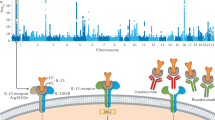
Abstract
An assessment of the number of molecular targets that represent an opportunity for therapeutic intervention is crucial to the development of post-genomic research strategies within the pharmaceutical industry. Now that we know the size of the human genome, it is interesting to consider just how many molecular targets this opportunity represents. We start from the position that we understand the properties that are required for a good drug, and therefore must be able to understand what makes a good drug target.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Lipinski, C., Lombardo, F., Dominy, B. & Feeney, P. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 23, 3–25 (1997).
Drews, J. Genomic sciences and the medicine of tomorrow. Nature Biotechnol. 14, 1516–1518 (1996).
Drews, J. & Ryser, S. Classic drug targets. Nature Biotechnol. 15, 1318–1319 (1997).
Drews, J. Drug discovery: a historical perspective. Science 287, 1960–1964 (2000).
Bailey, D., Zanders, E. & Dean, P. The end of the beginning for genomic medicine. Nature Biotechnol. 19, 207–209 (2001).
Apweiler, R. et al. The InterPro database, an integrated documentation resource for protein families, domains and functional sites. Nucleic Acids Res. 29, 37–40 (2001).
Proteome Analysis Database [online], (version analysed with release date 29 Oct 01) 〈http://www.ebi.ac.uk/proteome/〉 (2001).
Lander, E. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Tweeddale, H., Notley-McRobb, L. & Ferenci, T. Effect of slow growth on metabolism of Escherichia coli, as revealed by global metabolite pool ('metabolome') analysis. J. Bacteriol. 180, 5109–5119 (1998).
Fiehn, O. et al. Metabolite profiling for plant functional genomics. Nature Biotechnol. 18, 1157–1161 (2000).
Raamsdonk, L. et al. A functional genomics strategy that uses metabolome data to reveal the phenotype of silent mutations. Nature Biotechnol. 19, 45–50 (2001).
Claverie, J.-M. What if there are only 30,000 human genes? Science 291, 1255–1257 (2001).
Walke, D. W. et al. In vivo drug target discovery: identifying the best targets from the genome. Curr. Opin. Biotechnol. 12, 626–631 (2001)
Lehman Brothers. The Fruits of Genomics (Lehman Brothers, 2001).
Rubin, G.et al Comparative genomics of the eukaryotes. Science 287, 2204–2215 (2000).
Acknowledgements
We are indebted to J. P. Overington (Inpharmatica, London), A. Alex and L. Beeley for their contributions to the ideas on the physico-chemical limits for protein-binding sites. We also thank R. W. Spencer and C. Lipinski (Pfizer, Groton, Connecticut, USA) for much stimulating discussion.
Author information
Authors and Affiliations
Corresponding author
Related links
Rights and permissions
About this article
Cite this article
Hopkins, A., Groom, C. The druggable genome. Nat Rev Drug Discov 1, 727–730 (2002). https://doi.org/10.1038/nrd892
Issue Date:
DOI: https://doi.org/10.1038/nrd892
This article is cited by
-
MHESMMR: a multilevel model for predicting the regulation of miRNAs expression by small molecules
BMC Bioinformatics (2024)
-
Delivery of nucleic acids using nanomaterials
Molecular Biomedicine (2023)
-
The effects of methylphenidate and atomoxetine on Drosophila brain at single-cell resolution and potential drug repurposing for ADHD treatment
Molecular Psychiatry (2023)
-
A simple method for developing lysine targeted covalent protein reagents
Nature Communications (2023)
-
Molecular chameleons in drug discovery
Nature Reviews Chemistry (2023)