Key Points
-
The control of transcription initiation is a primary mechanism for regulating gene expression, especially in governing developmental and cell-specific programmes of gene expression.
-
For much of the past 25 years the model of a largely universal, highly conserved and monolithic core promoter recognition 'pre-initiation complex' composed of general factors that is responsible for initiating transcription in all eukaryotes prevailed.
-
Gene and cell-type-specific transcriptional regulation has traditionally been viewed as the role of sequence-specific enhancer and promoter DNA binding activators/repressors.
-
New evidence suggests that contrary to the conventional textbook models, there may be major diversification and switching of core promoter recognition complexes during cellular differentiation in eukaryotes.
-
Initial evidence for loss of the prototypical transcriptor factor IID (TFIID) and utilization of non-prototypical or orphan TATA-box-binding protein (TBP)-associated factors (TAFs) and TBP-related factors (TRFs) in directing cell-type-specific transcription initiation arose from studies of myogenesis, oogenesis and spermatogenesis.
-
TRF2 and TRF3 are highly expressed in germ cells and are important for germ cell differentiation.
-
There is a growing body of evidence pointing to novel functions and cell-type-specific activities of different subunits and components of the core transcription machinery.
-
Switching of the composition of the core promoter machinery during terminal differentiation of cell types may provide a key mechanism during development of eukaryotes that has evolved to accommodate highly diversified gene expression programmes in multicellular organisms.
Abstract
The eukaryotic core promoter recognition complex was generally thought to play an essential but passive role in the regulation of gene expression. However, recent evidence now indicates that core promoter recognition complexes together with 'non-prototypical' subunits may have a vital regulatory function in driving cell-specific programmes of transcription during development. Furthermore, new roles for components of these complexes have been identified beyond development; for example, in mediating interactions with chromatin and in maintaining active gene expression across cell divisions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Thomas, M. C. & Chiang, C. M. The general transcription machinery and general cofactors. Crit. Rev. Biochem. Mol. Biol. 41, 105–178 (2006).
Naar, A. M., Lemon, B. D. & Tjian, R. Transcriptional coactivator complexes. Annu. Rev. Biochem. 70, 475–501 (2001).
Conaway, R. C. & Conaway, J. W. Transcription initiated by RNA polymerase II and purified transcription factors from liver: transcription factors alpha, beta gamma, and delta promote formation of intermediates in assembly of the functional preinitiation complex. J. Biol. Chem. 265, 7559–7563 (1990).
Biggin, M. D. & Tjian, R. Transcription factors that activate the Ultrabithorax promoter in developmentally staged extracts. Cell 53, 699–711 (1988).
Wasylyk, B., Kedinger, C., Corden, J., Brison, O. & Chambon, P. Specific in vitro initiation of transcription on conalbumin and ovalbumin genes and comparison with adenovirus-2 early and late genes. Nature 285, 367–373 (1980).
Horikoshi, M., Carey, M. F., Kakidani, H. & Roeder, R. G. Mechanism of action of a yeast activator: direct effect of GAL4 derivatives on mammalian TFIID-promoter interactions. Cell 54, 665–669 (1988).
Lin, Y.-S., Carey, M. F., Ptashne, M. & Green, M. R. GAL4 derivatives function alone and synergistically with mammalian activators in vitro. Cell 54, 659–664 (1988).
Carey, M., Lin, Y. S., Green, M. R. & Ptashne, M. A mechanism of synergistic activation of a mammalian gene by GAL4 derivatives. Nature 345, 361–364 (1990).
Wang, W., Gralla, J. D. & Carey, M. The acidic activator GAL4-AH can stimulate polymerase II transcription by promoting assembly of a closed complex requiring TFIID and TFIIA. Genes Dev. 6, 1716–1727 (1992).
Goodrich, J. A., Hoey, T., Thut, C. J., Admon, A. & Tjian, R. Drosophila TAFII40 interacts with both a VP16 activation domain and the basal transcription factor TFIIB. Cell 75, 519–530 (1993).
Juven-Gershon, T., Hsu, J. Y. & Kadonaga, J. T. Perspectives on the RNA polymerase II core promoter. Biochem. Soc. Trans. 34, 1047–1050 (2006).
Juven-Gershon, T. & Kadonaga, J. T. Regulation of gene expression via the core promoter and the basal transcriptional machinery. Dev. Biol. 339, 225–229 (2009).
Liu, W. L. et al. Structures of three distinct activator–TFIID complexes. Genes Dev. 23, 1510–1521 (2009).
Rabenstein, M. D., Zhou, S., Lis, J. T. & Tjian, R. TATA box-binding protein (TBP)-related factor 2 (TRF2), a third member of the TBP family. Proc. Natl Acad. Sci. USA 96, 4791–4796 (1999).
Teichmann, M. et al. Human TATA-binding protein-related factor-2 (hTRF2) stably associates with hTFIIA in HeLa cells. Proc. Natl Acad. Sci. USA 96, 13720–13725 (1999).
Moore, P. A. et al. A human TATA binding protein-related protein with altered DNA binding specificity inhibits transcription from multiple promoters and activators. Mol. Cell. Biol. 19, 7610–7620 (1999).
Maldonado, E. Transcriptional functions of a new mammalian TATA-binding protein-related factor. J. Biol. Chem. 274, 12963–12966 (1999).
Ohbayashi, T., Makino, Y. & Tamura, T. A. Identification of a mouse TBP-like protein (TLP) distantly related to the Drosophila TBP-related factor. Nucleic Acids Res. 27, 750–755 (1999).
Zhang, D., Penttila, T. L., Morris, P. L., Teichmann, M. & Roeder, R. G. Spermiogenesis deficiency in mice lacking the Trf2 gene. Science 292, 1153–1155 (2001).
Martianov, I. et al. Late arrest of spermiogenesis and germ cell apoptosis in mice lacking the TBP-like TLF/TRF2 gene. Mol. Cell 7, 509–515 (2001).
Martianov, I. et al. Distinct functions of TBP and TLF/TRF2 during spermatogenesis: requirement of TLF for heterochromatic chromocenter formation in haploid round spermatids. Development 129, 945–955 (2002).
Kaltenbach, L., Horner, M. A., Rothman, J. H. & Mango, S. E. The TBP-like factor CeTLF is required to activate RNA polymerase II transcription during C. elegans embryogenesis. Mol. Cell 6, 705–713 (2000).
Veenstra, G. J., Weeks, D. L. & Wolffe, A. P. Distinct roles for TBP and TBP-like factor in early embryonic gene transcription in Xenopus. Science 290, 2312–2315 (2000). Showed that TRF2 is highly expressed in X. laevis oocytes and embryos and that knock down of TRF2 stops embryos from developing past the mid-blastula stage. This provided early evidence that non-prototypical core promoter recognition factors function in embryogenesis.
Jacobi, U. G. et al. TBP paralogs accommodate metazoan- and vertebrate-specific developmental gene regulation. EMBO J. 26, 3900–3909 (2007).
Kopytova, D. V. et al. Two isoforms of Drosophila TRF2 are involved in embryonic development, premeiotic chromatin condensation, and proper differentiation of germ cells of both sexes. Mol. Cell. Biol. 26, 7492–7505 (2006).
Bashirullah, A., Lam, G., Yin, V. P. & Thummel, C. S. dTrf2 is required for transcriptional and developmental responses to ecdysone during Drosophila metamorphosis. Dev. Dyn. 236, 3173–3179 (2007).
Persengiev, S. P. et al. TRF3, a TATA-box-binding protein-related factor, is vertebrate-specific and widely expressed. Proc. Natl Acad. Sci. USA 100, 14887–14891 (2003). Identified TRF3 in the human, mouse and frog genomes, and that TRF3 protein is expressed in many human and mouse tissues and cell lines. The identification of TRF3 set the stage for many other studies of this protein and its function in development.
Bartfai, R. et al. TBP2, a vertebrate-specific member of the TBP family, is required in embryonic development of zebrafish. Curr. Biol. 14, 593–598 (2004).
Jallow, Z., Jacobi, U. G., Weeks, D. L., Dawid, I. B. & Veenstra, G. J. Specialized and redundant roles of TBP and a vertebrate-specific TBP paralog in embryonic gene regulation in Xenopus. Proc. Natl Acad. Sci. USA 101, 13525–13530 (2004).
Xiao, L., Kim, M. & DeJong, J. Developmental and cell type-specific regulation of core promoter transcription factors in germ cells of frogs and mice. Gene Expr. Patterns 6, 409–419 (2006).
Gazdag, E., Rajkovic, A., Torres-Padilla, M. E. & Tora, L. Analysis of TATA-binding protein 2 (TBP2) and TBP expression suggests different roles for the two proteins in regulation of gene expression during oogenesis and early mouse development. Reproduction 134, 51–62 (2007).
Gazdag, E. et al. TBP2 is essential for germ cell development by regulating transcription and chromatin condensation in the oocyte. Genes Dev. 23, 2210–2223 (2009).
Hiller, M. A., Lin, T. Y., Wood, C. & Fuller, M. T. Developmental regulation of transcription by a tissue-specific TAF homolog. Genes Dev. 15, 1021–1030 (2001). Showed that the protein encoded by the Drosophila cannonball gene is a homologue of TAF5. Cannonball is expressed in Drosophila testes where it controls expression of a series of genes in spermatocytes. After these studies were published additional testes-specific TAFs were discovered that also function in spermatogenesis.
Hiller, M. et al. Testis-specific TAF homologs collaborate to control a tissue-specific transcription program. Development 131, 5297–5308 (2004).
Chen, X., Hiller, M., Sancak, Y. & Fuller, M. T. Tissue-specific TAFs counteract Polycomb to turn on terminal differentiation. Science 310, 869–872 (2005).
Dikstein, R., Zhou, S. & Tjian, R. Human TAFII105 is a cell type-specific TFIID subunit related to hTAFII130. Cell 87, 137–146 (1996). This paper documented the discovery of the first cell-type-specific TAF, TAF II 105 (now known as TAF4b), which is similar in sequence to TAF4. TAF4b is associated with TFIID in a B-cell line, but not in other cell lines. This paper raised awareness of the possibility of cell-type-specific general transcription factors; subsequently, many other cell-type-specific TAFs were discovered.
Freiman, R. N. et al. Requirement of tissue-selective TBP-associated factor TAFII105 in ovarian development. Science 293, 2084–2087 (2001). Showed that knock out of Taf4b in mice is required for development of the ovary. Taf4b is expressed in granulosa cells and is necessary for expression of ovarian-specific genes during folliculogenesis.
Voronina, E. et al. Ovarian granulosa cell survival and proliferation requires the gonad-selective TFIID subunit TAF4b. Dev. Biol. 303, 715–726 (2007).
Lovasco, L. A. et al. Accelerated ovarian aging in the absence of the transcription regulator TAF4B in mice. Biol. Reprod. 82, 23–34 (2010).
Geles, K. G. et al. Cell-type-selective induction of c-jun by TAF4b directs ovarian-specific transcription networks. Proc. Natl Acad. Sci. USA 103, 2594–2599 (2006).
Liu, W. L. et al. Structural changes in TAF4b-TFIID correlate with promoter selectivity. Mol. Cell 29, 81–91 (2008).
Falender, A. E. et al. Maintenance of spermatogenesis requires TAF4b, a gonad-specific subunit of TFIID. Genes Dev. 19, 794–803 (2005).
Pointud, J. C. et al. The intracellular localisation of TAF7L, a paralogue of transcription factor TFIID subunit TAF7, is developmentally regulated during male germ-cell differentiation. J. Cell Sci. 116, 1847–1858 (2003).
Cheng, Y. et al. Abnormal sperm in mice lacking the Taf7l gene. Mol. Cell. Biol. 27, 2582–2589 (2007).
Hart, D. O., Raha, T., Lawson, N. D. & Green, M. R. Initiation of zebrafish haematopoiesis by the TATA-box-binding protein-related factor Trf3. Nature 450, 1082–1085 (2007). Showed that knock out of trf3 in zebrafish is required for haematopoiesis and that Trf3 binds to the promoter of the mespa gene and facilitates its expression. In the absence of mespa , haematopoiesis does not occur.
Hart, D. O., Santra, M. K., Raha, T. & Green, M. R. Selective interaction between Trf3 and Taf3 required for early development and hematopoiesis. Dev. Dyn. 238, 2540–2549 (2009).
Deato, M. D. & Tjian, R. Switching of the core transcription machinery during myogenesis. Genes Dev. 21, 2137–2149 (2007). Showed that an unprecedented exchange of core promoter recognition machinery occurs when myoblasts differentiate into myotubes. Specifically, multiple subunits of TFIID decrease, whereas levels of TAF3 remained constant. TAF3 binds TRF3 in myotubes and the two factors associate with the core promoter of the myogenin gene where they direct its transcription, thereby facilitating myogenesis.
Deato, M. D. et al. MyoD targets TAF3/TRF3 to activate myogenin transcription. Mol. Cell 32, 96–105 (2008).
Akhtar, W. & Veenstra, G. J. TBP2 is a substitute for TBP in Xenopus oocyte transcription. BMC Biol. 7, 45 (2009).
Guermah, M., Ge, K., Chiang, C. M. & Roeder, R. G. The TBN protein, which is essential for early embryonic mouse development, is an inducible TAFII implicated in adipogenesis. Mol. Cell 12, 991–1001 (2003).
Mohan, W. S. J., Scheer, E., Wendling, O., Metzger, D. & Tora, L. TAF10 (TAFII30) is necessary for TFIID stability and early embryogenesis in mice. Mol. Cell. Biol. 23, 4307–4318 (2003).
Tatarakis, A. et al. Dominant and redundant functions of TFIID involved in the regulation of hepatic genes. Mol. Cell 31, 531–543 (2008).
Hochheimer, A., Zhou, S., Zheng, S., Holmes, M. C. & Tjian, R. TRF2 associates with DREF and directs promoter-selective gene expression in Drosophila. Nature 420, 439–445 (2002).
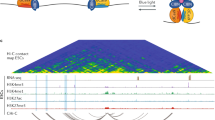
Isogai, Y., Keles, S., Prestel, M., Hochheimer, A. & Tjian, R. Transcription of histone gene cluster by differential core-promoter factors. Genes Dev. 21, 2936–2949 (2007). Showed that TRF2 directs the transcription of histone H1 genes, but not adjacent core histone genes, in the Drosophila histone gene cluster. TRF2 binds the promoter of the TATA-less histone H1 gene, whereas TBP binds the promoters of the core histone genes. Genome-wide studies revealed that TRF2 widely binds TATA-less genes and drives their transcription.
Holmgren, P., Johansson, T., Lambertsson, A. & Rasmuson, B. Content of histone H1 and histone phosphorylation in relation to the higher order structures of chromatin in Drosophila. Chromosoma 93, 123–131 (1985).
Ruddell, A. & Jacobs-Lorena, M. Biphasic pattern of histone gene expression during Drosophila oogenesis. Proc. Natl Acad. Sci. USA 82, 3316–3319 (1985).
Bernstein, B. E. et al. Genomic maps and comparative analysis of histone modifications in human and mouse. Cell 120, 169–181 (2005).
Heintzman, N. D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nature Genet. 39, 311–318 (2007).
Vermeulen, M. et al. Selective anchoring of TFIID to nucleosomes by trimethylation of histone H3 lysine 4. Cell 131, 58–69 (2007). Demonstrated that TAF3 binds a specific post-translationally modified form of histone H3 (H3K4me3) and recruits TFIID to chromatin containing this modification. Importantly, H3K4me3 is a known mark for transcriptionally active genes. These studies prompt us to reconsider the definition of a core promoter to include not just DNA elements, but also the specific post-translational modifications on histones associated with core promoter regions.
Millar, C. B. & Grunstein, M. Genome-wide patterns of histone modifications in yeast. Nature Rev. Mol. Cell Biol. 7, 657–666 (2006).
Jacobson, R. H., Ladurner, A. G., King, D. S. & Tjian, R. Structure and function of a human TAFII250 double bromodomain module. Science 288, 1422–1425 (2000).
Michelotti, E. F., Sanford, S. & Levens, D. Marking of active genes on mitotic chromosomes. Nature 388, 895–899 (1997).
Segil, N., Guermah, M., Hoffmann, A., Roeder, R. G. & Heintz, N. Mitotic regulation of TFIID: inhibition of activator-dependent transcription and changes in subcellular localization. Genes Dev. 10, 2389–2400 (1996).
Christova, R. & Oelgeschlager, T. Association of human TFIID-promoter complexes with silenced mitotic chromatin in vivo. Nature Cell Biol. 4, 79–82 (2002).
Chen, D., Hinkley, C. S., Henry, R. W. & Huang, S. TBP dynamics in living human cells: constitutive association of TBP with mitotic chromosomes. Mol. Biol. Cell 13, 276–284 (2002).
Kieffer-Kwon, P., Martianov, I. & Davidson, I. Cell-specific nucleolar localization of TBP-related factor 2. Mol. Biol. Cell 15, 4356–4368 (2004).
Xing, H., Vanderford, N. L. & Sarge, K. D. The TBP–PP2A mitotic complex bookmarks genes by preventing condensin action. Nature Cell Biol. 10, 1318–1323 (2008). Showed that TBP functions in mitotic bookmarking by binding PP2A and the CAP-G subunit of condensin at core promoters of genes that are active prior to mitosis. This assembly causes dephosphorylation of CAP-G, inactivation of condensin and blocks the compaction of chromatin containing these core promoters. Once cells exit mitosis, transcription of the bookmarked genes is reactivated.
Kimmins, S., Kotaja, N., Davidson, I. & Sassone-Corsi, P. Testis-specific transcription mechanisms promoting male germ-cell differentiation. Reproduction 128, 5–12 (2004).
Acknowledgements
We apologize to those whose relevant research we were unable to discuss owing to space limitations. J.A.G. was funded by Grant R01 GM55235 from the National Institute of General Medical Sciences and R.T. was partly funded by R37 CA25417 from the National Cancer Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
FURTHER INFORMATION
Nature Reviews Genetics series on Modes of transcriptional regulation
Glossary
- RNA polymerase II
-
(RNAPII). The enzyme that synthesizes mRNA in eukaryotic cells. RNAPII is composed of 12 protein subunits (RPB1–RPB12). The binding of RNAPII to promoters and the initiation of transcription requires many general transcription factors (TFs), for example, TFIIA, TFIIB, TFIID, TFIIE, TFIIF and TFIIH.
- Core promoter
-
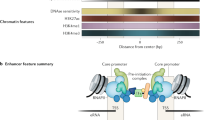
The region of a gene to which RNA polymerase II and the general transcription factors (TFs) bind to initiate transcription. Core promoters span from approximately 40 base pairs upstream to 40 base pairs downstream of the transcription start site and are composed of DNA elements to which subunits of TFIID (or TFIIB) bind.
- Pre-initiation complex
-
The assembly of general transcription factors and RNA polymerase II on core promoter DNA. This complex, which can be assembled in the absence of nucleotide triphosphates in vitro, is competent to initiate transcription in the presence of nucleotides.
- Transcription factor IID
-
(TFIID). A transcription factor for RNA polymerase II that binds core promoters. The TFIID complex is composed of the TATA-box-binding protein (TBP) and 13 or 14 TBP-associated factors.
- TATA-box-binding protein
-
(TBP). The central subunit of transcription factor IID. TBP binds TATA boxes found in the core promoters of some eukaryotic mRNA genes.
- TBP-associated factor
-
(TAF). All subunits of the transcription factor IID (TFIID) complex other than TATA-box-binding protein (TBP) are TAFs. There are 13 or 14 TAFs in the prototypical TFIID complex. There are also several proteins with sequence similarity to the prototypical TAFs, which are referred to as non-prototypical TAFs.
- Core promoter recognition factor
-
A protein or multi-subunit complex that binds with sequence specificity to core promoter elements. The prototypical core promoter recognition factor for mRNA genes in eukaryotes is transcription factor IID, subunits of which recognize multiple core promoter elements.
- TBP-related factor
-
(TRF). A protein that is highly related in sequence to TATA-box-binding protein (TBP). Two TRF proteins are discussed in this Review: TRF2 (also known as TLF, TLP, TRP or TBPL1) and TRF3 (also known as TBPL2).
- Mitotic bookmarking
-
The process by which genes that are active before mitosis are marked such that transcription begins again at these genes when cells exit mitosis and enter the G1 phase of the cell cycle.
Rights and permissions
About this article
Cite this article
Goodrich, J., Tjian, R. Unexpected roles for core promoter recognition factors in cell-type-specific transcription and gene regulation. Nat Rev Genet 11, 549–558 (2010). https://doi.org/10.1038/nrg2847
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg2847
This article is cited by
-
The late-evolving salmon and trout join the GnRH1 club
Histochemistry and Cell Biology (2023)
-
50+ years of eukaryotic transcription: an expanding universe of factors and mechanisms
Nature Structural & Molecular Biology (2019)
-
TAF1, associated with intellectual disability in humans, is essential for embryogenesis and regulates neurodevelopmental processes in zebrafish
Scientific Reports (2019)
-
Chromatin dependencies in cancer and inflammation
Nature Reviews Molecular Cell Biology (2018)
-
BIM and NOXA are mitochondrial effectors of TAF6δ-driven apoptosis
Cell Death & Disease (2018)