Abstract
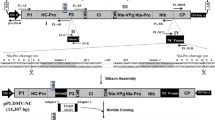
Virus-based transgene expression systems have become particularly valuable for recombinant protein production in plants. The dual-module in-plant activation (INPACT) expression platform consists of a uniquely designed split-gene cassette incorporating the cis replication elements of Tobacco yellow dwarf geminivirus (TYDV) and an ethanol-inducible activation cassette encoding the TYDV Rep and RepA replication-associated proteins. The INPACT system is essentially tailored for recombinant protein production in stably transformed plants and provides both inducible and high-level transient transgene expression with the potential to be adapted to diverse crop species. The construction of a novel split-gene cassette, the inducible nature of the system and the ability to amplify transgene expression via rolling-circle replication differentiates this system from other DNA- and RNA-based virus vector systems used for stable or transient recombinant protein production in plants. Here we provide a detailed protocol describing the design and construction of a split-gene INPACT cassette, and we highlight factors that may influence optimal activation and amplification of gene expression in transgenic plants. By using Nicotiana tabacum, the protocol takes 6–9 months to complete, and recombinant proteins expressed using INPACT can accumulate to up to 10% of the leaf total soluble protein.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Sainsbury, F., Thuenemann, E.C. & Lomonossoff, G.P. pEAQ: versatile expression vectors for easy and quick transient expression of heterologous proteins in plants. Plant Biotechnol. J. 7, 682–693 (2009).
Marillonnet, S. et al. Systemic Agrobacterium tumefaciens-mediated transfection of viral replicons for efficient transient expression in plants. Nat. Biotechnol. 23, 718–723 (2005).
Zhang, X.R. & Mason, H. Bean yellow dwarf virus replicons for high-level transgene expression in transgenic plants and cell cultures. Biotechnol. Bioeng. 93, 271–279 (2006).
Mori, M. et al. Inducible high-level mRNA amplification system by viral replicase in transgenic plants. Plant J. 27, 79–86 (2001).
Werner, S. et al. High-level recombinant protein expression in transgenic plants by using a double-inducible viral vector. Proc. Natl. Acad. Sci. USA 108, 14061–14066 (2011).
Lai, H.F. et al. Robust production of virus-like particles and monoclonal antibodies with geminiviral replicon vectors in lettuce. Plant Biotechnol. J. 10, 95–104 (2012).
Toth, R.L., Pogue, G.P. & Chapman, S. Improvement of the movement and host range properties of a plant virus vector through DNA shuffling. Plant J. 30, 593–600 (2002).
Lechtenberg, B. et al. Neither inverted repeat T-DNA configurations nor arrangements of tandemly repeated transgenes are sufficient to trigger transgene silencing. Plant J. 34, 507–517 (2003).
Jorgensen, R.A. et al. Chalcone synthase cosuppression phenotypes in petunia flowers: Comparison of sense vs. antisense constructs and single-copy vs complex T-DNA sequences. Plant Mol. Biol. 31, 957–973 (1996).
Muskens, M.W.M. et al. Role of inverted DNA repeats in transcriptional and post-transcriptional gene silencing. Plant Mol. Biol. 43, 243–260 (2000).
Lindbo, J.A. et al. Induction of a highly specific antiviral state in transgenic plants: implications for regulation of gene expression and virus resistance. Plant Cell 5, 1749–1759 (1993).
Schubert, D. et al. Silencing in Arabidopsis T-DNA transformants: the predominant role of a gene-specific RNA sensing mechanism versus position effects. Plant Cell 16, 2561–2572 (2004).
Vaucheret, H. et al. Transgene-induced gene silencing in plants. Plant J. 16, 651–659 (1998).
Dugdale, B. et al. In Plant Activation: an inducible, hyperexpression platform for recombinant protein production in plants. Plant Cell 25, 2429–2443 (2013).
Caddick, M.X. et al. An ethanol-inducible gene switch for plants used to manipulate carbon metabolism. Nat. Biotechnol. 16, 177–180 (1998).
Stenger, D.C., Davis, K.R. & Bisaro, D.M. Limited replication of tomato golden mosaic virus DNA in explants of nonhost species. Mol. Plant Microbe Interact. 5, 525–527 (1992).
Teng, K.L. et al. Involvement of C4 protein of beet severe curly top virus (family Geminiviridae) in virus movement. PLoS ONE 5, e11280 (2010).
Kinkema, M. et al. An improved chemically inducible gene switch that functions in the monocotyledonous plant sugar cane. Plant Mol. Biol. 84, 443–454 (2013).
Wright, E.A. et al. Splicing features in maize streak virus virion- and complementary-sense gene expression. Plant J. 12, 1285–1297 (1997).
Boulton, M.I. Functions and interactions of Mastrevirus gene products. Physiol. Mol. Plant Pathol. 60, 243–255 (2002).
Horvath, G.V. et al. Prediction of functional regions of the maize streak virus replication-associated proteins by protein-protein interaction analysis. Plant Mol. Biol. 38, 699–712 (1998).
Liu, L. et al. Bean yellow dwarf virus RepA, but not Rep, binds to maize retinoblastoma protein, and the virus tolerates mutations in the consensus binding motif. Virology 256, 270–279 (1999).
Xie, Q., Suarezlopez, P. & Gutierrez, C. Identification and analysis of a retinoblastoma binding motif in the replication protein of a plant DNA virus: requirement for efficient viral DNA replication. EMBO J. 14, 4073–4082 (1995).
Salter, M.G. et al. Characterisation of the ethanol-inducible alc gene expression system for transgenic plants. Plant J. 16, 127–132 (1998).
Deveaux, Y. et al. The ethanol switch: a tool for tissue-specific gene induction during plant development. Plant J. 36, 918–930 (2003).
Laufs, P. et al. Separable roles of UFO during floral development revealed by conditional restoration of gene function. Development 130, 785–796 (2003).
Roslan, H.A. et al. Characterization of the ethanol-inducible alc gene-expression system in Arabidopsis thaliana. Plant J. 28, 225–235 (2001).
Filichkin, S.A. et al. Alcohol-inducible gene expression in transgenic Populus. Plant Cell Rep. 25, 660–667 (2006).
Garoosi, G.A. et al. Characterization of the ethanol-inducible alc gene expression system in tomato. J. Exp. Botany 56, 1635–1642 (2005).
Sweetman, J.P. et al. Ethanol vapor is an efficient inducer of the alc gene expression system in model and crop plant species. Plant Physiol. 129, 943–948 (2002).
Moore, I., Samalova, M. & Kurup, S. Transactivated and chemically inducible gene expression in plants. Plant J. 45, 651–683 (2006).
Jia, H. et al. Combination of the ALCR/alcA ethanol switch and GAL4/VP16-UAS enhancer trap system enables spatial and temporal control of transgene expression in Arabidopsis. Plant Biotechnol. J. 5, 477–482 (2007).
Outchkourov, N.S. et al. The promoter-terminator of chrysanthemunm rbcS1 directs very high expression levels in plants. Planta 216, 1003–1012 (2003).
Kay, R. et al. Duplication of CaMV 35S promoter sequences creates a strong enhancer for plant genes. Science 236, 1299–1302 (1987).
Dey, N. & Maiti, I.B. Structure and promoter/leader deletion analysis of mirabilis mosaic virus (MMV) full-length transcript promoter in transgenic plants. Plant Mol. Biol. 40, 771–782 (1999).
Dey, N. & Maiti, I.B. Further characterization and expression analysis of mirabilis mosaic caulimovirus (MMV) full-length transcript promoter with single and double enhancer domains in transgenic plants. Transgenics 3, 61 (1999).
Maiti, I.B. et al. Promoter/leader deletion analysis and plant expression vectors with the figwort mosaic virus (FMV) full length transcript (FLt) promoter containing single or double enhancer domains. Transgenic Res. 6, 143–156 (1997).
Maiti, I.B. & Shepherd, R.J. Isolation and expression analysis of peanut chlorotic streak caulimovirus (PCISV) full-length transcript (FLt) promoter in transgenic plants. Biochem. Biophys. Res. Commun. 244, 440–444 (1998).
Gallie, D.R. et al. A comparison of eukaryotic viral 5′-leader sequences as enhancers of mRNA expression in vivo. Nucleic Acids Res. 15, 8693–8711 (1987).
Datla, R.S.S. et al. Improved high-level constitutive foreign gene expression in plants using an AMV RNA4 untranslated leader sequence. Plant Sci. 94, 139–149 (1993).
Sainsbury, F. & Lomonossoff, G.P. Extremely high-level and rapid transient protein production in plants without the use of viral replication. Plant Physiol. 148, 1212–1218 (2008).
Donson, J. et al. A putative primer for second-strand DNA synthesis of maize streak virus is virion-associated. EMBO J. 3, 3069–3073 (1984).
Hayes, R.J. et al. Priming of complementary DNA synthesis in vitro by small DNA molecules tightly bound to virion DNA of wheat dwarf virus. J. General Virol. 69, 1345–1350 (1988).
Morris, B.A.M. et al. The nucleotide sequence of the infectious cloned DNA component of tobacco yellow dwarf virus reveals features of geminiviruses infecting monocotyledonous plants. Virology 187, 633–642 (1992).
Maclean, J. et al. Optimization of human papillomavirus type 16 (HPV-16) L1 expression in plants: comparison of the suitability of different HPV-16 L1 gene variants and different cell-compartment localization. J. General Virol. 88, 1460–1469 (2007).
Rose, A.B. The effect of intron location on intron-mediated enhancement of gene expression in Arabidopsis. Plant J. 40, 744–751 (2004).
Goodall, G.J. & Filipowicz, W. The AU-rich sequences present in the introns of plant nuclear pre-mRNAs are required for splicing. Cell 58, 473–483 (1989).
Baynton, C.E. et al. U-rich tracts enhance 3′ splice site recognition in plant nuclei. Plant J. 10, 703–711 (1996).
Simpson, C.G. et al. Dual functionality of a plant U-rich intronic sequence element. Plant J. 37, 82–91 (2004).
Wang, B.B. & Brendel, V. Genomewide comparative analysis of alternative splicing in plants. Proc. Natl. Acad. Sci. USA 103, 7175–7180 (2006).
Luo, Z. & Chen, Z. Improperly terminated, unpolyadenylated mRNA of sense transgenes is targeted by RDR6-mediated RNA silencing in Arabidopsis. Plant Cell 19, 943–958 (2007).
Southern, E. Southern blotting. Nat. Protoc. 1, 518–525 (2006).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular Cloning: A labratory Manual (Cold Spring Harbor Labratory Press, 1989).
Acknowledgements
We thank the Australian Research Council for funding this research.
Author information
Authors and Affiliations
Contributions
B.D. and C.L.M. designed and performed experiments, analyzed data and wrote the manuscript. M.K. and T.A.J. designed and performed experiments and analyzed data. J.L.D. and R.M.H. designed experiments, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Data
Syntron and LIR sequence (PDF 96 kb)
Rights and permissions
About this article
Cite this article
Dugdale, B., Mortimer, C., Kato, M. et al. Design and construction of an in-plant activation cassette for transgene expression and recombinant protein production in plants. Nat Protoc 9, 1010–1027 (2014). https://doi.org/10.1038/nprot.2014.068
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.068
This article is cited by
-
Screening of a multi-virus resistant RNAi construct in cowpea through transient vacuum infiltration method
VirusDisease (2019)
-
Cotton Leaf Curl Multan Betasatellite DNA as a Tool to Deliver and Express the Human B-Cell Lymphoma 2 (Bcl-2) Gene in Plants
Molecular Biotechnology (2016)
-
Current status of viral expression systems in plants and perspectives for oral vaccines development
Plant Molecular Biology (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.