Key Points
-
Whether metastasis tends to occur early or late in tumour development remains controversial, and whether metastases descend directly from the primary tumour or give rise to each other is unclear
-
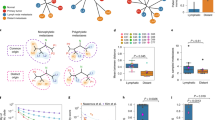
In the linear progression model, metastatic precursors leave the primary tumour at late stages of disease, after clonal evolution has given rise to a cell with metastatic ability; consequently, primary tumours and metastases are genetically closely related
-
The parallel progression model, on the other hand, assumes that metastasis occurs in early stages of carcinogenesis, and that metastases and primary tumour evolve independently, resulting in genetic disparity between them
-
Comparative genomics studies in different cancer types illustrate a variety of possible progression trajectories for systemic disease, but analysis of more patients is needed to arrive at generalizable conclusions
-
Problems complicating the interpretation of comparative genetics data include the unknown contributions of the tissue-specific background mutational burden, self-seeding and tumour-cell dormancy, and the extensive heterogeneity of primary tumours
-
A variety of experimental approaches beyond next-generation DNA sequencing are available for lineage tracing in human cancer
Abstract
In cancer, much uncertainty remains regarding the origins of metastatic disease. Models of metastatic progression offer competing views on when dissemination occurs (at an early or late stage of tumour development), whether metastases at different sites arise independently and directly from the primary tumour or give rise to each other, and whether dynamic cell exchange occurs between synchronously growing lesions. Although it is probable that many routes can lead to the establishment of systemic disease, clinical observations suggest that distinct modes of metastasis might prevail in different tumour types. Gaining a more-comprehensive understanding of the evolutionary processes that underlie metastasis is not only relevant from a basic biological perspective, but also has profound clinical implications. The 'tree of life' of metastatic cancer contains answers to many outstanding questions about the development of systemic disease, but has only been reconstructed in a limited number of patients. Here we review available data on the phylogenetic relationships between primary solid tumours and their metastases, and examine to what degree they support different models of metastatic progression. We provide a description of experimental methods for lineage tracing in human cancer, ranging from broad DNA-sequencing approaches to more-targeted techniques, and discuss their respective benefits and caveats. Finally, we propose future research questions in the area of cancer phylogenetics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Holohan, C., Van Schaeybroeck, S., Longley, D. B. & Johnston, P. G. Cancer drug resistance: an evolving paradigm. Nat. Rev. Cancer 13, 714–726 (2013).
Navin, N. E. & Hicks, J. Tracing the tumor lineage. Mol. Oncol. 4, 267–283 (2010).
Swanton, C. Intratumor heterogeneity: evolution through space and time. Cancer Res. 72, 4875–4882 (2012).
Marusyk, A., Almendro, V. & Polyak, K. Intra-tumour heterogeneity: a looking glass for cancer? Nat. Rev. Cancer 12, 323–334 (2012).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Sankin, A. et al. The impact of genetic heterogeneity on biomarker development in kidney cancer assessed by multiregional sampling. Cancer Med. http://dx.doi.org/10.1002/cam4.293 (2014).
Simmons, C. et al. Does confirmatory tumor biopsy alter the management of breast cancer patients with distant metastases? Ann. Oncol. 20, 1499–1504 (2009).
Niikura, N. et al. Latest biopsy approach for suspected metastases in patients with breast cancer. Nat. Rev. Clin. Oncol. 10, 711–719 (2013).
Jain, R. K. Normalizing tumor microenvironment to treat cancer: bench to bedside to biomarkers. J. Clin. Oncol. 31, 2205–2218 (2013).
Weiss, L. Concepts of metastasis. Cancer Metastasis Rev. 19, 219–234 (2000).
Klein, C. A. Parallel progression of primary tumours and metastases. Nat. Rev. Cancer 9, 302–312 (2009).
Klein, C. A. Framework models of tumor dormancy from patient-derived observations. Curr. Opin. Genet. Dev. 21, 42–49 (2011).
Hess, K. R., Pusztai, L., Buzdar, A. U. & Hortobagyi, G. N. Estrogen receptors and distinct patterns of breast cancer relapse. Breast Cancer Res. Treat. 78, 105–118 (2003).
Tsai, M. S. et al. Clinicopathological features and prognosis in resectable synchronous and metachronous colorectal liver metastasis. Ann. Surg. Oncol. 14, 786–794 (2007).
Bragado, P., Sosa, M. S., Keely, P., Condeelis, J. & Aguirre-Ghiso, J. A. Microenvironments dictating tumor cell dormancy. Recent Results Cancer Res. 195, 25–39 (2012).
Talmadge, J. E. & Fidler, I. J. AACR centennial series: the biology of cancer metastasis: historical perspective. Cancer Res. 70, 5649–5669 (2010).
Duda, D. G. et al. Malignant cells facilitate lung metastasis by bringing their own soil. Proc. Natl Acad. Sci. USA 107, 21677–21682 (2010).
Teng, M. W., Swann, J. B., Koebel, C. M., Schreiber, R. D. & Smyth, M. J. Immune-mediated dormancy: an equilibrium with cancer. J. Leukoc. Biol. 84, 988–993 (2008).
McAllister, S. S. & Weinberg, R. A. The tumour-induced systemic environment as a critical regulator of cancer progression and metastasis. Nat. Cell Biol. 16, 717–727 (2014).
Cairns, J. Mutation selection and the natural history of cancer. Nature 255, 197–200 (1975).
Bross, I. D., Viadana, E. & Pickren, J. Do generalized metastases occur directly from the primary? J. Chronic Dis. 28, 149–159 (1975).
Weinberg, R. A. Mechanisms of malignant progression. Carcinogenesis 29, 1092–1095 (2008).
Disibio, G. & French, S. W. Metastatic patterns of cancers: results from a large autopsy study. Arch. Pathol. Lab. Med. 132, 931–939 (2008).
Hellman, S. Karnofsky Memorial Lecture. Natural history of small breast cancers. J. Clin. Oncol. 12, 2229–2234 (1994).
Fisher, B. Laboratory and clinical research in breast cancer—a personal adventure: the David, A. Karnofsky memorial lecture. Cancer Res. 40, 3863–3874 (1980).
Giuliano, A. E. et al. Axillary dissection vs no axillary dissection in women with invasive breast cancer and sentinel node metastasisa randomized clinical trial. JAMA 305, 569–575 (2011).
Giuliano, A. E. et al. Locoregional recurrence after sentinel lymph node dissection with or without axillary dissection in patients with sentinel lymph node metastases: the American College of Surgeons Oncology Group Z0011 randomized trial. Ann. Surg. 252, 426–432 (2010).
Klein, C. A. Selection and adaptation during metastatic cancer progression. Nature 501, 365–372 (2013).
Husemann, Y. et al. Systemic spread is an early step in breast cancer. Cancer Cell 13, 58–68 (2008).
Kim, M.-Y. et al. Tumor self-seeding by circulating cancer cells. Cell 139, 1315–1326 (2009).
Aguirre-Ghiso, J. A. Models, mechanisms and clinical evidence for cancer dormancy. Nat. Rev. Cancer 7, 834–846 (2007).
Jones, S. et al. Comparative lesion sequencing provides insights into tumor evolution. Proc. Natl Acad. Sci. USA 105, 4283–4288 (2008).
Yachida, S. et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 467, 1114–1117 (2010).
Meng, S. et al. Circulating tumor cells in patients with breast cancer dormancy. Clin. Cancer Res. 10, 8152–8162 (2004).
Nagrath, S. et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 450, 1235–1239 (2007).
Riethdorf, S., Wikman, H. & Pantel, K. Review: Biological relevance of disseminated tumor cells in cancer patients. Int. J. Cancer 123, 1991–2006 (2008).
Cristofanilli, M. et al. Circulating tumor cells, disease progression, and survival in metastatic breast cancer. N. Engl. J. Med. 351, 781–791 (2004).
Danila, D. C. et al. Circulating tumor cell number and prognosis in progressive castration-resistant prostate cancer. Clin. Cancer Res. 13, 7053–7058 (2007).
Krebs, M. G. et al. Evaluation and prognostic significance of circulating tumor cells in patients with non-small-cell lung cancer. J. Clin. Oncol. 29, 1556–1563 (2011).
Crowley, E., Di Nicolantonio, F., Loupakis, F. & Bardelli, A. Liquid biopsy: monitoring cancer-genetics in the blood. Nat. Rev. Clin. Oncol. 10, 472–484 (2013).
Alix-Panabières, C. & Pantel, K. Circulating tumor cells: liquid biopsy of cancer. Clin. Chem. 59, 110–118 (2013).
Schmidt-Kittler, O. et al. From latent disseminated cells to overt metastasis: genetic analysis of systemic breast cancer progression. Proc. Natl Acad. Sci. USA 100, 7737–7742 (2003).
Weckermann, D. et al. Perioperative activation of disseminated tumor cells in bone marrow of patients with prostate cancer. J. Clin. Oncol. 27, 1549–1556 (2009).
Schardt, J. A. et al. Genomic analysis of single cytokeratin-positive cells from bone marrow reveals early mutational events in breast cancer. Cancer Cell 8, 227–239 (2005).
Klein, C. A. et al. Comparative genomic hybridization, loss of heterozygosity, and DNA sequence analysis of single cells. Proc. Natl Acad. Sci. USA 96, 4494–4499 (1999).
Klein, C. A. et al. Genetic heterogeneity of single disseminated tumour cells in minimal residual cancer. Lancet 360, 683–689 (2002).
Wang, Y. et al. Clonal evolution in breast cancer revealed by single nucleus genome sequencing. Nature 512, 155–160 (2014).
Xu, X. et al. Single-cell exome sequencing reveals single-nucleotide mutation characteristics of a kidney tumor. Cell 148, 886–895 (2012).
Navin, N. et al. Tumour evolution inferred by single-cell sequencing. Nature 472, 90–94 (2011).
Brannon, A. et al. Comparative sequencing analysis reveals high genomic concordance between matched primary and metastatic colorectal cancer lesions. Genome Biol. 15, 454 (2014).
Campbell, P. J. et al. The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature 467, 1109–1113 (2010).
Liu, W. et al. Copy number analysis indicates monoclonal origin of lethal metastatic prostate cancer. Nat. Med. 15, 559–565 (2009).
Letouzé, E., Allory, Y., Bollet, M. A., Radvanyi, F. & Guyon, F. Analysis of the copy number profiles of several tumor samples from the same patient reveals the successive steps in tumorigenesis. Genome Biol. 11, R76 (2010).
Haffner, M. C. et al. Tracking the clonal origin of lethal prostate cancer. J. Clin. Invest. 123, 4918–4922 (2013).
Ding, L. et al. Genome remodelling in a basal-like breast cancer metastasis and xenograft. Nature 464, 999–1005 (2010).
Aceto, N. et al. Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell 158, 1110–1122 (2014).
Shah, S. P. et al. Mutational evolution in a lobular breast tumour profiled at single nucleotide resolution. Nature 461, 809–813 (2009).
Wu, X. et al. Clonal selection drives genetic divergence of metastatic medulloblastoma. Nature 482, 529–533 (2012).
Vermaat, J. S. et al. Primary colorectal cancers and their subsequent hepatic metastases are genetically different: implications for selection of patients for targeted treatment. Clin. Cancer Res. 18, 688–699 (2012).
Kuukasjarvi, T. et al. Genetic heterogeneity and clonal evolution underlying development of asynchronous metastasis in human breast cancer. Cancer Res. 57, 1597–1604 (1997).
Schmid, K. et al. EGFR/KRAS/BRAF mutations in primary lung adenocarcinomas and corresponding locoregional lymph node metastases. Clin. Cancer Res. 15, 4554–4560 (2009).
Colombino, M. et al. BRAF/NRAS mutation frequencies among primary tumors and metastases in patients with melanoma. J. Clin. Oncol. 30, 2522–2529 (2012).
Baldus, S. E. et al. Prevalence and heterogeneity of KRAS, BRAF, and PIK3CA mutations in primary colorectal adenocarcinomas and their corresponding metastases. Clin. Cancer Res. 16, 790–799 (2010).
Stoecklein, N. H. & Klein, C. A. Genetic disparity between primary tumours, disseminated tumour cells, and manifest metastasis. Int. J. Cancer 126, 589–598 (2010).
Tomasetti, C., Vogelstein, B. & Parmigiani, G. Half or more of the somatic mutations in cancers of self-renewing tissues originate prior to tumor initiation. Proc. Natl Acad. Sci. USA 110, 1999–2004 (2013).
Welch, J. S. et al. The origin and evolution of mutations in acute myeloid leukemia. Cell 150, 264–278 (2012).
van't Veer, L. J. et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 415, 530–536 (2002).
Comen, E., Norton, L. & Massagué, J. Clinical implications of cancer self-seeding. Nat. Rev. Clin. Oncol. 8, 369–377 (2011).
Jain, R. K., Martin, J. D. & Stylianopoulos, T. The role of mechanical forces in tumor growth and therapy. Annu. Rev. Biomed. Eng. 16, 321–346 (2014).
Geurts, T. W. et al. Pulmonary squamous cell carcinoma following head and neck squamous cell carcinoma: metastasis or second primary? Clin. Cancer Res. 11, 6608–6614 (2005).
Sequist, L. V. et al. Genotypic and histological evolution of lung cancers acquiring resistance to EGFR inhibitors. Sci. Transl. Med. 3, 75ra26 (2011).
Davoli, T. et al. Cumulative haploinsufficiency and triplosensitivity drive aneuploidy patterns and shape the cancer genome. Cell 155, 948–962 (2013).
Beroukhim, R. et al. Assessing the significance of chromosomal aberrations in cancer: methodology and application to glioma. Proc. Natl Acad. Sci. USA 104, 20007–20012 (2007).
Mani, R. S. et al. Induced chromosomal proximity and gene fusions in prostate cancer. Science 326, 1230 (2009).
Behjati, S. et al. Genome sequencing of normal cells reveals developmental lineages and mutational processes. Nature 513, 422–425 (2014).
Going, J. J., Abd El-Monem, H. M. & Craft, J. A. Clonal origins of human breast cancer. J. Pathol. 194, 406–412 (2001).
Katona, T. M. et al. Genetically heterogeneous and clonally unrelated metastases may arise in patients with cutaneous melanoma. Am. J. Surg. Pathol. 31, 1029–1037 (2007).
Novelli, M. et al. X-inactivation patch size in human female tissue confounds the assessment of tumor clonality. Proc. Natl Acad. Sci. USA 100, 3311–3314 (2003).
Jäger, N. et al. Hypermutation of the inactive X chromosome is a frequent event in cancer. Cell 155, 567–581 (2013).
Shibata, D. & Tavaré, S. Counting divisions in a human somatic cell tree: how, what and why? Cell Cycle 5, 610–614 (2006).
Shibata, D., Navidi, W., Salovaara, R., Li, Z. H. & Aaltonen, L. A. Somatic microsatellite mutations as molecular tumor clocks. Nat. Med. 2, 676–681 (1996).
Nicolas, P., Kim, K. M., Shibata, D. & Tavare, S. The stem cell population of the human colon crypt: analysis via methylation patterns. PLoS Comput. Biol. 3, e28 (2007).
Yatabe, Y., Tavare, S. & Shibata, D. Investigating stem cells in human colon by using methylation patterns. Proc. Natl Acad. Sci. USA 98, 10839–10844 (2001).
Woo, Y. J., Siegmund, K. D., Tavare, S. & Shibata, D. Older individuals appear to acquire mitotically older colorectal cancers. J. Pathol. 217, 483–488 (2009).
Siegmund, K. D., Marjoram, P., Woo, Y. J., Tavare, S. & Shibata, D. Inferring clonal expansion and cancer stem cell dynamics from DNA methylation patterns in colorectal cancers. Proc. Natl Acad. Sci. USA 106, 4828–4833 (2009).
Siegmund, K. D., Marjoram, P., Tavare, S. & Shibata, D. High DNA methylation pattern intratumoral diversity implies weak selection in many human colorectal cancers. PLoS ONE 6, e21657 (2011).
Baylin, S. B. & Jones, P. A. A decade of exploring the cancer epigenome—biological and translational implications. Nat. Rev. Cancer 11, 726–734 (2011).
Feinberg, A. P. & Vogelstein, B. Hypomethylation distinguishes genes of some human cancers from their normal counterparts. Nature 301, 89–92 (1983).
Ellegren, H. Microsatellites: simple sequences with complex evolution. Nat. Rev. Genet. 5, 435–445 (2004).
Strand, M., Prolla, T. A., Liskay, R. M. & Petes, T. D. Destabilization of tracts of simple repetitive DNA in yeast by mutations affecting DNA mismatch repair. Nature 365, 274–276 (1993).
Lynch, M. Evolution of the mutation rate. Trends Genet. 26, 345–352 (2010).
Boyer, J. C. et al. Sequence dependent instability of mononucleotide microsatellites in cultured mismatch repair proficient and deficient mammalian cells. Hum. Mol. Genet. 11, 707–713 (2002).
Ionov, Y., Peinado, M. A., Malkhosyan, S., Shibata, D. & Perucho, M. Ubiquitous somatic mutations in simple repeated sequences reveal a new mechanism for colonic carcinogenesis. Nature 363, 558–561 (1993).
Aaltonen, L. A. et al. Clues to the pathogenesis of familial colorectal cancer. Science 260, 812–816 (1993).
Samowitz, W. S. et al. Microsatellite instability in sporadic colon cancer is associated with an improved prognosis at the population level. Cancer Epidemiol. Biomarkers Prev. 10, 917–923 (2001).
Bonadona, V. et al. Cancer risks associated with germline mutations in MLH1, MSH2, and MSH6 genes in Lynch syndrome. JAMA 305, 2304–2310 (2011).
Fishel, R. et al. The human mutator gene homolog MSH2 and its association with hereditary nonpolyposis colon cancer. Cell 75, 1027–1038 (1993).
Tsao, J. L. et al. Genetic reconstruction of individual colorectal tumor histories. Proc. Natl Acad. Sci. USA 97, 1236–1241 (2000).
Wasserstrom, A. et al. Reconstruction of cell lineage trees in mice. PLoS ONE 3, e1939 (2008).
Reizel, Y. et al. Colon stem cell and crypt dynamics exposed by cell lineage reconstruction. PLoS Genet. 7, e1002192 (2011).
Reizel, Y. et al. Cell lineage analysis of the mammalian female germline. PLoS Genet. 8, e1002477 (2012).
Frumkin, D. et al. Cell lineage analysis of a mouse tumor. Cancer Res. 68, 5924–5931 (2008).
Shlush, L. I. et al. Cell lineage analysis of acute leukemia relapse uncovers the role of replication-rate heterogeneity and microsatellite instability. Blood 120, 603–612 (2012).
Salipante, S. J. & Horwitz, M. S. Phylogenetic fate mapping. Proc. Natl Acad. Sci. USA 103, 5448–5453 (2006).
Salipante, S. J., Kas, A., McMonagle, E. & Horwitz, M. S. Phylogenetic analysis of developmental and postnatal mouse cell lineages. Evol. Dev. 12, 84–94 (2010).
Salipante, S. J., Thompson, J. M. & Horwitz, M. S. Phylogenetic fate mapping: theoretical and experimental studies applied to the development of mouse fibroblasts. Genetics 178, 967–977 (2008).
Zhou, W. et al. Use of somatic mutations to quantify random contributions to mouse development. BMC Genomics 14, 39 (2013).
Salk, J. J. & Horwitz, M. S. Passenger mutations as a marker of clonal cell lineages in emerging neoplasia. Semin. Cancer Biol. 20, 294–303 (2010).
Salk, J. J. et al. Clonal expansions in ulcerative colitis identify patients with neoplasia. Proc. Natl Acad. Sci. USA 106, 20871–20876 (2009).
Naxerova, K. et al. Hypermutable DNA chronicles the evolution of human colon cancer. Proc. Natl Acad. Sci. USA 111, E1889–E1898 (2014).
de Bruin, E. C. et al. Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 346, 251–256 (2014).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Johnson, B. E. et al. Mutational analysis reveals the origin and therapy-driven evolution of recurrent glioma. Science 343, 189–193 (2014).
Maley, C. C. et al. Genetic clonal diversity predicts progression to esophageal adenocarcinoma. Nat. Genet. 38, 468–473 (2006).
Burrell, R. A. & Swanton, C. Tumour heterogeneity and the evolution of polyclonal drug resistance. Mol. Oncol. 8, 1095–1111 (2014).
Shibata, D. Cancer. Heterogeneity and tumor history. Science 336, 304–305 (2012).
Slack, N. H. & Bross, I. D. The influence of site of metastasis on tumour growth and response to chemotherapy. Br. J. Cancer 32, 78–86 (1975).
Kodack, D. P. et al. Combined targeting of HER2 and VEGFR2 for effective treatment of HER2-amplified breast cancer brain metastases. Proc. Natl Acad. Sci. USA 109, E3119–E3127 (2012).
Merlo, L. M. F., Pepper, J. W., Reid, B. J. & Maley, C. C. Cancer as an evolutionary and ecological process. Nat. Rev. Cancer 6, 924–935 (2006).
Acknowledgements
The work of the authors is supported in part by funding from the US Department of Defence (grant W81XWH-11-1-0146 to K.N.), the Breast Cancer Innovator Award (grant W81XWH-10-1-0016 to R.K.J.), the US National Institutes of Health (grants P01CA080124 and R01CA163815 to R.K.J.), the Proton Beam/Federal Share Program (R.K.J.), and the National Foundation for Cancer Research (R.K.J.).
Author information
Authors and Affiliations
Contributions
K.N. and R.K.J. made substantial contributions to discussion of content and review/editing of the manuscript before submission. K.N. researched the data for the article and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Naxerova, K., Jain, R. Using tumour phylogenetics to identify the roots of metastasis in humans. Nat Rev Clin Oncol 12, 258–272 (2015). https://doi.org/10.1038/nrclinonc.2014.238
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrclinonc.2014.238
This article is cited by
-
Biology, vulnerabilities and clinical applications of circulating tumour cells
Nature Reviews Cancer (2023)
-
Unraveling the Drivers of Tumorigenesis in the Context of Evolution: Theoretical Models and Bioinformatics Tools
Journal of Molecular Evolution (2023)
-
Is it time for redefining oligometastatic disease? Analysis of lung metastases CT in ten tumor types
Discover Oncology (2023)
-
Targeting SOST using a small-molecule compound retards breast cancer bone metastasis
Molecular Cancer (2022)
-
Characterization of evolution trajectory and immune profiling of brain metastasis in lung adenocarcinoma
npj Precision Oncology (2021)