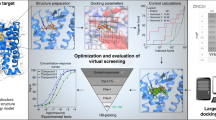
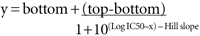Key Points
-
In recent years, the emphasis in high-throughput screening has shifted from increasing screening capacity to improving the quality of the resulting data.
-
This article discusses the issues that are crucial to successful screening, including:
-
What makes a good assay?
-
Types of screen and readouts
-
Compound selection by filtering screening libraries to remove compounds with undesirable properties, and focusing libraries to target types to enhance hit rates.
Abstract
The basic goal of small-molecule screening is the identification of chemically 'interesting' starting points for elaboration towards a drug. A number of innovative approaches for pursuing this goal have evolved, and the right approach is dictated by the target class being pursued and the capabilities of the organization involved. A recent trend in high-throughput screening has been to place less emphasis on the number of data points that can be produced, and to focus instead on the quality of the data obtained. Several computational and technological advances have aided in the selection of compounds for screening and widened the variety of assay formats available for screening. The effect on the efficiency of the screening process is discussed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kapur, R. Fluorescence imaging and engineered biosensors functional and activity-based sensing using high content screening. Ann. NY Acad. Sci. 961, 196–197 (2002).
Bleicher, K. H. Chemogenomics: bridging a drug discovery gap. Curr. Med. Chem. 23, 2077–2084 (2002).
Namchuk, M. Finding the molecules to fuel chemogenomics. Targets 1, 125–129 (2002).
Zhang, J. H., Chung, T. D. & Oldenburg, K. R. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J. Biomol. Screen. 4, 27–32 (1999). Describes a widely used metric for defining the quality of high-throughput screening assays.
Miller, A. Discovery of cytokine mimics using cell-based systems. Drug Discov. Today 5, 77–83 (2000).
Selvin, P. R. Principles and biophysical applications of lanthanide-based probes. Annu. Rev. Biophys. Biomol. Struct. 31, 275–302 (2002).
Lipinski, C. A., Lombardo, F., Dominy, B. W. & Feeney, P. J. Experimental and computational approaches to estimate solubility and permeablity in drug discovery. Adv. Drug Deliv. Rev. 23, 3–25 (1997). Introduces Lipinski's rule-of-five.
Ghose, A. K., Viswanadhan, V. N. & Wendelowski, J. J. A knowledge-based approach in designing combinatorial or medicinal chemistry libraries for drug discovery. 1. A qualitative characterization of known drug databases. J. Comb. Chem. 1, 55–67 (1999).
Oprea, T. I. Property distribution of drug-related chemical databases. J. Comput. Aided Mol. Des. 14, 251–264. (2000).
Walters, W. P., Ajay, A. & Murcko, M. A. Recognizing molecules with drug-like properties. Curr. Opin. Chem. Biol. 3, 384–387 (1999).
Egan, W. J. & Lauri, G. Prediction of intestinal permeability. Adv. Drug Deliv. Rev. 54, 273–289 (2002).
Clark, D. E. & Grootenhuis, P. D. J. Progress in computational methods for the predicition of ADMET properties. Curr. Opin. Drug. Discov. Devel. 5, 382–390 (2002).
Hann, M. M., Leach, A. R. & Harper G. Molecular complexity and its impact on the probability of finding leads for drug discovery. J. Chem. Inf. Comput. Sci. 41, 856–864 (2001).
Oprea, T. I. Current trends in lead discovery: are we looking for the appropriate properties? J. Comput. Aided Mol. Des. 16, 325–34 (2002).
Muegge, I., Heald, S. L. & Brittelli, D. Simple selection criteria for drug-like chemical matter. J. Med. Chem. 44, 1841–1846 (2001).
Mitchell, T. M. Machine Learning (McGraw-Hill, New York, NY, 1997).
Zupan, J. & Gasteiger, J. Neural Networks for Chemists (VCH, Weinheim, 1993).
Ajay, A., Walters, W. P. & Murcko, M. A. Can we learn to distinguish between 'drug-like' and 'nondrug-like' molecules? J. Med. Chem. 41, 3314–3324 (1998).
Sadowski, J. & Kubinyi, H. A scoring scheme for discriminating between drugs and nondrugs. J. Med. Chem. 41, 3325–3329 (1998). References 18 and 19 describe the application of machine-learning techniques to the prediction of drug-likeness.
Aronov, A. M., Munagala, N. R., Kuntz, I. D. & Wang, C. C. Virtual screening of combinatorial libraries accross a gene family: in search of inhibitors of Giardia lamblia guanine phosphorribosyltransferase. Antimicrob. Agents Chemother. 45, 2571–2576 (2001).
Diller, D. J. & Merz, K. M. High throughput docking for library design and library prioritization. Proteins 43, 113–124 (2001).
Stahura, F. L., Xue, L., Gooden, J. W. & Bajorath, J. Molecular scaffold-based design and comparison of combinatorial libraries focused on the ATP-binding site of protein kinases. J. Mol. Graph. Model. 17, 1–9 (1999).
Balakin, K. V. et al. Property-based design of GPCR-targeted library. J. Chem. Inf. Comput. Sci. 42, 1332–1342 (2002).
Muegge, I. & Rarey, M. in Reviews in Computational Chemistry Volume 17 (eds Lipkowitz, K. B. & Boyd, D. B.) 1–60 (Wiley–VCH, New York, 2001).
Charifson, P. S., Corkery, J. J., Murcko, M. A. & Walters, W. P. Consensus scoring: a method for obtaining improved hit rates from docking databases of three-dimensional structures into proteins. J. Med. Chem. 42, 5100–5109 (1999).
Ajay, A. & Murcko, M. A. Computational methods to predict binding free energy in ligand-receptor complexes. J. Med. Chem. 38, 4953–4967 (1995).
Hull, R. D. et al. Chemical similarity searches using latent semantic structural indexing (LaSSI) and comparison to TOPOSIM. J. Med. Chem. 44, 1185–1191 (2001).
Kearsley, S. K, & Sheridan, R. P. Why do we need so many chemical similarity search methods? Drug Discov. Today 7, 903–911 (2002).
Fox, T. et al. A single amino acid substitution makes ERK2 susceptible to pyridinyl imidazole inhibitors of p38 MAP kinase. Protein Sci. 7, 2249–2255 (1998).
Stills, M. et al. Comparison of assay technologies for tyrosine kinase assay generates different results in high throughput screening. J. Biomol. Screen. 7, 191–214 (2002).
Lipinski, C. A. Drug-like properties and the causes of poor solubility and poor permeability. J Pharmacol. Toxicol. Methods 44, 235–249 (2000).
Segal, I. Enzyme Kinetics: Behavior and Analysis of Rapid Equilibrium and Steady-State Enzyme Systems. 465–504 (John Wiley and Sons, New York, 1993).
Motalsky, H. Analysing Data with Graphpad Prism, 173–174 (1998).
Author information
Authors and Affiliations
Corresponding author
Related links
Glossary
- HILL SLOPE
-
In the program Prism, the curves are fitted to the equation:
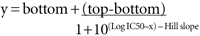
Rights and permissions
About this article
Cite this article
Walters, W., Namchuk, M. Designing screens: how to make your hits a hit. Nat Rev Drug Discov 2, 259–266 (2003). https://doi.org/10.1038/nrd1063
Issue Date:
DOI: https://doi.org/10.1038/nrd1063
This article is cited by
-
Accelerating inhibitor discovery for deubiquitinating enzymes
Nature Communications (2023)
-
Discovery of new PKN2 inhibitory chemotypes via QSAR-guided selection of docking-based pharmacophores
Molecular Diversity (2023)
-
Discovery of novel antagonists targeting the DNA binding domain of androgen receptor by integrated docking-based virtual screening and bioassays
Acta Pharmacologica Sinica (2022)
-
Predictive validity in drug discovery: what it is, why it matters and how to improve it
Nature Reviews Drug Discovery (2022)
-
A practical guide to large-scale docking
Nature Protocols (2021)