Abstract
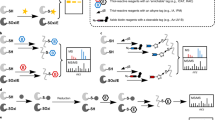
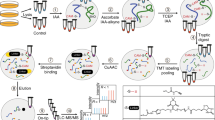
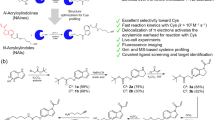
Reversible modifications of cysteine thiols have a key role in redox signaling and regulation. A number of reversible redox modifications, including disulfide formation, S-nitrosylation (SNO) and S-glutathionylation (SSG), have been recognized for their significance in various physiological and pathological processes. Here we describe a procedure for the enrichment of peptides containing reversible cysteine modifications. Starting with tissue or cell lysate samples, all of the unmodified free thiols are blocked using N-ethylmaleimide (NEM). This is followed by the selective reduction of those cysteines bearing the reversible modification(s) of interest. The reduction is achieved by using different reducing reagents that react specifically with each type of cysteine modification (e.g., ascorbate for SNO). This protocol serves as a general approach for enrichment of thiol-containing proteins or peptides derived from reversibly modified proteins. The approach uses a commercially available thiol-affinity resin (thiopropyl Sepharose 6B) to directly capture free thiol-containing proteins through a disulfide exchange reaction, followed by on-resin protein digestion and multiplexed isobaric labeling to facilitate liquid chromatography-tandem mass spectrometry (LC-MS/MS)-based quantitative site-specific analysis of cysteine-based reversible modifications. The overall approach requires a simpler workflow with increased specificity compared with the commonly used biotinylation-based assays. The procedure for selective enrichment and analyses of SNO and the level of total reversible cysteine modifications (or total oxidation) is presented to demonstrate the utility of this general strategy. The entire protocol requires ∼3 d for sample processing with an additional day for LC-MS/MS and data analysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Giron, P., Dayon, L. & Sanchez, J.C. Cysteine tagging for MS-based proteomics. Mass Spectrom. Rev. 30, 366–395 (2011).
Held, J.M. & Gibson, B.W. Regulatory control or oxidative damage? Proteomic approaches to interrogate the role of cysteine oxidation status in biological processes. Mol. Cell Proteomics 11, R111.013037 (2012).
Antelmann, H. & Helmann, J.D. Thiol-based redox switches and gene regulation. Antioxid. Redox Signal. 14, 1049–1063 (2011).
Bachi, A., Dalle-Donne, I. & Scaloni, A. Redox proteomics: chemical principles, methodological approaches and biological/biomedical promises. Chem. Rev. 113, 596–698 (2013).
Sato, Y. & Inaba, K. Disulfide bond formation network in the three biological kingdoms, bacteria, fungi and mammals. FEBS J. 279, 2262–2271 (2012).
Derakhshan, B., Wille, P.C. & Gross, S.S. Unbiased identification of cysteine S-nitrosylation sites on proteins. Nat. Protoc. 2, 1685–1691 (2007).
Greco, T.M. et al. Identification of S-nitrosylation motifs by site-specific mapping of the S-nitrosocysteine proteome in human vascular smooth muscle cells. Proc. Natl. Acad. Sci. USA 103, 7420–7425 (2006).
Hess, D.T., Matsumoto, A., Kim, S.O., Marshall, H.E. & Stamler, J.S. Protein S-nitrosylation: purview and parameters. Nat. Rev. Mol. Cell. Biol. 6, 150–166 (2005).
Paulsen, C.E. et al. Peroxide-dependent sulfenylation of the EGFR catalytic site enhances kinase activity. Nat. Chem. Biol. 8, 57–64 (2012).
Wan, J., Roth, A.F., Bailey, A.O. & Davis, N.G. Palmitoylated proteins: purification and identification. Nat. Protoc. 2, 1573–1584 (2007).
Roth, A.F. et al. Global analysis of protein palmitoylation in yeast. Cell 125, 1003–1013 (2006).
Brandes, N., Schmitt, S. & Jakob, U. Thiol-based redox switches in eukaryotic proteins. Antioxid. Redox Signal. 11, 997–1014 (2009).
Jortzik, E., Wang, L. & Becker, K. Thiol-based posttranslational modifications in parasites. Antioxid. Redox Signal. 17, 657–673 (2012).
Jaffrey, S.R., Erdjument-Bromage, H., Ferris, C.D., Tempst, P. & Snyder, S.H. Protein S-nitrosylation: a physiological signal for neuronal nitric oxide. Nat. Cell. Biol. 3, 193–197 (2001).
Jaffrey, S.R. & Snyder, S.H. The biotin switch method for the detection of S-nitrosylated proteins. Sci. STKE 2001, pl1 (2001).
Lind, C. et al. Identification of S-glutathionylated cellular proteins during oxidative stress and constitutive metabolism by affinity purification and proteomic analysis. Arch. Biochem. Biophys. 406, 229–240 (2002).
Reynaert, N.L. et al. In situ detection of S-glutathionylated proteins following glutaredoxin-1 catalyzed cysteine derivatization. Biochim. Biophys. Acta 1760, 380–387 (2006).
Leichert, L.I. et al. Quantifying changes in the thiol redox proteome upon oxidative stress in vivo. Proc. Natl. Acad. Sci. USA 105, 8197–8202 (2008).
Hao, G., Derakhshan, B., Shi, L., Campagne, F. & Gross, S.S. SNOSID, a proteomic method for identification of cysteine S-nitrosylation sites in complex protein mixtures. Proc. Natl. Acad. Sci. USA 103, 1012–1017 (2006).
Liu, T. et al. Improved proteome coverage using high-efficiency cysteinyl peptide enrichment: the mammary epithelial cell proteome. Proteomics 5, 1263–1273 (2005).
Liu, T. et al. High-throughput comparative proteome analysis using a quantitative cysteinyl-peptide enrichment technology. Anal. Chem. 76, 5345–5353 (2004).
Forrester, M.T. et al. Proteomic analysis of S-nitrosylation and denitrosylation by resin-assisted capture. Nat. Biotechnol. 27, 557–559 (2009).
Su, D. et al. Quantitative site-specific reactivity profiling of S-nitrosylation in mouse skeletal muscle using cysteinyl peptide enrichment coupled with mass spectrometry. Free Radic. Biol. Med. 57, 68–78 (2013).
Liu, M. et al. Site-specific proteomics approach for study protein s-nitrosylation. Anal. Chem. 82, 7160–7168 (2010).
Forrester, M.T. et al. Site-specific analysis of protein S-acylation by resin-assisted capture. J. Lipid Res. 52, 393–398 (2011).
Paulech, J. et al. Large-scale capture of peptides containing reversibly oxidized cysteines by thiol-disulfide exchange applied to the myocardial redox proteome. Anal. Chem. 85, 3774–3780 (2013).
Ross, P.L. et al. Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol. Cell. Proteomics 3, 1154–1169 (2004).
Dayon, L. et al. Relative quantification of proteins in human cerebrospinal fluids by MS/MS using 6-plex isobaric tags. Anal. Chem. 80, 2921–2931 (2008).
Shelton, M.D., Chock, P.B. & Mieyal, J.J. Glutaredoxin: role in reversible protein s-glutathionylation and regulation of redox signal transduction and protein translocation. Antioxid. Redox Signal. 7, 348–366 (2005).
Zhang, C., Rodriguez, C., Circu, M.L., Aw, T.Y. & Feng, J. S-Glutathionyl quantification in the attomole range using glutaredoxin-3-catalyzed cysteine derivatization and capillary gel electrophoresis with laser-induced fluorescence detection. Anal. Bioanal. Chem. 401, 2165–2175 (2011).
Mustafa, A.K. et al. H2S signals through protein S-sulfhydration. Sci. Signal. 2, ra72 (2009).
Pan, J. & Carroll, K.S. Persulfide reactivity in the detection of protein S-sulfhydration. ACS Chem. Biol. 2013, 1110–1116 (2013).
Forrester, M.T., Foster, M.W. & Stamler, J.S. Assessment and application of the biotin switch technique for examining protein S-nitrosylation under conditions of pharmacologically induced oxidative stress. J. Biol. Chem. 282, 13977–13983 (2007).
Forrester, M.T., Foster, M.W., Benhar, M. & Stamler, J.S. Detection of protein S-nitrosylation with the biotin-switch technique. Free Radic. Biol. Med. 46, 119–126 (2009).
Ong, S.E. et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell Proteomics 1, 376–386 (2002).
Murray, C.I., Uhrigshardt, H., O'Meally, R.N., Cole, R.N. & Van Eyk, J.E. Identification and quantification of S-nitrosylation by cysteine reactive tandem mass tag switch assay. Mol. Cell Proteomics 11, M111 013441 (2012).
Foster, M.W. Methodologies for the characterization, identification and quantification of S-nitrosylated proteins. Biochim. Biophys. Acta 1820, 675–683 (2012).
Zhang, Y., Ficarro, S.B., Li, S. & Marto, J.A. Optimized orbitrap HCD for quantitative analysis of phosphopeptides. J. Am. Soc. Mass Spectrom. 20, 1425–1434 (2009).
Kelly, R.T. et al. Chemically etched open tubular and monolithic emitters for nanoelectrospray ionization mass spectrometry. Anal. Chem. 78, 7796–7801 (2006).
Livesay, E.A. et al. Fully automated four-column capillary LC-MS system for maximizing throughput in proteomic analyses. Anal. Chem. 80, 294–302 (2008).
Eng, J.K., Mccormack, A.L. & Yates, J.R. An approach to correlate tandem mass-spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Elias, J.E. & Gygi, S.P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Qian, W.J. et al. Probability-based evaluation of peptide and protein identifications from tandem mass spectrometry and SEQUEST analysis: the human proteome. J. Proteome Res. 4, 53–62 (2005).
Kim, S., Gupta, N. & Pevzner, P.A. Spectral probabilities and generating functions of tandem mass spectra: a strike against decoy databases. J. Proteome Res. 7, 3354–3363 (2008).
Viner, R.I., Williams, T.D. & Schoneich, C. Peroxynitrite modification of protein thiols: oxidation, nitrosylation, and S-glutathiolation of functionally important cysteine residue(s) in the sarcoplasmic reticulum Ca-ATPase. Biochemistry 38, 12408–12415 (1999).
Knoefler, D. et al. Quantitative in vivo redox sensors uncover oxidative stress as an early event in life. Mol. Cell 47, 767–776 (2012).
Acknowledgements
Portions of this work were supported by the US National Institutes of Health (NIH) Director's New Innovator Award Program DP2OD006668 and a US Department of Energy (DOE) Early Career Research Award (to W.-J.Q.), NIH P41 GM103493 (to R.D.S.), and the DOE Office of Biological and Environmental Research Genome Sciences Program under the Pan-omics project. The experimental work was performed in the Environmental Molecular Science Laboratory, a DOE/Biological and Environmental Research (BER) national scientific user facility at the Pacific Northwest National Laboratory (PNNL) in Richland, Washington. PNNL is operated by Battelle for the DOE under contract no. DE-AC05-76RLO-1830.
Author information
Authors and Affiliations
Contributions
J.G., M.J.G. and D.S. performed the experiments and optimized the protocol; T.L. developed the initial enrichment method; D.G.C. and R.D.S. contributed to development of the measurement capabilities used; W.-J.Q. conceived and supervised the project. J.G., M.J.G. and W.-J.Q. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Conditions for selective reduction, negative and positive controls for multiple reversible cysteine modifications. (PDF 223 kb)
Supplementary Data
Quantification of total cysteine oxidation levels in RAW cells treated with 0.1 mM and 0.5 mM diamide. (XLSX 220 kb)
Source data
Rights and permissions
About this article
Cite this article
Guo, J., Gaffrey, M., Su, D. et al. Resin-assisted enrichment of thiols as a general strategy for proteomic profiling of cysteine-based reversible modifications. Nat Protoc 9, 64–75 (2014). https://doi.org/10.1038/nprot.2013.161
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.161
This article is cited by
-
Oxidative stress-induced FABP5 S-glutathionylation protects against acute lung injury by suppressing inflammation in macrophages
Nature Communications (2021)
-
A quantitative thiol reactivity profiling platform to analyze redox and electrophile reactive cysteine proteomes
Nature Protocols (2020)
-
Proteome-wide detection of S-nitrosylation targets and motifs using bioorthogonal cleavable-linker-based enrichment and switch technique
Nature Communications (2019)
-
Mass spectrometry-based proteomics for system-level characterization of biological responses to engineered nanomaterials
Analytical and Bioanalytical Chemistry (2018)
-
A simple isotopic labeling method to study cysteine oxidation in Alzheimer’s disease: oxidized cysteine-selective dimethylation (OxcysDML)
Analytical and Bioanalytical Chemistry (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.