Abstract
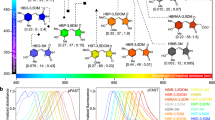
Naturally occurring fluorescent proteins (FPs) cloned from marine organisms often suffer from many drawbacks for cell biology applications, including poor folding efficiency at 37 °C, slow chromophore formation and obligatory quaternary structure. Many of these drawbacks can be minimized or eliminated by using protein engineering and directed evolution, resulting in superior probes for use in live-cell fluorescence microscopy. In this protocol, we provide methods for engineering a monomeric FP, for enhancing its brightness by directed evolution, and for thoroughly characterizing the optimized variant. Variations on this procedure can be used to select for many other desirable features, such as a red-shifted emission spectrum or enhanced photostability. Although the length of the procedure is dependent on the degree of optimization desired, the basic steps can be accomplished in 4–6 weeks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Prasher, D.C., Eckenrode, V.K., Ward, W.W., Prendergast, F.G. & Cormier, M.J. Primary structure of the Aequorea victoria green-fluorescent protein. Gene 111, 229–233 (1992).
Chalfie, M., Tu, Y., Euskirchen, G., Ward, W.W. & Prasher, D.C. Green fluorescent protein as a marker for gene expression. Science 263, 802–805 (1994).
Inouye, S. & Tsuji, F.I. Aequorea green fluorescent protein. Expression of the gene and fluorescence characteristics of the recombinant protein. FEBS Lett. 341, 277–280 (1994).
Matz, M.V. et al. Fluorescent proteins from nonbioluminescent Anthozoa species. Nat. Biotechnol. 17, 969–973 (1999).
Campbell, R.E. et al. A monomeric red fluorescent protein. Proc. Natl. Acad. Sci. USA 99, 7877–7882 (2002).
Shaner, N.C. et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat. Biotechnol. 22, 1567–1572 (2004).
Ai, H., Henderson, J.N., Remington, S.J. & Campbell, R.E. Directed evolution of a monomeric, bright and photostable version of Clavularia cyan fluorescent protein: structural characterization and applications in fluorescence imaging. Biochem. J. 400, 531–540 (2006).
Hoi, H. et al. An engineered monomeric Zoanthus sp. Yellow fluorescent protein. Chem. Biol. 20, 1296–1304 (2013).
Baird, G.S., Zacharias, D.A. & Tsien, R.Y. Biochemistry, mutagenesis, and oligomerization of DsRed, a red fluorescent protein from coral. Proc. Natl. Acad. Sci. USA 97, 11984–11989 (2000).
Lauf, U., Lopez, P. & Falk, M.M. Expression of fluorescently tagged connexins: a novel approach to rescue function of oligomeric DsRed-tagged proteins. FEBS Lett. 498, 11–15 (2001).
Gavin, P., Devenish, R.J. & Prescott, M. An approach for reducing unwanted oligomerisation of DsRed fusion proteins. Biochem. Biophys. Res. Commun. 298, 707–713 (2002).
Soling, A., Simm, A. & Rainov, N. Intracellular localization of herpes simplex virus type 1 thymidine kinase fused to different fluorescent proteins depends on choice of fluorescent tag. FEBS Lett. 527, 153–158 (2002).
Shaner, N.C. et al. Improving the photostability of bright monomeric orange and red fluorescent proteins. Nat. Methods 5, 545–551 (2008).
Karasawa, S., Araki, T., Yamamoto-Hino, M. & Miyawaki, A. A green-emitting fluorescent protein from Galaxeidae coral and its monomeric version for use in fluorescent labeling. J. Biol. Chem. 278, 34167–34171 (2003).
Shaner, N.C. et al. A bright monomeric green fluorescent protein derived from Branchiostoma lanceolatum. Nat. Methods 10, 407–409 (2013).
Karasawa, S., Araki, T., Nagai, T., Mizuno, H. & Miyawaki, A. Cyan-emitting and orange-emitting fluorescent proteins as a donor/acceptor pair for fluorescence resonance energy transfer. Biochem. J. 381, 307–312 (2004).
Wiedenmann, J. et al. EosFP, a fluorescent marker protein with UV-inducible green-to-red fluorescence conversion. Proc. Natl. Acad. Sci. USA 101, 15905–15910 (2004).
Gurskaya, N.G. et al. Engineering of a monomeric green-to-red photoactivatable fluorescent protein induced by blue light. Nat. Biotechnol. 24, 461–465 (2006).
Ando, R., Mizuno, H. & Miyawaki, A. Regulated fast nucleocytoplasmic shuttling observed by reversible protein highlighting. Science 306, 1370–1373 (2004).
Kogure, T. et al. A fluorescent variant of a protein from the stony coral Montipora facilitates dual-color single-laser fluorescence cross-correlation spectroscopy. Nat. Biotechnol. 24, 577–581 (2006).
Shagin, D.A. et al. GFP-like proteins as ubiquitous metazoan superfamily: evolution of functional features and structural complexity. Mol. Biol. Evol. 21, 841–850 (2004).
Alieva, N.O. et al. Diversity and evolution of coral fluorescent proteins. PLoS ONE 3, e2680 (2008).
Pavoor, T.V., Cho, Y.K. & Shusta, E.V. Development of GFP-based biosensors possessing the binding properties of antibodies. Proc. Natl. Acad. Sci. USA 106, 11895–11900 (2009).
Wang, L., Jackson, W.C., Steinbach, P.A. & Tsien, R.Y. Evolution of new nonantibody proteins via iterative somatic hypermutation. Proc. Natl. Acad. Sci. USA 101, 16745–16749 (2004).
Arakawa, H. et al. Protein evolution by hypermutation and selection in the B cell line DT40. Nucleic Acids Res. 36, e1 (2008).
Cormack, B.P., Valdivia, R.H. & Falkow, S. FACS-optimized mutants of the green fluorescent protein (GFP). Gene 173, 33–38 (1996).
Nguyen, A.W. & Daugherty, P.S. Evolutionary optimization of fluorescent proteins for intracellular FRET. Nat. Biotechnol. 23, 355–360 (2005).
Mena, M.A., Treynor, T.P., Mayo, S.L. & Daugherty, P.S. Blue fluorescent proteins with enhanced brightness and photostability from a structurally targeted library. Nat. Biotechnol. 24, 1569–1571 (2006).
Dai, M. et al. The creation of a novel fluorescent protein by guided consensus engineering. Protein Eng. Des. Sel. 20, 69–79 (2007).
Hoi, H. et al. A monomeric photoconvertible fluorescent protein for imaging of dynamic protein localization. J. Mol. Biol. 401, 776–791 (2010).
Treynor, T.P., Vizcarra, C.L., Nedelcu, D. & Mayo, S.L. Computationally designed libraries of fluorescent proteins evaluated by preservation and diversity of function. Proc. Natl. Acad. Sci. USA 104, 48–53 (2007).
Chica, R.A., Moore, M.M., Allen, B.D. & Mayo, S.L. Generation of longer emission wavelength red fluorescent proteins using computationally designed libraries. Proc. Natl. Acad. Sci. USA 107, 20257–20262 (2010).
Stemmer, W.P., Crameri, A., Ha, K.D., Brennan, T.M. & Heyneker, H.L. Single-step assembly of a gene and entire plasmid from large numbers of oligodeoxyribonucleotides. Gene 164, 49–53 (1995).
Bessette, P.H., Mena, M.A., Nguyen, A.W. & Daugherty, P.S. Construction of designed protein libraries using gene assembly mutagenesis. In Directed Evolution Library Creation: Methods and Protocols 1st edn., Vol. 231 (eds. Arnold, F.H. & Georgiou, G.) 29–37 (Humana Press, 2003).
Yang, T.T., Cheng, L. & Kain, S.R. Optimized codon usage and chromophore mutations provide enhanced sensitivity with the green fluorescent protein. Nucleic Acids Res. 24, 4592–4593 (1996).
Tsien, R.Y. The green fluorescent protein. Annu. Rev. Biochem. 67, 509–544 (1998).
Yarbrough, D., Wachter, R.M., Kallio, K., Matz, M.V. & Remington, S.J. Refined crystal structure of DsRed, a red fluorescent protein from coral, at 2.0-Å resolution. Proc. Natl. Acad. Sci. USA 98, 462–467 (2001).
Ormo, M. et al. Crystal structure of the Aequorea victoria green fluorescent protein. Science 273, 1392–1395 (1996).
Fromant, M., Blanquet, S. & Plateau, P. Direct random mutagenesis of gene-sized DNA fragments using polymerase chain reaction. Anal. Biochem. 224, 347–353 (1995).
Cirino, P.C., Mayer, K.M. & Umeno, D. Generating mutant libraries using error-prone PCR. In Directed Evolution Library Creation: Methods and Protocols 1st edn., Vol. 231 (eds. Arnold, F.H. & Georgiou, G.) 3–9 (Humana Press, 2003).
Ho, S.N., Hunt, H.D., Horton, R.M., Pullen, J.K. & Pease, L.R. Site-directed mutagenesis by overlap extension using the polymerase chain reaction. Gene 77, 51–59 (1989).
Heckman, K.L. & Pease, L.R. Gene splicing and mutagenesis by PCR-driven overlap extension. Nat. Protoc. 2, 924–932 (2007).
Georgescu, R., Bandara, G. & Sun, L. Saturation mutagenesis. In Directed Evolution Library Creation: Methods and Protocols 1st edn., Vol. 231 (eds. Arnold, F.H. & Georgiou, G.) 75–83 (Humana Press, 2003).
Aguinaldo, A.M. & Arnold, F.H. Staggered extension process (StEP) in vitro recombination. In Directed Evolution Library Creation: Methods and Protocols 1st edn., Vol. 231 (eds. Arnold, F.H. & Georgiou, G.) 105–110 (Humana Press, 2003).
Zhao, H. & Zha, W. In vitro 'sexual' evolution through the PCR-based staggered extension process (StEP). Nat. Protoc. 1, 1865–1871 (2006).
Trudeau, D.L., Smith, M.A. & Arnold, F.H. Innovation by homologous recombination. Curr. Opin. Chem. Biol. 17 (2013), 902–909.
Lindenburg, L.H., Hessels, A.M., Ebberink, E.H., Arts, R. & Merkx, M. Robust red FRET sensors using self-associating fluorescent domains. ACS Chem. Biol. 8, 2133–2139 (2013).
Shaner, N.C., Steinbach, P.A. & Tsien, R.Y. A guide to choosing fluorescent proteins. Nat. Methods 2, 905–909 (2005).
Crameri, A., Whitehorn, E.A., Tate, E. & Stemmer, W.P. Improved green fluorescent protein by molecular evolution using DNA shuffling. Nat. Biotechnol. 14, 315–319 (1996).
Berman, H.M. et al. The protein data bank. Nucleic Acids Res. 28, 235–242 (2000).
Strack, R.L. et al. A rapidly maturing far-red derivative of DsRed-Express2 for whole-cell labeling. Biochemistry 48, 8279–8281 (2009).
Adams, M.J., Highfield, J.G. & Kirkbright, G.F. Determination of absolute fluorescence quantum efficiency of quinine bisulfate in aqueous-medium by optoacoustic spectrometry. Anal. Chem. 49, 1850–1852 (1977).
Brannon, J.H. & Magde, D. Absolute quantum yield determination by thermal blooming. Fluorescein. J. Phys. Chem. 82, 705–709 (1978).
Karstens, T. & Kobs, K. Rhodamine-B and rhodamine-101 as reference substances for fluorescence quantum yield measurements. J. Phys. Chem. 84, 1871–1872 (1980).
Ward, W.W. Biochemical and physical properties of green fluorescent protein. In Green Fluorescent Protein: Properties, Applications, and Protocols 2nd edn., Vol. 47 (eds. Chalfie, M. & Kain, S.R.) 39–65 (John Wiley & Sons, 2006).
Ai, H., Olenych, S.G., Wong, P., Davidson, M.W. & Campbell, R.E. Hue-shifted monomeric variants of Clavularia cyan fluorescent protein: identification of the molecular determinants of color and applications in fluorescence imaging. BMC Biol. 6, 13 (2008).
Shu, X., Shaner, N.C., Yarbrough, C.A., Tsien, R.Y. & Remington, S.J. Novel chromophores and buried charges control color in mFruits. Biochemistry 45, 9639–9647 (2006).
Acknowledgements
This research was made possible with financial support from the University of Alberta, the Canada Foundation for Innovation, the Natural Sciences and Engineering Research Council of Canada, and Alberta Ingenuity (scholarship to Y.S. and a New Faculty Award to R.E.C.). R.E.C. holds a Canada Research Chair in Bioanalytical Chemistry.
Author information
Authors and Affiliations
Contributions
M.W.D. and M.A.B. acquired data and images shown in Figures 3 and 6. H.A., M.A.B., Y.S., M.W.D. and R.E.C. developed the protocols and wrote and edited the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Ai, Hw., Baird, M., Shen, Y. et al. Engineering and characterizing monomeric fluorescent proteins for live-cell imaging applications. Nat Protoc 9, 910–928 (2014). https://doi.org/10.1038/nprot.2014.054
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.054
This article is cited by
-
Absolute measurement of cellular activities using photochromic single-fluorophore biosensors and intermittent quantification
Nature Communications (2022)
-
Tutorial: guidance for quantitative confocal microscopy
Nature Protocols (2020)
-
HriGFP Novel Flourescent Protein: Expression and Applications
Molecular Biotechnology (2020)
-
A genetically encoded Ca2+ indicator based on circularly permutated sea anemone red fluorescent protein eqFP578
BMC Biology (2018)
-
A robotic multidimensional directed evolution approach applied to fluorescent voltage reporters
Nature Chemical Biology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.