Abstract
The pig has emerged as an important large animal model in biomedical and pharmaceutical research. We describe a protocol for high-efficiency germline transgenesis and sustained transgene expression in pigs by using the Sleeping Beauty (SB) transposon system. The protocol is based on co-injection of a plasmid encoding the SB100X hyperactive transposase, together with a second plasmid carrying a transgene flanked by binding sites for the transposase, into the cytoplasm of porcine zygotes. The transposase mediates excision of the transgene cassette from the plasmid vector and its permanent insertion into the genome to produce stable transgenic animals. This method compares favorably in terms of both efficiency and reliable transgene expression to classic pronuclear microinjection or somatic cell nuclear transfer (SCNT), and it offers comparable efficacies to lentiviral approaches, without limitations on vector design, issues of transgene silencing and the toxicity and biosafety concerns of working with viral vectors. Microinjection of the vectors into zygotes and transfer of the embryos to recipient animals can be performed in 1 d; generation of germline-transgenic lines by using this protocol takes ∼1 year.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Kuzmuk, K.N. & Schook, L.B. Pigs as a model for biomedical sciences. in The Genetics of the Pig, Vol. 2 (eds. Rothschild, M.F. & Ruvinsky, A.) 426–444 (CAB International, 2011).
Kues, W.A. & Niemann, H. Advances in farm animal transgenesis. Prev. Vet. Med. 102, 146–156 (2011).
Whyte, J.J. & Prather, R.S. Genetic modifications of pigs for medicine and agriculture. Mol. Reprod. Dev. 78, 879–891 (2011).
Cibelli, J. et al. Strategies for improving animal models for regenerative medicine. Cell Stem Cell 12, 271–274 (2013).
Garrels, W., Ivics, Z. & Kues, W.A. Precision genetic engineering in large mammals. Trends Biotechnol. 30, 386–393 (2012).
Petters, R.M. et al. Genetically engineered large animal model for studying cone photoreceptor survival and degeneration in retinitis pigmentosa. Nat. Biotechnol. 15, 965–970 (1997).
Rogers, C.S. et al. Disruption of the CFTR gene produces a model of cystic fibrosis in newborn pigs. Science 321, 1837–1841 (2008).
Kragh, P.M. et al. Hemizygous minipigs produced by random gene insertion and handmade cloning express the Alzheimer's disease-causing dominant mutation APPsw. Transgenic Res. 18, 545–558 (2009).
Yang, D. et al. Expression of Huntington's disease protein results in apoptotic neurons in the brains of cloned transgenic pigs. Hum. Mol. Genet. 19, 3983–3994 (2010).
Flisikowska, T. et al. A porcine model of familial adenomatous polyposis. Gastroenterology 143, 1173–1175 e1-7 (2012).
Suzuki, S. et al. Il2rg gene-targeted severe combined immunodeficiency pigs. Cell Stem Cell 10, 753–758 (2012).
Yeom, H.J. et al. Generation and characterization of human heme oxygenase-1 transgenic pigs. PLoS ONE 7, e46646 (2012).
Hofmann, A. et al. Epigenetic regulation of lentiviral transgene vectors in a large animal model. Mol. Ther. 13, 59–66 (2006).
Kues, W.A. et al. Epigenetic silencing and tissue independent expression of a novel tetracycline inducible system in double-transgenic pigs. FASEB J. 20, 1200–1202 (2006).
Hall, V.J. Early development of the porcine embryo: the importance of cell signalling in development of pluripotent cell lines. Reprod. Fertil. Dev. 25, 94–102 (2012).
Esteban, M.A. et al. Generation of induced pluripotent stem cell lines from Tibetan miniature pig. J. Biol. Chem. 284, 17634–17640 (2009).
Ezashi, T. et al. Derivation of induced pluripotent stem cells from pig somatic cells. Proc. Natl. Acad. Sci. USA 106, 10993–10998 (2009).
Wu, Z. et al. Generation of pig induced pluripotent stem cells with a drug-inducible system. J. Mol. Cell Biol. 1, 46–54 (2009).
Kues, W.A. et al. Derivation and characterization of sleeping beauty transposon-mediated porcine induced pluripotent stem cells. Stem Cells Dev. 22, 124–135 (2013).
Montserrat, N. et al. Generation of pig iPS cells: a model for cell therapy. J. Cardiovasc. Transl. Res. 4, 121–130 (2011).
West, F.D. et al. Brief report: chimeric pigs produced from induced pluripotent stem cells demonstrate germline transmission and no evidence of tumor formation in young pigs. Stem Cells 29, 1640–1643 (2011).
Fujishiro, S.H. et al. Generation of naive-like porcine-induced pluripotent stem cells capable of contributing to embryonic and fetal development. Stem Cells Dev. 22, 473–482 (2013).
Fan, N. et al. Piglets cloned from induced pluripotent stem cells. Cell Res. 23, 162–166 (2013).
Hofmann, A. et al. Efficient transgenesis in farm animals by lentiviral vectors. EMBO Rep. 4, 1054–1060 (2003).
Whitelaw, C.B. et al. Efficient generation of transgenic pigs using equine infectious anaemia virus (EIAV) derived vector. FEBS Lett. 571, 233–236 (2004).
Kurome, M. et al. Effects of sperm pretreatment on efficiency of ICSI-mediated gene transfer in pigs. J. Reprod. Dev. 53, 1217–1226 (2007).
Watanabe, M. et al. The creation of transgenic pigs expressing human proteins using BAC-derived, full-length genes and intracytoplasmic sperm injection-mediated gene transfer. Transgenic Res. 21, 605–618 (2012).
Ivics, Z., Hackett, P.B., Plasterk, R.H. & Izsvák, Z. Molecular reconstruction of Sleeping Beauty, a Tc1-like transposon from fish, and its transposition in human cells. Cell 91, 501–510 (1997).
Ivics, Z. et al. Transposon-mediated genome manipulation in vertebrates. Nat. Methods 6, 415–422 (2009).
Zayed, H., Izsvák, Z., Walisko, O. & Ivics, Z. Development of hyperactive Sleeping Beauty transposon vectors by mutational analysis. Mol. Ther. 9, 292–304 (2004).
Rostovskaya, M. et al. Transposon-mediated BAC transgenesis in human ES cells. Nucleic Acids Res. 40, e150 (2012).
Voigt, K. et al. Retargeting Sleeping Beauty transposon insertions by engineered zinc finger DNA-binding domains. Mol. Ther. 20, 1852–1862 (2012).
Moldt, B. et al. Comparative genomic integration profiling of Sleeping Beauty transposons mobilized with high efficacy from integrase-defective lentiviral vectors in primary human cells. Mol. Ther. 19, 1499–1510 (2011).
Grabundzija, I. et al. Comparative analysis of transposable element vector systems in human cells. Mol. Ther. 18, 1200–1209 (2010).
Ammar, I. et al. Retargeting transposon insertions by the adeno-associated virus Rep protein. Nucleic Acids Res. 40, 6693–6712 (2012).
Ivics, Z. & Izsvák, Z. The expanding universe of transposon technologies for gene and cell engineering. Mob. DNA 1, 25 (2010).
Ammar, I., Izsvák, Z. & Ivics, Z. The Sleeping Beauty transposon toolbox. Methods Mol. Biol. 859, 229–240 (2012).
Mates, L. et al. Molecular evolution of a novel hyperactive Sleeping Beauty transposase enables robust stable gene transfer in vertebrates. Nat. Genet. 41, 753–761 (2009).
Katter, K. et al. Transposon-mediated transgenesis, transgenic rescue, and tissue-specific gene expression in rodents and rabbits. FASEB J. 27, 930–941 (2013).
Ivics, Z. et al. Germline transgenesis in rodents by pronuclear microinjection of Sleeping Beauty transposons. Nat. Protoc. 9, 773–793 (2014).
Ivics, Z. et al. Germline transgenesis in rabbits by pronuclear microinjection of Sleeping Beauty transposons. Nat. Protoc. 9, 794–809 (2014).
Garrels, W. et al. Germline transgenic pigs by Sleeping Beauty transposition in porcine zygotes and targeted integration in the pig genome. PLoS ONE 6, e23573 (2011).
Ellis, J. Silencing and variegation of gammaretrovirus and lentivirus vectors. Hum. Gene Ther. 16, 1241–1246 (2005).
Jahner, D. et al. De novo methylation and expression of retroviral genomes during mouse embryogenesis. Nature 298, 623–628 (1982).
Wolf, D. & Goff, S.P. Embryonic stem cells use ZFP809 to silence retroviral DNAs. Nature 458, 1201–1204 (2009).
Park, F. Lentiviral vectors: are they the future of animal transgenesis? Physiol. Genomics 31, 159–173 (2007).
Turan, S. & Bode, J. Site-specific recombinases: from tag-and-target- to tag-and-exchange-based genomic modifications. FASEB J. 25, 4088–4107 (2011).
Garrels, W. et al. Genotype-independent transmission of transgenic fluorophore protein by boar spermatozoa. PLoS ONE 6, e27563 (2011).
Claeys Bouuaert, C., Lipkow, K., Andrews, S.S., Liu, D. & Chalmers, R. The autoregulation of a eukaryotic DNA transposon. Elife 2, e00668 (2013).
Jakobsen, J.E. et al. Pig transgenesis by Sleeping Beauty DNA transposition. Transgenic Res. 20, 533–545 (2011).
Carlson, D.F. et al. Strategies for selection marker-free swine transgenesis using the Sleeping Beauty transposon system. Transgenic Res. 20, 1125–1137 (2011).
Al-Mashhadi, R.H. et al. Familial hypercholesterolemia and atherosclerosis in cloned minipigs created by DNA transposition of a human PCSK9 gain-of-function mutant. Sci. Transl. Med. 5, 166ra1 (2013).
Wu, Z. et al. Pig transgenesis by piggyBac transposition in combination with somatic cell nuclear transfer. Transgenic Res. 22, 1107–1118 (2013).
Iqbal, K. et al. Cytoplasmic injection of circular plasmids allows targeted expression in mammalian embryos. Biotechniques 47, 959–968 (2009).
Iqbal, K. et al. Species-specific telomere length differences between blastocyst cell compartments and ectopic telomere extension in early bovine embryos by human telomerase reverse transcriptase. Biol. Reprod. 84, 723–733 (2011).
O'Malley, R.C., Alonso, J.M., Kim, C.J., Leisse, T.J. & Ecker, J.R. An adapter ligation-mediated PCR method for high-throughput mapping of T-DNA inserts in the Arabidopsis genome. Nat. Protoc. 2, 2910–2917 (2007).
Ivics, Z., Izsvák, Z., Medrano, G., Chapman, K.M. & Hamra, F.K. Sleeping Beauty transposon mutagenesis in rat spermatogonial stem cells. Nat. Protoc. 6, 1521–1535 (2011).
Hyttel, P., Sinowatz, F., Vejlsted, M. & Betteridge,, K. Essentials of Domestic Animal Embryology (Saunders Elsevier, 2010).
Garrels, W., Cleve, N., Niemann, H. & Kues, W.A. Rapid non-invasive genotyping of reporter transgenic mammals. Biotechniques 10.2144/000113874 (2012).
Acknowledgements
This work was financially supported by grants of the Deutsche Forschungsgemeinschaft to W.A.K. and Z. Ivics (KU 1586/2-1 and IV 21/6-1). The expert support and critical input by J.W. Carnwath, S. Holler, B. Barg-Kues, N. Cleve, M. Ziegler and M. Diederich are also gratefully acknowledged. Whole-animal images under fluorescence excitation were done by the Friedrich Loeffler Institute's photographer D.B., and the video was recorded by P. Köhler.
Author information
Authors and Affiliations
Contributions
W.A.K., T.R., M.P., A.G., Z. Ivics and Z. Izsvák designed the study; W.G., L.M., T.Y.Y., S.B., V.Z., V.L., A.G., Z. Ivics and W.A.K. performed the experiments; W.A.K., Z. Ivics and W.G. evaluated the data; and W.A.K., Z. Ivics, T.R., W.G. and Z. Izsvák wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
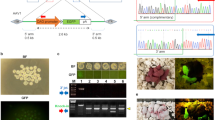
Supplementary Figure 1 Identification of genomic transgene integration by PCR.
(a) Identification of integrated transposon sequences from mouse genomic DNA samples by PCR with primers that amplify the left ITR of SB. Lanes: 1) H2O; 2) mouse genomic DNA, negative control; 3) mouse genomic DNA, positive control; 4) transgenic founder #1; 5-9) F1 offspring of transgenic founder #1; 10) transgenic founder #2; 11-13) F1 offspring of transgenic founder #2; M) 100-bp molecular size marker. (b) Ouline of the LMPCR procedure. Digestion of genomic DNA with the frequently cutting restriction enzymes BfaI and DpnII and ligation of linkers with a known sequence allows for specific LMPCR amplification of transposon/genomic DNA junctions using primers specific to the transposon ITR (blue arrows) and the linkers (green arrows). GOI, gene of interest; ITR, inverted terminal repeat. (c) Agarose gel with genomic DNA samples. Lanes 1-4) genomic DNA samples of rat founders, 500 ng each; M) DNA size marker. (d) Agarose gel with genomic DNA samples digested with BfaI restriction endonuclease. Lanes 1-4) BfaI-digested DNA samples of rat founders, 200 ng each; M) DNA size marker. (e) Agarose gel with LMPCR products. Lanes 1-4) result of nested PCR following BfaI linker ligation; M) DNA size marker.
Supplementary Figure 2 Locus-specific PCR test of a rat founder and its F1 descendants.
The founder of these F1 animals carried three SB integrations (in chr2, chr4 and chr16), which were transmitted to 13 descendants in different combinations. M, DNA size marker.
Supplementary information
Supplementary Figure 1
Identification of genomic transgene integration by PCR. (PDF 488 kb)
Supplementary Figure 2
Locus-specific PCR test of a rat founder and its F1 descendants. (PDF 361 kb)
Copy number-dependent fluorescence in F2-generation piglets.
The F2 litter of piglets are derived from the crossbreeding of two lines. Each founder carried three monomeric Venus-transposons, which segregated individually during meiosis. The piglets shown carry 0 to 5 copies of the Venus-transposon, and the fluorescent intensity correlates directly with the genotype. The “bluish” piglet in the back is a non-transgenic littermate. (MPG 1838 kb)
Rights and permissions
About this article
Cite this article
Ivics, Z., Garrels, W., Mátés, L. et al. Germline transgenesis in pigs by cytoplasmic microinjection of Sleeping Beauty transposons. Nat Protoc 9, 810–827 (2014). https://doi.org/10.1038/nprot.2014.010
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.010
This article is cited by
-
Improvement of Sleeping Beauty Transposon System Enabling Efficient and Stable Protein Production
Biotechnology and Bioprocess Engineering (2022)
-
Improvement of Tol2 Transposon System by Modification of Tol2 Transposase
Biotechnology and Bioprocess Engineering (2022)
-
One-step genome editing of porcine zygotes through the electroporation of a CRISPR/Cas9 system with two guide RNAs
In Vitro Cellular & Developmental Biology - Animal (2020)
-
Long-term health and germline transmission in transgenic cattle following transposon-mediated gene transfer
BMC Genomics (2018)
-
Changes in Skeletal Muscle and Body Weight on Sleeping Beauty Transposon-Mediated Transgenic Mice Overexpressing Pig mIGF-1
Biochemical Genetics (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.