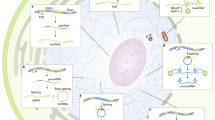
Key Points
-
Although long non-coding RNAs (lncRNAs) and mRNAs share many common features, several types of lncRNAs are distinguished from mRNAs by unique features of biogenesis, form and function.
-
lncRNAs exhibit more highly specific expression patterns than mRNAs.
-
Many lncRNAs undergo special processing events, such as backspliced circularization, 5′- and 3′-bookending by processed small nucleolar RNAs (snoRNAs), and cleavage by RNase P.
-
lncRNAs are more enriched in the nucleus than the cytoplasm relative to mRNAs, and although cytoplasmic lncRNAs associate with the ribosome, few are productively translated.
-
Certain classes of lncRNAs are preferentially subject to degradation by nonsense-mediated decay and the nuclear exosome, and the elongation of divergent ncRNA transcripts is co-transcriptionally terminated by premature polyadenylation.
-
lncRNAs are uniquely capable of cis action on the genome and chromatin. This feature of lncRNAs enables such biological phenomena as gene imprinting, dosage compensation of sex chromosomes, transcriptional enhancement, chromosome looping and antisense regulation.
Abstract
Long non-coding RNAs (lncRNAs) are a diverse class of RNAs that engage in numerous biological processes across every branch of life. Although initially discovered as mRNA-like transcripts that do not encode proteins, recent studies have revealed features of lncRNAs that further distinguish them from mRNAs. In this Review, we describe special events in the lifetimes of lncRNAs — before, during and after transcription — and discuss how these events ultimately shape the unique characteristics and functional roles of lncRNAs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012).
Berretta, J. & Morillon, A. Pervasive transcription constitutes a new level of eukaryotic genome regulation. EMBO Rep. 10, 973–982 (2009).
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 22, 1775–1789 (2012). Comprehensive comparative analysis between mRNAs and lncRNAs in humans.
Carninci, P. et al. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005).
Mattick, J. S., Taft, R. J. & Faulkner, G. J. A global view of genomic information — moving beyond the gene and the master regulator. Trends Genet. 26, 21–28 (2010).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Clark, M. B. et al. Quantitative gene profiling of long noncoding RNAs with targeted RNA sequencing. Nat. Methods 12, 339–342 (2015).
Rinn, J. L. & Chang, H. Y. Genome regulation by long noncoding RNAs. Annu. Rev. Biochem. 81, 145–166 (2012).
Ponting, C. P., Oliver, P. L. & Reik, W. Evolution and functions of long noncoding RNAs. Cell 136, 629–641 (2009).
Flynn, R. A. & Chang, H. Y. Long noncoding RNAs in cell-fate programming and reprogramming. Cell Stem Cell 14, 752–761 (2014).
Batista, P. J. & Chang, H. Y. Long noncoding RNAs: cellular address codes in development and disease. Cell 152, 1298–1307 (2013).
Esteller, M. Non-coding RNAs in human disease. Nat. Rev. Genet. 12, 861–874 (2011).
Ulitsky, I. & Bartel, D. P. lincRNAs: genomics, evolution, and mechanisms. Cell 154, 26–46 (2013).
Geisler, S. & Coller, J. RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. Nat. Rev. Mol. Cell Biol. 14, 699–712 (2013).
Kornienko, A. E., Guenzl, P. M., Barlow, D. P. & Pauler, F. M. Gene regulation by the act of long non-coding RNA transcription. BMC Biol. 11, 59 (2013).
Banfai, B. et al. Long noncoding RNAs are rarely translated in two human cell lines. Genome Res. 22, 1646–1657 (2012).
Gascoigne, D. K. et al. Pinstripe: a suite of programs for integrating transcriptomic and proteomic datasets identifies novel proteins and improves differentiation of protein-coding and non-coding genes. Bioinformatics 28, 3042–3050 (2012).
Cabili, M. N. et al. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 25, 1915–1927 (2011).
Salzman, J., Gawad, C., Wang, P. L., Lacayo, N. & Brown, P. O. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE 7, e30733 (2012).
Lyle, R. et al. The imprinted antisense RNA at the Igf2r locus overlaps but does not imprint Mas1. Nat. Genet. 25, 19–21 (2000).
Hutchinson, J. N. et al. A screen for nuclear transcripts identifies two linked noncoding RNAs associated with SC35 splicing domains. BMC Genomics 8, 39 (2007).
Mercer, T. R., Dinger, M. E., Sunkin, S. M., Mehler, M. F. & Mattick, J. S. Specific expression of long noncoding RNAs in the mouse brain. Proc. Natl Acad. Sci. USA 105, 716–721 (2008).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1774 (2012).
Brunner, A. L. et al. Transcriptional profiling of long non-coding RNAs and novel transcribed regions across a diverse panel of archived human cancers. Genome Biol. 13, R75 (2012).
Gupta, R. A. et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464, 1071–1076 (2010).
Lee, C. S. et al. Transcriptome sequencing in Sezary syndrome identifies Sezary cell and mycosis fungoides-associated lncRNAs and novel transcripts. Blood 120, 3288–3297 (2012).
Hu, G. et al. Expression and regulation of intergenic long noncoding RNAs during T cell development and differentiation. Nat. Immunol. 14, 1190–1198 (2013).
Amaral, P. P. & Mattick, J. S. Noncoding RNA in development. Mamm. Genome 19, 454–492 (2008).
Zheng, G. X., Do, B. T., Webster, D. E., Khavari, P. A. & Chang, H. Y. Dicer-microRNA-Myc circuit promotes transcription of hundreds of long noncoding RNAs. Nat. Struct. Mol. Biol. 21, 585–590 (2014). Demonstrates that lncRNAs can be regulated as a class differently from protein-coding genes.
Alcid, E. A. & Tsukiyama, T. ATP-dependent chromatin remodeling shapes the long noncoding RNA landscape. Genes Dev. 28, 2348–2360 (2014).
Cheung, V. et al. Chromatin- and transcription-related factors repress transcription from within coding regions throughout the Saccharomyces cerevisiae genome. PLoS Biol. 6, e277 (2008).
Marquardt, S. et al. A chromatin-based mechanism for limiting divergent noncoding transcription. Cell 157, 1712–1723 (2014).
Grzechnik, P., Tan-Wong, S. M. & Proudfoot, N. J. Terminate and make a loop: regulation of transcriptional directionality. Trends Biochem. Sci. 39, 319–327 (2014).
Sigova, A. A. et al. Divergent transcription of long noncoding RNA/mRNA gene pairs in embryonic stem cells. Proc. Natl Acad. Sci. USA 110, 2876–2881 (2013).
Seila, A. C. et al. Divergent transcription from active promoters. Science 322, 1849–1851 (2008).
Core, L. J., Waterfall, J. J. & Lis, J. T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008).
Preker, P. et al. RNA exosome depletion reveals transcription upstream of active human promoters. Science 322, 1851–1854 (2008).
Almada, A. E., Wu, X., Kriz, A. J., Burge, C. B. & Sharp, P. A. Promoter directionality is controlled by U1 snRNP and polyadenylation signals. Nature 499, 360–363 (2013).
Wu, X. & Sharp, P. A. Divergent transcription: a driving force for new gene origination? Cell 155, 990–996 (2013).
Guil, S. & Esteller, M. Cis-acting noncoding RNAs: friends and foes. Nat. Struct. Mol. Biol. 19, 1068–1075 (2012). A review of the cis -encoded functions of lncRNAs.
Descostes, N. et al. Tyrosine phosphorylation of RNA polymerase II CTD is associated with antisense promoter transcription and active enhancers in mammalian cells. eLife 3, e02105 (2014).
Hsin, J. P., Li, W., Hoque, M., Tian, B. & Manley, J. L. RNAP II CTD tyrosine 1 performs diverse functions in vertebrate cells. eLife 3, e02112 (2014).
Zhang, Y., Yang, L. & Chen, L. L. Life without a tail: new formats of long noncoding RNAs. Int. J. Biochem. Cell Biol. 54, 338–349 (2014).
Wilusz, J. E. Long noncoding RNAs: re-writing dogmas of RNA processing and stability. Biochim. Biophys. Acta http://dx.doi.org/10.1016/j.bbagrm.2015.06.003 (2015).
Williams, A. S. & Marzluff, W. F. The sequence of the stem and flanking sequences at the 3′ end of histone mRNA are critical determinants for the binding of the stem-loop binding protein. Nucleic Acids Res. 23, 654–662 (1995).
Marvin, M. C. et al. Accumulation of noncoding RNA due to an RNase P defect in Saccharomyces cerevisiae. RNA 17, 1441–1450 (2011).
Ulitsky, I., Shkumatava, A., Jan, C. H., Sive, H. & Bartel, D. P. Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell 147, 1537–1550 (2011).
Wilusz, J. E., Freier, S. M. & Spector, D. L. 3′ end processing of a long nuclear-retained noncoding RNA yields a tRNA-like cytoplasmic RNA. Cell 135, 919–932 (2008).
Wilusz, J. E. et al. A triple helix stabilizes the 3′ ends of long noncoding RNAs that lack poly(A) tails. Genes Dev. 26, 2392–2407 (2012).
Tripathi, V. et al. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol. Cell 39, 925–938 (2010).
Brown, J. A., Valenstein, M. L., Yario, T. A., Tycowski, K. T. & Steitz, J. A. Formation of triple-helical structures by the 3′-end sequences of MALAT1 and MENβ noncoding RNAs. Proc. Natl Acad. Sci. USA 109, 19202–19207 (2012).
Sunwoo, H. et al. MEN ε/β nuclear-retained non-coding RNAs are up-regulated upon muscle differentiation and are essential components of paraspeckles. Genome Res. 19, 347–359 (2009).
Sasaki, Y. T., Ideue, T., Sano, M., Mituyama, T. & Hirose, T. MENɛ/β noncoding RNAs are essential for structural integrity of nuclear paraspeckles. Proc. Natl Acad. Sci. USA 106, 2525–2530 (2009).
Nakagawa, S. et al. The lncRNA Neat1 is required for corpus luteum formation and the establishment of pregnancy in a subpopulation of mice. Development 141, 4618–4627 (2014).
Standaert, L. et al. The long noncoding RNA Neat1 is required for mammary gland development and lactation. RNA 20, 1844–1849 (2014).
Tycowski, K. T., Shu, M. D., Borah, S., Shi, M. & Steitz, J. A. Conservation of a triple-helix-forming RNA stability element in noncoding and genomic RNAs of diverse viruses. Cell Rep. 2, 26–32 (2012).
Brown, J. A. et al. Structural insights into the stabilization of MALAT1 noncoding RNA by a bipartite triple helix. Nat. Struct. Mol. Biol. 21, 633–640 (2014).
Conrad, N. K. & Steitz, J. A. A Kaposi's sarcoma virus RNA element that increases the nuclear abundance of intronless transcripts. EMBO J. 24, 1831–1841 (2005).
Mitton-Fry, R. M., DeGregorio, S. J., Wang, J., Steitz, T. A. & Steitz, J. A. Poly(A) tail recognition by a viral RNA element through assembly of a triple helix. Science 330, 1244–1247 (2010).
Jeck, W. R. et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19, 141–157 (2013).
Guo, J. U., Agarwal, V., Guo, H. & Bartel, D. P. Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 15, 409 (2014).
Chen, C. Y. & Sarnow, P. Initiation of protein synthesis by the eukaryotic translational apparatus on circular RNAs. Science 268, 415–417 (1995).
Wang, Y. & Wang, Z. Efficient backsplicing produces translatable circular mRNAs. RNA 21, 172–179 (2015).
Danan, M., Schwartz, S., Edelheit, S. & Sorek, R. Transcriptome-wide discovery of circular RNAs in Archaea. Nucleic Acids Res. 40, 3131–3142 (2012).
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338 (2013).
Zhang, Y. et al. Circular intronic long noncoding RNAs. Mol. Cell 51, 792–806 (2013).
Li, Z. et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 22, 256–264 (2015).
Yin, Q. F. et al. Long noncoding RNAs with snoRNA ends. Mol. Cell 48, 219–230 (2012).
Cassidy, S. B., Schwartz, S., Miller, J. L. & Driscoll, D. J. Prader–Willi syndrome. Genet. Med. 14, 10–26 (2012).
Dhir, A., Dhir, S., Proudfoot, N. J. & Jopling, C. L. Microprocessor mediates transcriptional termination of long noncoding RNA transcripts hosting microRNAs. Nat. Struct. Mol. Biol. 22, 319–327 (2015).
Mercer, T. R. et al. Expression of distinct RNAs from 3′ untranslated regions. Nucleic Acids Res. 39, 2393–2403 (2011).
Affymetrix & Cold Spring Harbor Laboratory ENCODE Transcriptome Project. Post-transcriptional processing generates a diversity of 5′-modified long and short RNAs. Nature 457, 1028–1032 (2009).
Lai, F., Gardini, A., Zhang, A. & Shiekhattar, R. Integrator mediates the biogenesis of enhancer RNAs. Nature 525, 399–403 (2015).
Tsuiji, H. et al. Competition between a noncoding exon and introns: Gomafu contains tandem UACUAAC repeats and associates with splicing factor-1. Genes Cells 16, 479–490 (2011).
Zhang, B. et al. A novel RNA motif mediates the strict nuclear localization of a long noncoding RNA. Mol. Cell. Biol. 34, 2318–2329 (2014).
Coccia, E. M. et al. Regulation and expression of a growth arrest-specific gene (gas5) during growth, differentiation, and development. Mol. Cell. Biol. 12, 3514–3521 (1992).
Anderson, D. M. et al. A micropeptide encoded by a putative long noncoding RNA regulates muscle performance. Cell 160, 595–606 (2015). Functional characterization of a micropeptide encoded by a transcript previously annotated as a lncRNA, thus illustrating the rich diversity of ORFs and cautioning against computational definitions of lncRNAs.
Kondo, T. et al. Small peptides switch the transcriptional activity of Shavenbaby during Drosophila embryogenesis. Science 329, 336–339 (2010).
Colak, D. et al. Promoter-bound trinucleotide repeat mRNA drives epigenetic silencing in fragile X syndrome. Science 343, 1002–1005 (2014).
Khalil, A. M. et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl Acad. Sci. USA 106, 11667–11672 (2009).
Mondal, T., Rasmussen, M., Pandey, G. K., Isaksson, A. & Kanduri, C. Characterization of the RNA content of chromatin. Genome Res. 20, 899–907 (2010).
Ulveling, D., Francastel, C. & Hube, F. When one is better than two: RNA with dual functions. Biochimie 93, 633–644 (2011).
Kretz, M. et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature 493, 231–235 (2013).
Tay, Y., Rinn, J. & Pandolfi, P. P. The multilayered complexity of ceRNA crosstalk and competition. Nature 505, 344–352 (2014).
Faghihi, M. A. et al. Evidence for natural antisense transcript-mediated inhibition of microRNA function. Genome Biol. 11, R56 (2010).
van Heesch, S. et al. Extensive localization of long noncoding RNAs to the cytosol and mono- and polyribosomal complexes. Genome Biol. 15, R6 (2014).
Cabili, M. N. et al. Localization and abundance analysis of human lncRNAs at single-cell and single-molecule resolution. Genome Biol. 16, 20 (2015). A wide survey of the diverse localization patterns of lncRNAs.
Ingolia, N. T., Lareau, L. F. & Weissman, J. S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011).
Guttman, M. et al. lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature 477, 295–300 (2011).
Pircher, A., Bakowska-Zywicka, K., Schneider, L., Zywicki, M. & Polacek, N. An mRNA-derived noncoding RNA targets and regulates the ribosome. Mol. Cell 54, 147–155 (2014).
Guttman, M., Russell, P., Ingolia, N. T., Weissman, J. S. & Lander, E. S. Ribosome profiling provides evidence that large noncoding RNAs do not encode proteins. Cell 154, 240–251 (2013). Reanalyses the original ribosome-profiling experiments to find that lncRNAs are not productively translated and lack the signature of ribosome release.
Lee, S. et al. Global mapping of translation initiation sites in mammalian cells at single-nucleotide resolution. Proc. Natl Acad. Sci. USA 109, E2424–E2432 (2012).
Ivanov, I. P., Loughran, G. & Atkins, J. F. uORFs with unusual translational start codons autoregulate expression of eukaryotic ornithine decarboxylase homologs. Proc. Natl Acad. Sci. USA 105, 10079–10084 (2008).
Clark, M. B. et al. Genome-wide analysis of long noncoding RNA stability. Genome Res. 22, 885–898 (2012).
Houseley, J. Form and function of eukaryotic unstable non-coding RNAs. Biochem. Soc. Trans. 40, 836–841 (2012).
Davis, C. A. & Ares, M. Jr. Accumulation of unstable promoter-associated transcripts upon loss of the nuclear exosome subunit Rrp6p in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 103, 3262–3267 (2006).
Houseley, J. & Tollervey, D. The many pathways of RNA degradation. Cell 136, 763–776 (2009).
Thompson, D. M. & Parker, R. Cytoplasmic decay of intergenic transcripts in Saccharomyces cerevisiae. Mol. Cell. Biol. 27, 92–101 (2007).
Arigo, J. T., Eyler, D. E., Carroll, K. L. & Corden, J. L. Termination of cryptic unstable transcripts is directed by yeast RNA-binding proteins Nrd1 and Nab3. Mol. Cell 23, 841–851 (2006).
LaCava, J. et al. RNA degradation by the exosome is promoted by a nuclear polyadenylation complex. Cell 121, 713–724 (2005).
Thiebaut, M., Kisseleva-Romanova, E., Rougemaille, M., Boulay, J. & Libri, D. Transcription termination and nuclear degradation of cryptic unstable transcripts: a role for the nrd1-nab3 pathway in genome surveillance. Mol. Cell 23, 853–864 (2006).
Wyers, F. et al. Cryptic pol II transcripts are degraded by a nuclear quality control pathway involving a new poly(A) polymerase. Cell 121, 725–737 (2005).
Camblong, J. et al. Trans-acting antisense RNAs mediate transcriptional gene cosuppression in S. cerevisiae. Genes Dev. 23, 1534–1545 (2009).
Berretta, J., Pinskaya, M. & Morillon, A. A cryptic unstable transcript mediates transcriptional trans-silencing of the Ty1 retrotransposon in S. cerevisiae. Genes Dev. 22, 615–626 (2008).
Marquardt, S., Hazelbaker, D. Z. & Buratowski, S. Distinct RNA degradation pathways and 3′ extensions of yeast non-coding RNA species. Transcription 2, 145–154 (2011).
Xu, Z. et al. Bidirectional promoters generate pervasive transcription in yeast. Nature 457, 1033–1037 (2009).
van Dijk, E. L. et al. XUTs are a class of Xrn1-sensitive antisense regulatory non-coding RNA in yeast. Nature 475, 114–117 (2011).
Grzechnik, P., Gdula, M. R. & Proudfoot, N. J. Pcf11 orchestrates transcription termination pathways in yeast. Genes Dev. 29, 849–861 (2015).
Beaulieu, Y. B., Kleinman, C. L., Landry-Voyer, A. M., Majewski, J. & Bachand, F. Polyadenylation-dependent control of long noncoding RNA expression by the poly(A)-binding protein nuclear 1. PLoS Genet. 8, e1003078 (2012).
Flynn, R. A., Almada, A. E., Zamudio, J. R. & Sharp, P. A. Antisense RNA polymerase II divergent transcripts are P-TEFb dependent and substrates for the RNA exosome. Proc. Natl Acad. Sci. USA 108, 10460–10465 (2011).
Niazi, F. & Valadkhan, S. Computational analysis of functional long noncoding RNAs reveals lack of peptide-coding capacity and parallels with 3′ UTRs. RNA 18, 825–843 (2012).
Tani, H., Torimura, M. & Akimitsu, N. The RNA degradation pathway regulates the function of GAS5 a non-coding RNA in mammalian cells. PLoS ONE 8, e55684 (2013).
Kurihara, Y. et al. Genome-wide suppression of aberrant mRNA-like noncoding RNAs by NMD in Arabidopsis. Proc. Natl Acad. Sci. USA 106, 2453–2458 (2009).
Mendell, J. T., Sharifi, N. A., Meyers, J. L., Martinez-Murillo, F. & Dietz, H. C. Nonsense surveillance regulates expression of diverse classes of mammalian transcripts and mutes genomic noise. Nat. Genet. 36, 1073–1078 (2004).
Mitrovich, Q. M. & Anderson, P. mRNA surveillance of expressed pseudogenes in C. elegans. Curr. Biol. 15, 963–967 (2005).
Ruiz-Orera, J., Messeguer, X., Subirana, J. A. & Alba, M. M. Long non-coding RNAs as a source of new peptides. eLife 3, e03523 (2014).
Shlyueva, D., Stampfel, G. & Stark, A. Transcriptional enhancers: from properties to genome-wide predictions. Nat. Rev. Genet. 15, 272–286 (2014).
De Santa, F. et al. A large fraction of extragenic RNA pol II transcription sites overlap enhancers. PLoS Biol. 8, e1000384 (2010).
Kim, T. K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010).
Natoli, G. & Andrau, J. C. Noncoding transcription at enhancers: general principles and functional models. Annu. Rev. Genet. 46, 1–19 (2012).
Orom, U. A. et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 143, 46–58 (2010).
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Wang, K. C. et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 472, 120–124 (2011).
Quagliata, L. et al. Long noncoding RNA HOTTIP/HOXA13 expression is associated with disease progression and predicts outcome in hepatocellular carcinoma patients. Hepatology 59, 911–923 (2014).
Xiang, J. F. et al. Human colorectal cancer-specific CCAT1-L lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res. 24, 513–531 (2014).
Trimarchi, T. et al. Genome-wide mapping and characterization of Notch-regulated long noncoding RNAs in acute leukemia. Cell 158, 593–606 (2014).
Lai, F. et al. Activating RNAs associate with Mediator to enhance chromatin architecture and transcription. Nature 494, 497–501 (2013).
Schaukowitch, K. et al. Enhancer RNA facilitates NELF release from immediate early genes. Mol. Cell 56, 29–42 (2014).
Di Croce, L. & Helin, K. Transcriptional regulation by Polycomb group proteins. Nat. Struct. Mol. Biol. 20, 1147–1155 (2013).
Herzog, V. A. et al. A strand-specific switch in noncoding transcription switches the function of a Polycomb/Trithorax response element. Nat. Genet. 46, 973–981 (2014).
Barlow, D. P. & Bartolomei, M. S. Genomic imprinting in mammals. Cold Spring Harb. Perspect. Biol. 6 a018382 (2014).
Zhang, Y. & Tycko, B. Monoallelic expression of the human H19 gene. Nat. Genet. 1, 40–44 (1992).
Adalsteinsson, B. T. & Ferguson-Smith, A. C. Epigenetic control of the genome-lessons from genomic imprinting. Genes (Basel) 5, 635–655 (2014).
Royo, H. & Cavaille, J. Non-coding RNAs in imprinted gene clusters. Biol. Cell 100, 149–166 (2008).
Mancini-Dinardo, D., Steele, S. J., Levorse, J. M., Ingram, R. S. & Tilghman, S. M. Elongation of the Kcnq1ot1 transcript is required for genomic imprinting of neighboring genes. Genes Dev. 20, 1268–1282 (2006).
Sleutels, F., Zwart, R. & Barlow, D. P. The non-coding Air RNA is required for silencing autosomal imprinted genes. Nature 415, 810–813 (2002).
Wutz, A. et al. Imprinted expression of the Igf2r gene depends on an intronic CpG island. Nature 389, 745–749 (1997).
Latos, P. A. et al. Airn transcriptional overlap, but not its lncRNA products, induces imprinted Igf2r silencing. Science 338, 1469–1472 (2012).
Pandey, R. R. et al. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol. Cell 32, 232–246 (2008).
Williamson, C. M. et al. Uncoupling antisense-mediated silencing and DNA methylation in the imprinted Gnas cluster. PLoS Genet. 7, e1001347 (2011).
Lin, S. P. et al. Asymmetric regulation of imprinting on the maternal and paternal chromosomes at the Dlk1-Gtl2 imprinted cluster on mouse chromosome 12. Nat. Genet. 35, 97–102 (2003).
Lee, J. T. Epigenetic regulation by long noncoding RNAs. Science 338, 1435–1439 (2012).
Conrad, T. & Akhtar, A. Dosage compensation in Drosophila melanogaster: epigenetic fine-tuning of chromosome-wide transcription. Nat. Rev. Genet. 13, 123–134 (2011).
Grant, J. et al. Rsx is a metatherian RNA with Xist-like properties in X-chromosome inactivation. Nature 487, 254–258 (2012).
Wang, X., Douglas, K. C., Vandeberg, J. L., Clark, A. G. & Samollow, P. B. Chromosome-wide profiling of X-chromosome inactivation and epigenetic states in fetal brain and placenta of the opossum, Monodelphis domestica. Genome Res. 24, 70–83 (2014).
Lee, J. T. & Jaenisch, R. Long-range cis effects of ectopic X-inactivation centres on a mouse autosome. Nature 386, 275–279 (1997).
Pelechano, V. & Steinmetz, L. M. Gene regulation by antisense transcription. Nat. Rev. Genet. 14, 880–893 (2013).
Yap, K. L. et al. Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol. Cell 38, 662–674 (2010).
Sopher, B. L. et al. CTCF regulates ataxin-7 expression through promotion of a convergently transcribed, antisense noncoding RNA. Neuron 70, 1071–1084 (2011).
Hongay, C. F., Grisafi, P. L., Galitski, T. & Fink, G. R. Antisense transcription controls cell fate in Saccharomyces cerevisiae. Cell 127, 735–745 (2006).
Heo, J. B. & Sung, S. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science 331, 76–79 (2011).
Sun, Q., Csorba, T., Skourti-Stathaki, K., Proudfoot, N. J. & Dean, C. R-loop stabilization represses antisense transcription at the Arabidopsis FLC locus. Science 340, 619–621 (2013).
Tian, D., Sun, S. & Lee, J. T. The long noncoding RNA, Jpx, is a molecular switch for X chromosome inactivation. Cell 143, 390–403 (2010).
Sun, B. K., Deaton, A. M. & Lee, J. T. A transient heterochromatic state in Xist preempts X inactivation choice without RNA stabilization. Mol. Cell 21, 617–628 (2006).
Beltran, M. et al. A natural antisense transcript regulates Zeb2/Sip1 gene expression during Snail1-induced epithelial–mesenchymal transition. Genes Dev. 22, 756–769 (2008).
Oberle, I. et al. Instability of a 550-base pair DNA segment and abnormal methylation in fragile X syndrome. Science 252, 1097–1102 (1991).
Blume, S. W., Meng, Z., Shrestha, K., Snyder, R. C. & Emanuel, P. D. The 5′-untranslated RNA of the human dhfr minor transcript alters transcription pre-initiation complex assembly at the major (core) promoter. J. Cell Biochem. 88, 165–180 (2003).
Martianov, I., Ramadass, A., Serra Barros, A., Chow, N. & Akoulitchev, A. Repression of the human dihydrofolate reductase gene by a non-coding interfering transcript. Nature 445, 666–670 (2007).
Jiang, H., Mankodi, A., Swanson, M. S., Moxley, R. T. & Thornton, C. A. Myotonic dystrophy type 1 is associated with nuclear foci of mutant RNA, sequestration of muscleblind proteins and deregulated alternative splicing in neurons. Hum. Mol. Genet. 13, 3079–3088 (2004).
Quinn, J. J. et al. Revealing long noncoding RNA architecture and functions using domain-specific chromatin isolation by RNA purification. Nat. Biotechnol. 32, 933–940 (2014).
Lim, C. K. & Kelley, R. L. Autoregulation of the Drosophila noncoding roX1 RNA gene. PLoS Genet. 8, e1002564 (2012).
Ponjavic, J., Ponting, C. P. & Lunter, G. Functionality or transcriptional noise? Evidence for selection within long noncoding RNAs. Genome Res. 17, 556–565 (2007).
Diederichs, S. The four dimensions of noncoding RNA conservation. Trends Genet. 30, 121–123 (2014).
Hezroni, H. et al. Principles of long noncoding RNA evolution derived from direct comparison of transcriptomes in 17 species. Cell Rep. 11, 1110–1122 (2015).
Mortimer, S. A., Kidwell, M. A. & Doudna, J. A. Insights into RNA structure and function from genome-wide studies. Nat. Rev. Genet. 15, 469–479 (2014).
Byron, K., Cervantes, M. C., Wang, J. T. L., Lin, W. C. & Park, Y. Mining roX1 RNA in Drosophila genomes using covariance models. Int. J. Comp. Biosci. 1, 22–32 (2010).
Smith, M. A., Gesell, T., Stadler, P. F. & Mattick, J. S. Widespread purifying selection on RNA structure in mammals. Nucleic Acids Res. 41, 8220–8236 (2013).
Eddy, S. R. Computational analysis of conserved RNA secondary structure in transcriptomes and genomes. Annu. Rev. Biophys. 43, 433–456 (2014).
Guttman, M. & Rinn, J. L. Modular regulatory principles of large non-coding RNAs. Nature 482, 339–346 (2012).
Somarowthu, S. et al. HOTAIR forms an intricate and modular secondary structure. Mol. Cell 58, 353–361 (2015).
Tsai, M. C. et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 329, 689–693 (2010).
Ilik, I. A. et al. Tandem stem-loops in roX RNAs act together to mediate X chromosome dosage compensation in Drosophila. Mol. Cell 51, 156–173 (2013).
Krueger, B. J., Varzavand, K., Cooper, J. J. & Price, D. H. The mechanism of release of P-TEFb and HEXIM1 from the 7SK snRNP by viral and cellular activators includes a conformational change in 7SK. PLoS ONE 5, e12335 (2010).
Buenrostro, J. D. et al. Quantitative analysis of RNA–protein interactions on a massively parallel array reveals biophysical and evolutionary landscapes. Nat. Biotechnol. 32, 562–568 (2014).
Spitale, R. C. et al. Structural imprints in vivo decode RNA regulatory mechanisms. Nature 519, 486–490 (2015).
Feng, J. et al. The RNA component of human telomerase. Science 269, 1236–1241 (1995).
Nowacki, M. et al. RNA-mediated epigenetic programming of a genome-rearrangement pathway. Nature 451, 153–158 (2008).
Sauvageau, M. et al. Multiple knockout mouse models reveal lincRNAs are required for life and brain development. eLife 2, e01749 (2013). Genetic screen of 18 lncRNA knockouts in mice that demonstrates lncRNA roles in development.
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR–Cas9 complex. Nature 517, 583–588 (2015).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Amrein, H. & Axel, R. Genes expressed in neurons of adult male Drosophila. Cell 88, 459–469 (1997).
Conn, S. J. et al. The RNA binding protein Quaking regulates formation of circRNAs. Cell 160, 1125–1134 (2015).
Chang, T.-C. et al. Genome-wide annotation of microRNA primary transcript structures reveals novel regulatory mechanisms. Genome Res. 25, 1401–1409 (2015).
Sigova, A. A. et al. Transcription factor trapping by RNA in gene regulatory elements. Science 350, 978–981 (2015).
Acknowledgements
The authors thank P. J. Batista, R. A. Flynn and G. X. Zheng for constructive discussions and comments on this Review. Supported by US National Institutes of Health grants, HHMI (H.Y.C.), and the Stanford Bio-X Fellowship (J.J.Q.). The authors apologize to colleagues whose work could not be cited or discussed owing to space constraints.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
FURTHER INFORMATION
Glossary
- Long non-coding RNAs
-
(lncRNAs). RNA transcripts with a length of >200 nt that do not encode proteins.
- Imprinting
-
An epigenetic mechanism of regulating gene expression in an allele-specific manner.
- Cis or trans
-
Long non-coding RNAs can act on their neighbouring genomic environment (in cis) or diffuse to distant sites of action (in trans).
- Transcriptional noise
-
A hypothesis explaining pervasive transcription by which RNA polymerase II randomly initiates transcription throughout the genome.
- Biogenesis
-
The production by organisms of new biological material, such as RNAs, proteins or organelles.
- Open reading frames
-
(ORFs). Continuous stretches of codons that have the potential to encode a protein.
- MicroRNAs
-
(miRNAs). Short (~22 nt) non-coding RNAs that post-transcriptionally silence target RNAs by base pairing.
- Small nucleolar RNAs
-
(snoRNAs). A class of small RNAs that guide the chemical modification of ribosomal RNAs, tRNAs and others.
- Ribosome profiling
-
A biochemical technique for mapping ribosome-associated and translated RNAs.
- Chromatin
-
The compartment of the nucleus that compacts and organizes genomic DNA and regulates gene expression.
- Divergent transcription
-
The production of sense and antisense RNAs from bidirectional promoters.
- U1 snRNP
-
A ribonucleoprotein complex that coordinates the precise splicing of nascent RNA transcripts.
- Enhancers
-
DNA-encoded elements that activate the expression of nearby genes.
- RNase P
-
A ribonucleoprotein complex that cleaves tRNA precursors during their maturation.
- Secondary structure
-
The base-pairing interactions that dictate nucleic acid folds.
- Circular RNAs
-
(circRNAs). Chemically circular RNAs produced by nonsequential exon–exon back-splicing.
- Circular intronic long non-coding RNAs
-
(ciRNAs). Circular, branched intronic RNAs resulting from stabilized introns after canonical splicing.
- Exon-intron circRNAs
-
(EIciRNAs). A class of circular RNAs that retain unspliced introns.
- sno-lncRNA
-
Long non-coding RNAs that are capped on the 5′ and 3′ ends by processed small nucleolar RNAs.
- Bifunctional RNAs
-
RNAs with separable non-coding and protein-coding functions.
- Nonsense-mediated decay
-
(NMD). A genetic pathway for the elimination of RNAs that are defective in protein coding, primarily owing to premature stop codons.
- Exosome
-
A large protein complex that degrades RNA.
- Upstream antisense RNAs
-
(uaRNAs). Non-coding RNAs transcribed in the opposite direction from promoters by divergent transcription.
- Enhancer RNAs
-
(eRNAs). Non-coding RNAs transcribed from enhancers.
- Dosage compensation
-
The epigenetic process of balancing gene expression from sex chromosomes between males and females.
- DNA methylation
-
The chemical modification of cytosine residues on DNA that can be epigenetically inherited.
- CpG island
-
DNA sites of the sequence CG where the cytosine may be methylated in a regulated manner.
- Chromatin isolation by RNA purification
-
(ChIRP). A biochemical technique for mapping the genomic binding sites of an RNA of interest.
- Synteny
-
The physical proximity of genes within a single genomic locus.
- CRISPRi and CRISPRa
-
Sequence-specific interference (i) or activation (a) of gene expression using CRISPR technology. Typically, this involves a nuclease-deficient Cas9-mutant protein fused to transcriptional activator or repressor proteins; this fusion protein is then directed to a target genomic locus by an engineered guide RNA.
Rights and permissions
About this article
Cite this article
Quinn, J., Chang, H. Unique features of long non-coding RNA biogenesis and function. Nat Rev Genet 17, 47–62 (2016). https://doi.org/10.1038/nrg.2015.10
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg.2015.10
This article is cited by
-
Analysis of differentially expressed genes in torn rotator cuff tendon tissues in diabetic patients through RNA-sequencing
BMC Musculoskeletal Disorders (2024)
-
Long non-coding RNA (H19) in patients with spondyloarthritis: association with disease parameters and ultrasonographic findings
Egyptian Rheumatology and Rehabilitation (2024)
-
Lnc-PSMA8-1 activated by GEFT promotes rhabdomyosarcoma progression via upregulation of mTOR expression by sponging miR-144-3p
BMC Cancer (2024)
-
Non-coding RNAs regulating endothelial progenitor cells for venous thrombosis: promising therapy and innovation
Stem Cell Research & Therapy (2024)
-
Non-coding RNAs as potential therapeutic targets for receptor tyrosine kinase signaling in solid tumors: current status and future directions
Cancer Cell International (2024)