strategies in cell culture. Gene Ther. 9, 700–702 (2002).
Thorpe, P. H., Stevenson, B. J. & Porteous, D. J. Functional correction of episomal mutations with short DNA fragments and RNA–DNA oligonucleotides. J. Gene Med. 4, 195–204 (2002).
Yanez, R. J. & Porter, A. C. Gene targeting is enhanced in human cells overexpressing hRAD51. Gene Ther. 6, 1282–1290 (1999).
Yanez, R. J. & Porter, A. C. Influence of DNA delivery method on gene targeting frequencies in human cells. Somat. Cell Mol. Genet. 25, 27–31 (1999).
Mazin, A. V., Bornarth, C. J., Solinger, J. A., Heyer, W. D. & Kowalczykowski, S. C. Rad54 protein is targeted to pairing loci by the Rad51 nucleoprotein filament. Mol. Cell 6, 583–592 (2000).
Mazin, A. V., Zaitseva, E., Sung, P. & Kowalczykowski, S. C. Tailed duplex DNA is the preferred substrate for Rad51 protein-mediated homologous pairing. EMBO J. 19, 1148–1156 (2000).
Baumann, P., Benson, F. E. & West, S. C. Human Rad51 protein promotes ATP-dependent homologous pairing and strand transfer reactions in vitro. Cell 87, 757–766 (1996).
Igoucheva, O., Alexeev, V., Pryce, M. & Yoon, K. Transcription affects formation and processing of intermediates in oligonucleotide-mediated gene alteration. Nucleic Acids Res. 31, 2659–2670 (2003).
Lahue, R. S., Au, K. G. & Modrich, P. DNA mismatch correction in a defined system. Science 245, 160–164 (1989).
Buermeyer, A. B., Deschenes, S. M., Baker, S. M. & Liskay, R. M. Mammalian DNA mismatch repair. Annu. Rev. Genet. 33, 533–564 (1999).
Kolodner, R. D. & Marsischky, G. T. Eukaryotic DNA mismatch repair. Curr. Opin. Genet. Dev. 9, 89–96 (1999).
Rice, M. C., Bruner, M., Czymmek, K. & Kmiec, E. B. In vitro and in vivo nucleotide exchange directed by chimeric RNA/DNA oligonucleotides in Saccharomyces cerevisiae. Mol. Microbiol. 40, 857–868 (2001).
Batty, D. P. & Wood, R. D. Damage recognition in nucleotide excision repair of DNA. Gene 241, 193–204 (2000).
Wood, R. D. Nucleotide excision repair in mammalian cells. J. Biol. Chem. 272, 23465–23468 (1997).
Ellis, H. M., Yu, D., DiTizio, T. & Court, D. L. High efficiency mutagenesis, repair, and engineering of chromosomal DNA using single-stranded oligonucleotides. Proc. Natl Acad. Sci. USA 98, 6742–6746 (2001).
Cole-Strauss, A. et al. Correction of the mutation responsible for sickle cell anemia by an RNA–DNA oligonucleotide. Science 273, 1386–1389 (1996).
Xiang, Y., Cole-Strauss, A., Yoon, K., Gryn, J. & Kmiec, E. B. Targeted gene conversion in a mammalian CD34+-enriched cell population using a chimeric RNA/DNA oligonucleotide. J. Mol. Med. 75, 829–835 (1997).
Liu, H., Agarwal, S., Kmiec, E. & Davis, B. R. Targeted β-globin gene conversion in human hematopoietic CD34(+) and Lin(−)CD38(−)cells. Gene Ther. 9, 118–126 (2002).
Alexeev, V. & Yoon, K. Stable and inheritable changes in genotype and phenotype of albino melanocytes induced by an RNA–DNA oligonucleotide. Nature Biotechnol. 16, 1343–1346 (1998).
Alexeev, V., Igoucheva, O., Domashenko, A., Cotsarelis, G. & Yoon, K. Localized in vivo genotypic and phenotypic correction of the albino mutation in skin by RNA–DNA oligonucleotide. Nature Biotechnol. 18, 43–47 (2000).
Yoon, K., Cole-Strauss, A. & Kmiec, E. B. Targeted gene correction of episomal DNA in mammalian cells mediated by a chimeric RNA.DNA oligonucleotide. Proc. Natl Acad. Sci. USA 93, 2071–2076 (1996).
Kren, B. T., Cole-Strauss, A., Kmiec, E. B. & Steer, C. J. Targeted nucleotide exchange in the alkaline phosphatase gene of HuH-7 cells mediated by a chimeric RNA/DNA oligonucleotide. Hepatology 25, 1462–1468 (1997).
Santana, E., Peritz, A. E., Lyer, S., Uitto, J. & Yoon, K. Different frequency of gene targeting events by the RNA–DNA oligonucleotide among epithelial cells. J. Invest. Dermatol. 111, 1172–1177 (1998).
Kren, B. T., Bandyopadhyay, P. & Steer, C. J. In vivo site-directed mutagenesis of the factor IX gene by chimeric RNA/DNA oligonucleotides. Nature Med. 4, 285–290 (1998). This study presents in vivo evidence of chimeric oligonucleotide activity in animals, which is supported by strong genotypic and phenotypic data.
Bandyopadhyay, P., Ma, X., Linehan-Stieers, C., Kren, B. T. & Steer, C. J. Nucleotide exchange in genomic DNA of rat hepatocytes using RNA/DNA oligonucleotides. Targeted delivery of liposomes and polyethyleneimine to the asialoglycoprotein receptor. J. Biol. Chem. 274, 10163–10172 (1999).
Tagalakis, A. D., Graham, I. R., Riddell, D. R., Dickson, J. G. & Owen, J. S. Gene correction of the apolipoprotein (Apo) E2 phenotype to wild-type ApoE3 by in situ chimeraplasty. J. Biol. Chem. 276, 13226–13230 (2001).
Lai, L. -W. & Lien, Y. H. Homologous recombination based gene therapy. Expt. Nephrol. 7, 11–14 (1999).
Lai, L. W. & Lien, Y. H. Chimeric RNA/DNA oligonucleotide-based gene therapy. Kidney Int. 61 (Suppl.), 47–51 (2002).
Bartlett, R. J. et al. In vivo targeted repair of a point mutation in the canine dystrophin gene by a chimeric RNA/DNA oligonucleotide. Nature Biotechnol. 18, 615–622 (2000). This article describes gene repair in a dog model for muscular dystrophy, along with an excellent discussion of what each assay shows.
Rando, T. A., Disatnik, M. H. & Zhou, L. Z. Rescue of dystrophin expression in mdx mouse muscle by RNA/DNA oligonucleotides. Proc. Natl Acad. Sci. USA 97, 5363–5368 (2000). An excellent article that outlines gene repair in a well-characterized animal model and provides a good phenotypic analysis.
Bertoni, C. & Rando, T. A. Dystrophin gene repair in mdx muscle precursor cells in vitro and in vivo mediated by RNA–DNA chimeric oligonucleotides. Hum. Gene Ther. 13, 707–718 (2002).
Li, Z. H., Liu, D. P., Yin, W. X., Guo, Z. C. & Liang, C. C. Targeted correction of the point mutations of β-thalassemia and targeted mutagenesis of the nucleotide associated with HPFH by RNA/DNA oligonucleotides: potential for β-thalassemia gene therapy. Blood Cells Mol. Dis. 27, 530–538 (2001).
van der Steege, G. et al. Persistent failures in gene repair. Nature Biotechnol. 19, 305–306 (2001).
Thomas, K. R. & Capecchi, M. R. Recombinant DNA technique and sickle cell anemia research. Science 275, 1404–1405 (1997).
Graham, I. R. et al. Gene repair validation. Nature Biotechnol. 19, 507–508 (2001).
Thorpe, P. H., Stevens, B. J. & Porteous, D. J. Comparing two strategies for functional gene correction (2001).
Bandyopadhyay, P., Kren, B. T., Ma, X. & Steer, C. J. Enhanced gene transfer into HuH-7 cells and primary rat hepatocytes using targeted liposomes and polyethylenimine. Biotechniques 25, 282–292 (1998).
Degtyareva, N., Subramanian, D. & Griffith, J. D. Analysis of the binding of p53 to DNAs containing mismatched and bulged bases. J. Biol. Chem. 276, 8778–8784 (2001).
Dudenhoffer, C., Rohaly, G., Will, K., Deppert, W. & Wiesmuller, L. Specific mismatch recognition in heteroduplex intermediates by p53 suggests a role in fidelity control of homologous recombination. Mol. Cell Biol. 18, 5332–5342 (1998).
Colosimo, A., Goncz, K. K., Novelli, G., Dallapiccola, B. & Gruenert, D. C. Targeted correction of a defective selectable marker gene in human epithelial cells by small DNA fragments. Mol. Ther. 3, 178–185 (2001).
Krejci, L., Damborsky, J., Thomsen, B., Duno, M. & Bendixen, C. Molecular dissection of interactions between Rad51 and members of the recombination-repair group. Mol. Cell Biol. 21, 966–976 (2001).
Rice, K. P., Eggler, A. L., Sung, P. & Cox, M. M. DNA pairing and strand exchange by the Escherichia coli RecA and yeast Rad51 proteins without ATP hydrolysis: on the importance of not getting stuck. J. Biol. Chem. 276, 38570–38581 (2001).
Alexiadis, V. & Kadonaga, J. T. Strand pairing by Rad54 and Rad51 is enhanced by chromatin. Genes Dev. 16, 2767–2771 (2002).
Vispe, S., Cazaux, C., Lesca, C. & Defais, M. Overexpression of Rad51 protein stimulates homologous recombination and increases resistance of mammalian cells to ionizing radiation. Nucleic Acids Res. 26, 2859–2864 (1998).
Burke, D. T., Carle, G. F. & Olson, M. V. Cloning of large segments of exogenous DNA into yeast by means of artificial chromosome vectors. Science 236, 806–812 (1987).
Cimino, G. D., Gamper, H. B., Isaacs, S. T. & Hearst, J. E. Psoralens as photoactive probes of nucleic acid structure and function: organic chemistry, photochemistry, and biochemistry. Annu. Rev. Biochem. 54, 1151–1193 (1985).
Faruqi, A. F., Seidman, M. M., Segal, D. J., Carroll, D. & Glazer, P. M. Recombination induced by triple-helix-targeted DNA damage in mammalian cells. Mol. Cell Biol. 16, 6820–6828 (1996).
Sandor, Z. & Bredberg, A. Triple helix directed psoralen adducts induce a low frequency of recombination in an SV40 shuttle vector. Biochim. Biophys. Acta 1263, 235–240 (1995).
Vasquez, K. M., Wensel, T. G., Hogan, M. E. & Wilson, J. H. High-affinity triple helix formation by synthetic oligonucleotides at a site within a selectable mammalian gene. Biochemistry 34, 7243–7251 (1995).
Vasquez, K. M., Wensel, T. G., Hogan, M. E. & Wilson, J. H. High-efficiency triple-helix-mediated photo-cross-linking at a targeted site within a selectable mammalian gene. Biochemistry 35, 10712–10719 (1996).
Kren, B. T., Wong, P. Y. & Steer, C. J. Short, single-stranded oligonucleotides mediate targeted nucleotide conversion using extracts from isolated liver mitochondria. DNA Repair 2, 531–546 (2003).
Dekker, M., Brouwers, C. & Te, R. H. Targeted gene modification in mismatch-repair-deficient embryonic stem cells by single-stranded DNA oligonucleotides. Nucleic Acids Res. 31, 27 (2003). This paper represents an important step in the use of single-stranded DNA for targeting in mouse ES cells.
Fyodorov, D. V. & Kadonaga, J. T. The many faces of chromatin remodeling: SWItching beyond transcription. Cell 106, 523–525 (2001).
Britt, A. B. & May, G. D. Re-engineering plant gene targeting. Trends Plant Sci. 8, 90–95 (2003).
Lu, I. -L. et al. Correction/mutation of acid α-D-glucosidase gene by modified single-stranded oligonucleotides: in vitro and in vivo studies. Gene Therapy (in the press). One of the most complete studies using single-stranded vectors, with convincing genotype and phenotype evidence of conversion.
Parekh-Olmedo, H., Czymmek, K. & Kmiec, E. B. Targeted gene repair in mammalian cells using chimeric RNA/DNA oligonucleotides and modified single-stranded vectors. Sci. STKE 2001, 1 (2001).
Fortin, G. S. & Symington, L. S. Mutations in yeast Rad51 that partially bypass the requirement for Rad55 and Rad57 in DNA repair by increasing the stability of Rad51–DNA complexes. EMBO J. 21, 3160–3170 (2002).
Kren, B. T. et al. Correction of the UDP-glucuronosyltransferase gene defect in the gunn rat model of crigler-najjar syndrome type I with a chimeric oligonucleotide. Proc. Natl Acad. Sci. USA 96, 10349–10354 (1999).
Gamper, H. B. et al. A plausible mechanism for gene correction by chimeric oligonucleotides. Biochemistry 39, 5808–5816 (2000).
Chen, Z. et al. Mitochondria isolated from liver contain the essential factors required for RNA/DNA oligonucleotide-targeted gene repair. Biochem. Biophys. Res. Commun. 285, 188–194 (2001).
Kmiec, E. B., Johnson, C. & May, G. D. Chloroplast lysates support directed mutagenesis via modified DNA and chimeric RNA/DNA oligonucleotides. Plant J. 27, 267–274 (2001).
Rice, M. C., May, G. D., Kipp, P. B., Parekh, H. & Kmiec, E. B. Genetic repair of mutations in plant cell-free extracts directed by specific chimeric oligonucleotides. Plant Physiol. 123, 427–438 (2000).
Beetham, P. R., Kipp, P. B., Sawycky, X. L., Arntzen, C. J. & May, G. D. A tool for functional plant genomics: chimeric RNA/DNA oligonucleotides cause in vivo gene-specific mutations. Proc. Natl Acad. Sci. USA 96, 8774–8778 (1999). The first report of gene repair in plants using chimeric RNA–DNA oligonucleotides.
Kochevenko, A. & Willmitzer, L. Chimeric RNA/DNA oligonucleotide-based site-specific modification of the tobacco acetolactate syntase gene. Plant Physiol. 132, 174–184 (2003).
Zhu, T. et al. Targeted manipulation of maize genes in vivo using chimeric RNA/DNA oligonucleotides. Proc. Natl Acad. Sci. USA 96, 8768–8773 (1999).
Zhu, T., Mettenburg, K., Peterson, D. J., Tagliani, L. & Baszczynski, C. L. Engineering herbicide-resistant maize using chimeric RNA/DNA oligonucleotides. Nature Biotechnol. 18, 555–558 (2000).
Acknowledgements
We are grateful to many of our friends and colleagues who have been so supportive and encouraged us to be persistent. We thank the NIH and NaPro BioTherapeutics for financial support.
Author information
Authors and Affiliations
Corresponding author
Related links
Related links
Databases
LocusLink
OMIM
Saccharomyces Genome Database
Further Information
Glossary
- PHOSPHOROTHIOATE
-
A modification that provides nuclease resistance in an oligonucleotide in which a non-bridging oxygen is replaced by a sulphur atom in the oligophosphate backbone.
- EPISOMAL
-
In the context of transient transfection, this term refers to a plasmid target that is extra chromosomal.
- FLUORESCENCE-ACTIVATED CELL SORTING
-
(FACS). A method whereby dissociated and individual living cells are sorted, in a liquid stream, according to the intensity of fluorescence that they emit as they pass through a laser beam.
- PSORALEN
-
A photosensitizing chemical that is used for determining RNA–DNA structures in cells, and intercalates between two strands in duplex DNA. When attached to an oligonucleotide, psoralen forms interstrand crosslinks. When exposed to ultraviolet light, it forms photoadducts, crosslinked chemical bonds within adjacent bases.
- MONO-ADDUCTS
-
A form of DNA lesion induced by DNA damaging agents, such as ultraviolet radiation, which on longer exposure can be converted into covalent crosslinks in the DNA. Mono-adducts can, to an extent, induce recombination in yeast, mammalian and bacterial cells.
Rights and permissions
About this article
Cite this article
Liu, L., Parekh-Olmedo, H. & Kmiec, E. The development and regulation of gene repair. Nat Rev Genet 4, 679–689 (2003). https://doi.org/10.1038/nrg1156
Issue Date:
DOI: https://doi.org/10.1038/nrg1156
This article is cited by
-
Analyses of point mutation repair and allelic heterogeneity generated by CRISPR/Cas9 and single-stranded DNA oligonucleotides
Scientific Reports (2016)
-
Combinatorial gene editing in mammalian cells using ssODNs and TALENs
Scientific Reports (2014)
-
Progress and prospects: oligonucleotide-directed gene modification in mouse embryonic stem cells: a route to therapeutic application
Gene Therapy (2011)
-
Simultaneous targeted exchange of two nucleotides by single-stranded oligonucleotides clusters within a region of about fourteen nucleotides
BMC Molecular Biology (2008)
-
CD8 T cell memory development: CD4 T cell help is appreciated
Immunologic Research (2007)