Abstract
Induced pluripotent stem (iPS) cells hold the potential to revolutionize regenerative medicine through their capacity to generate cells of diverse lineages for future patient-specific cell-based therapies. To facilitate the transition of iPS cells to clinical practice, a variety of technologies have been developed for transgene-free pluripotency reprogramming. We recently reported efficient iPS cell generation from human fibroblasts using synthetic modified mRNAs. Here we describe a stepwise protocol for the generation of modified mRNA–derived iPS cells from primary human fibroblasts, focusing on the critical parameters including medium choice, quality control, and optimization steps needed for synthesizing modified mRNAs encoding reprogramming factors and introducing these into cells over the course of 2–3 weeks to ensure successful reprogramming. The protocol described herein is for reprogramming of human fibroblasts to pluripotency; however, the properties of modified mRNA make it a powerful platform for protein expression, which has broad applicability in directed differentiation, cell fate specification and therapeutic applications.
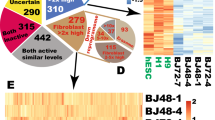
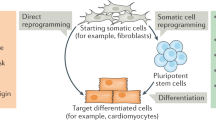
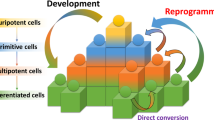
Similar content being viewed by others
Introduction
The seminal discovery by Yamanaka and colleagues1,2 that differentiated cell types could be reprogrammed to a state of pluripotency by a set of defined factors was a sea-change moment in stem cell biology that held enormous potential for regenerative medicine and future cell-based autologous therapies. Patient-derived iPS cells also provide a valuable cellular resource in which to study the cellular and molecular pathogenesis of diverse genetic diseases3. The fact that iPS cell–based disease modeling can be done in vitro has opened up the possibility of identifying disease-specific therapeutics through small-molecule screening and other high-throughput approaches4,5.
Development of nonintegrative approaches for iPS cell derivation
The initial approach used to reprogram mouse and human fibroblasts to iPS cells utilized retroviral vectors to deliver OCT4 (officially known as POU5F1), KLF4, SOX2 and c-MYC (officially known as MYC) to cells1,2. This approach has been widely adopted, in part, owing to the fact that retroviral-based methods are straightforward, reliable, inexpensive and easily adoptable by labs with basic skills in virus production and use. Although these properties make viral-based approaches ideal for basic research, cell-based therapeutic application of retrovirally derived iPS cells in patients is precluded because of the risks associated with the integration of viral sequences into the genome. To surmount this, a number of nonintegrating strategies for iPS cell generation have been developed using adenoviral vectors6,7, nonintegrating DNA plasmid-based vectors8,9,10,11,12,13,14,15,16,17, direct protein transduction18,19, Sendai viral vectors20,21 and the modified mRNA–based approach detailed here22. The strengths and weaknesses of these methodologies have been discussed previously5.
iPS cell derivation with synthetic modified mRNA
The labile nature and relatively short half-life of modified mRNA, combined with the fact that iPS cell derivation normally requires 3–4 weeks to complete, necessitates that a reprogramming strategy mediated by modified mRNA would involve repeated administration. Previous studies had, however, shown that transfection of RNA into mammalian cells resulted in severe cytotoxicity owing to the activation of innate immune responses by Toll-like receptors (TLR3, TLR7 and TLR8) and RNA sensors (RIG-I and PKR)23,24,25,26,27. Pioneering work from Kariko and Weissman26,28,29,30 at the University of Pennsylvania had shown, however, that incorporation of certain naturally occurring modified nucleosides into mRNA could largely abrogate immune activation. In light of these and other studies, modified mRNAs synthesized with complete substitution of cytidine and uridine with the modified nucleosides pseudouridine and 5-methylcytidine could be introduced repeatedly into cells with minimal activation of innate immune responses and limited cytotoxicity, thereby allowing robust and sustained protein expression22. Combined with medium supplementation of B18R, a Vaccinia virus–encoded neutralizing type I interferon receptor31 to quell residual interferon-mediated immune responses, daily transfection of modified mRNAs encoding the reprogramming factors (OCT4, SOX2, c-MYC, KLF4 and LIN28A) permitted successful reprogramming of human fibroblasts to pluripotency with high efficiency22.
Modified mRNA: applications beyond iPS cell generation
It must be recognized that iPS cells are only the starting point for patient-specific therapies, and that specification to diverse cell types is necessary for both disease modeling and to generate autologous tissues for cell-based therapies. Generally, protocols for differentiating pluripotent cells to defined fates are inefficient, and the derivation of many cell types is currently not possible. Thus, the ability to safely and efficiently control cell fate both during pluripotency reprogramming and during differentiation toward specific cell types would have a profound effect on bringing iPS cell technology closer to clinical application.
The modified-mRNA technology is endowed with a number of properties that make it a powerful platform for directing cell fate. These properties include the fact that modified mRNA can mediate robust, penetrant and dose-titratable expression of virtually any protein, in many different types of cells. Moreover, because multiple different modified mRNA species can be introduced into cells simultaneously, coexpression of multiple proteins can be achieved. Further, as exemplified in Figure 1 (for OCT4 and SOX2), by simply changing the dose of modified mRNA, the levels of any desired factors can be exquisitely controlled. This property may be used to explore the stoichiometry of various factors required during reprogramming or directed differentiation experiments. To our knowledge, no other reprogramming technology permits such control over reprogramming factor expression. Notably, the transient nature of protein expression mediated by modified mRNA—expression peaks within 10–16 h and diminishes by 24–36 h after transfection—opens up the possibility of experimentally activating developmentally important transcriptional programs through ectopic expression of transcription factors in a temporally controlled, stage-specific sequence in order to direct iPS cells to diverse fates. In proof-of-principle experiments, we have previously demonstrated that modified mRNA encoding MyoD could be used to direct the fate of iPS cells toward a myogenic fate with reasonable efficacy, and could also be used for transdifferentiation22. We anticipate that modified mRNA encoding lineage specification factors will emerge as an important tool for generating diverse cells types either by transdifferentiation or directed differentiation of iPS cells for experimental and therapeutic applications.
(a,b) Dermal fibroblasts transfected with the indicated modified mRNA showing protein expression by (a) immunostaining (a) and western blotting (WB) (b). The antibodies used are described in MATERIALS. DAPI, 4′,6-diamidino-2-phenylindole (a DNA-binding fluorescent stain); mES, mouse ES cell lysate. Scale bars, 50 μm.
Beyond applications for directing cell fate, recent studies have shown that modified mRNA can be used to express therapeutically active proteins in vivo. In a recent study, Rudolph and colleagues32 used pulmonary delivery of modified mRNA encoding surfactant protein B to rescue a mouse model of congenital lung disease caused by a lack of the surfactant protein B. In addition to this localized use, two groups have shown that modified mRNA can be used to produce systemic, biologically active erythropoietin in vivo32,33. Taken together, modified mRNA seems poised to emerge as a novel therapeutic paradigm for application in regenerative medicine and beyond.
Experimental design
The procedure has two major components: in vitro transcription (IVT) of modified mRNA for reprogramming factors (Steps 1–28) and generation of RiPS cells with modified mRNA (Steps 29 onward). The protocol for the generation of iPS cells using modified mRNA is labor intensive and costly. Any variations in the reagents involved greatly impede the outcome of the experiment. Given the labile nature of RNA, we recommend avoiding unnecessary delay between the steps while synthesizing modified mRNA. It is essential to work under RNase-free conditions. We always recommend including a modified mRNA encoding a reporter (e.g., GFP) every time modified mRNA is prepared as a surrogate for monitoring the IVT process. This preparation should then be tested for GFP expression by transfection into cells. To avoid repeated freeze-thaw cycles, modified mRNA or mRNA cocktail should be stored at −80 °C in workable volumes.
An ∼2-week timeline of a typical reprogramming experiment is outlined in Figure 2. Before starting a reprogramming experiment, ensure that all the reagents (media, transfection reagents and modified mRNA) are present in sufficient amounts. All reagents should be quality controlled before a reprogramming experiment is started, as we have encountered substantial lot-to-lot variability in commercially available reagents. In particular, transfection reagents should be tested using a suitable reporter–modified mRNA to ensure high transfection efficiency. Modified mRNA should be evaluated to ensure that encoded proteins are robustly produced (Step 28, Fig. 1). Changing the lot of commercially available reagents in the middle of an experiment (before testing) may adversely affect reprogramming. Culture medium has an enormous effect on the outcome of reprogramming efficacy. Although in our original publication we used Nutristem medium (Stemgent), we and others subsequently found that later lots of Nutristem were not supportive of modified mRNA–based reprogramming, probably resulting from changes in the manufacturing of this reagent. We and others34 have subsequently determined that another commercially available medium (Pluriton) is robustly supportive of modified-mRNA reprogramming, and we therefore recommend that all reprogramming experiments using modified mRNA be conducted in Pluriton medium.
(a) Schematic representation of a typical reprogramming experiment using modified mRNA indicating timing for key events. The medium used at each stage of the protocol is indicated: DMEM (complete) for feeder cell and fibroblast plating, Pluriton medium (complete) for modified-mRNA reprogramming, hES medium for maintenance and expansion of picked iPS cell clones on CF-1 feeder cells and mTeSR1 for adaptation and maintenance of iPS cell clones on Matrigel. The complete composition of each medium is detailed in the Reagent Setup section of the protocol. (b) Morphological changes observed during the reprogramming experiment. Areas of morphological changes characteristic of the mesenchymal to epithelial transition are marked with a yellow dashed circles, and emergent colonies are marked with a solid yellow arrows. Scale bars, 100 μm. (c) Live staining of emerging colonies at day 14 of reprogramming showing immunostaining of SSEA-4 and TRA-1-60. Low magnification showing one-fourth of a six-well plate (left) and individual colonies (right). Scale bars, 50 μm. (d) Matrigel-adapted established iPS cell colonies maintained in mTeSR1 medium, immunostained with NANOG and OCT4. Scale bars, 50 μm.
Cell density and cell growth kinetics have a crucial effect on modified mRNA–based reprogramming. This is mostly determined by the intrinsic proliferation potential of different fibroblasts. It is therefore advised to plate cells at varying densities in order to ensure that successful reprogramming can be achieved regardless of the specific cell growth kinetics of the fibroblasts used. The use of antibiotics is not recommended during transfection, and thus sterility should be maintained while handling the cells.
Two very important controls should always be included while performing reprogramming experiments using modified mRNA. (i) Include a control fibroblast line known to be reliably reprogrammable (such as Bj fibroblasts) as a positive control to ensure that all reagents, reprogramming media and mRNA cocktail are working. (ii) Include a negative control well transfected only with modified mRNA encoding a reporter (GFP) in order to monitor modified mRNA or transfection reagent toxicity and observable morphological changes during reprogramming. Modified-mRNA reprogramming requires daily transfection of the reprogramming cocktail, and skipping transfection even for a day is not recommended. We recommend doing the transfection every day at the same time, as this will keep the time interval between transfections constant throughout the experiment.
Materials
REAGENTS
Critical
We recommend using the reagents listed in this protocol for reprogramming; if alternative reagents from other vendors are used they should be tested by the end user before setting up a reprogramming experiment.
Reagents required for IVT
-
KAPA HiFi HotStart ReadyMix, 2× (Kapa Biosystems, cat. no. KK2601)
-
One Shot TOP10 chemically competent Escherichia coli (Invitrogen, cat. no. C4040-03)
-
SOC medium (Life Technologies, cat. no. 15544-034)
-
Difco LB medium, 20 g l−1 in water, autoclaved; (BD, cat. no. 240230)
-
LB agar plates with ampicillin
-
Ampicillin (Sigma, cat. no. A0166)
-
QIAquick PCR purification kit (Qiagen, cat. no. 28106)
-
QIAprep spin miniprep kit (Qiagen, cat. no. 27106)
-
MEGAscript T7 kit (Ambion, cat. no. AM1334)
-
MEGAclear kit (Ambion, cat. no. AM1908)
-
3′-O-Me-m7G(5′)ppp(5′)G RNA cap analog (New England Biolabs, cat. no. S1411S)
-
5-Methylcytidine-5′-triphosphate (Me-CTP; Trilink, cat. no. N1014)
-
Pseudouridine-5′-triphosphate (Pseudo-UTP; Trilink, cat. no. N1019)
-
Antarctic phosphatase (New England Biolabs, cat. no. M0289S)
-
TE buffer, pH 7.0 (Life Technologies, cat. no. AM9861)
-
SeaKam LE agarose (Lonza, cat. no. 50004)
-
Ambion RNaseZap (Life Technologies, cat. no. AM9780)
Plasmids
-
Plasmids encoding human KLF4, c-MYC, OCT4, SOX2, LIN28A and nuclear destabilized EGFP (NDG) in combination with 5′ and 3′ UTRs; these constructs are all available at the Addgene plasmid repository (http://www.addgene.org/Derrick_Rossi). A schematic of all the vectors is shown in Figure 3. The five reprogramming constructs (KMOSL) and reporter construct (NDG) including the T7 promoter, 5′ UTR and 3′ UTR were generated by a splint-ligation method in which the T7 promoter and the 5′ and 3′ UTRs were appended to the respective open reading frames (ORFs) using gene-specific oligonucleotide splints by repetitive cycles of annealing/ligation and PCR amplification22. The T7 promoter-5′UTR-ORF-3′UTR cassettes for each factor were cloned into pcDNA 3.3. Annotated sequence files of all constructs are available in the Supplementary Data.
Figure 3: IVT and quality control of modified mRNA. (a) A schematic representation of the reprogramming constructs (OKMSL) and NDG in pcDNA3.3. Note that the T7 promoter driving mRNA synthesis is upstream of the 5′ UTR. (b) Workflow for modified-mRNA synthesis, with the estimated timeline and critical quality control measures indicated. (c) Addition of poly-(A) tail to template DNA by tail-PCR using primers Xu-F1 and Xu-T120. Gel electrophoresis image showing the size of tailed templates for L, K, M, O and S. (d) Representative NanoDrop reading of a typical IVT preparation (red line), with a poor yield reaction included for comparison (blue line marked by *). (e) Analysis of modified mRNA by Bioanalyzer. L, LIN28A; K, KLF4; M, C-MYC; O, OCT4; S, SOX2; NDG, nuclear destabilized EGFP; Mr, marker (DNA ladder in c, RNA ladder in e).
Primers
-
Xu-F1: 5′-TTGGACCCTCGTACAGAAGCTAATACG-3′ and Xu-T120: 5′-TTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT TTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT TTTTCTTCCTACTCAGGCTTTATTCAAAGACCA-3′. The primers we used were synthesized at and purchased from Integrated DNA Technologies. Primer Xu-T120 was synthesized as Ultamer oligos at a 4-nmol scale.
Reagents required for transfection
-
Opti-MEM I reduced-serum medium (1×) liquid (Invitrogen, cat. no. 31985-070)
-
Lipofectamine RNAiMax (Invitrogen, cat. no. 56532)
-
Stemfect RNA transfection kit (Stemgent, cat. no. 00-0069)
Reagents required for generation of RiPS cells
-
DMEM, high-glucose (Gibco, cat. no. 11965-092)
-
EMEM (ATCC, cat. no. 302003)
-
Defined FBS (Atlas Biologicals, cat. no. F-0500-A)
-
GlutaMAX-1 (Gibco, cat. no. 35050-061)
-
Penicillin-streptomycin (100×; Gibco, cat. no. 15070-063)
-
MEM–non-essential amino acids (NEAA)(Gibco, cat. no. 11140-050)
-
Newborn human foreskin fibroblasts (NuFFs; mitotically inactivated via mitomycin treatment or irradiation; GlobalStem, cat. no. GSC-3001G or GSC-3001M)
-
CF-1 MEF 4M IRR feeder cells (GlobalStem, cat. no. GSC-6001G)
-
PBS without calcium and magnesium (Cellgro, cat. no. 21-040-CV)
-
EmbryoMax 0.1% (wt/vol) gelatin (Millipore, cat. no. ES-006-B)
-
Trypsin-EDTA, 1× (Cellgro, cat. no. 25-052-CI)
-
TrypLE Select (Invitrogen, cat. no. 12563-011)
-
ROCK inhibitor Y27632 (Stemgent, cat. no. 04-0012)
-
Stemgent Bj fibroblasts (early passage, p6; Stemgent, cat. no. 08-0027)
-
B18R recombinant protein carrier-free (eBioscience, cat. no. 34-8185-85)
-
DMEM-F12 (Stem Cell Technologies, cat. no. 36254)
-
mTeSR1 kit (Stem Cell Technologies, cat. no. 05850)
-
mFreSR (Stem Cell Technologies, cat. no. 05855)
-
KnockOut serum replacement (Life Technologies, cat. no. 10828-028)
-
2-Mercaptoethanol (Sigma, cat. no. M7522)
-
hFGF2 (Gemini, cat. no. 400-432P)
-
Pluriton (Stemgent, cat. no. 00-0070)
-
BD Matrigel, human embryonic stem cell (hESC)-qualified matrix (BD, cat. no. 354277)
-
Rabbit anti-mouse/human OCT4 antibody (Stemgent, cat. no. 09-0023; 1:1,000 for immunoblotting, 1:500 for immunofluorescence)
-
Rabbit anti-mouse/human SOX2 antibody (Stemgent, cat. no. 09-0024; 1:1,000 for immunoblotting, 1:500 for immunofluorescence)
-
Mouse anti-mouse/human KLF4 antibody (Stemgent, cat. no. 09-0021; 1:500 for immunofluorescence)
-
Mouse anti-human c-Myc antibody (Stemgent, cat. no. 09-0032; 1:500 for immunofluorescence)
-
Mouse anti-human LIN28A antibody (Stemgent, cat. no. 09-0022; 1:500 for immunofluorescence)
-
Alexa Fluor 555 anti-human SSEA-4 antibody (BD, cat. no. 560218; 1:100 for live staining)
-
Alexa Fluor 647 anti-human Tra-1-60 antibody (BD, cat. no. 560122; 1:75 for live staining)
-
Alexa Fluor 488 mouse anti-human Nanog antibody (BD, cat. no. 560791; 1:500 for immunofluorescence)
-
DyLight649 anti-rabbit IgG antibody (Jackson Immunoreagent, cat. no. 711-495-152; 1:1,000 for immunofluorescence)
-
Alexa Fluor-488 anti-mouse IgG antibody (Jackson Immunoreagent, cat. no. 715-546-151; 1:1,000 for immunofluorescence)
-
Goat anti-rabbit IgG HRP antibody (Abcam, cat. no. ab6721-1; 1:2,000)
-
Tissue culture plates, six-well (BD Falcon, cat. no. 353046)
-
Tissue culture plates, 12-well (BD Falcon, cat. no. 353043)
-
DNase- and RNase-free 1.5-ml sterile microcentrifuge tubes
-
RNase-free sterile 15-ml conical tubes
-
RNase-free sterile 50-ml conical tubes
-
RNase-free sterile aerosol-barrier tips (10, 20, 200 and 1,000 μl)
EQUIPMENT
-
NanoDrop spectrophotometer or equivalent
-
PCR thermocycler (Bio-Rad) or equivalent
-
Microfuge centrifuge (Eppendorf) or equivalent
-
Vortex (VWR) or equivalent
-
Thermomixer (Eppendorf) or equivalent
-
Gel apparatus for electrophoresis (Bio-Rad) or equivalent
-
Bioanalyzer (Agilent Technologies) or equivalent
-
Cell incubator, CO2 (5%) and O2 (5%) (Thermo Scientific) or equivalent
-
Fluorescence microscope (Nikon Eclipse Ti) or equivalent
REAGENT SETUP
Complete DMEM medium for fibroblast and feeder cell plating and maintenance
-
Combine 500 ml of 1× DMEM, 50 ml of FBS, 5 ml of GlutaMAX, 5 ml of MEM-NEAA and 5 ml of penicillin-streptomycin. Store the medium at 4 °C. The medium should be used within 1 month.
Pluriton complete medium for modified-mRNA transfection and reprogramming
-
Combine 50 ml of Pluriton basal medium, 20 μl of 2,500× supplement and 20 μl of B18R (stock 500 μg ml−1).
Critical
Freshly make up the medium in small aliquots according to the needs of the experiment, as B18R is unstable. We typically use the medium for 2–3 d only.
hES medium for maintenance of iPS cells on CF-1 feeder cells
-
Combine 200 ml of DMEM-F12, 50 ml of KnockOut serum replacement, 2.5 ml of GlutaMAX (200 mM stock), 2.5 ml of MEM-NEAA (10 mM stock), 1.75 μl of 2-mercaptoethanol (14.3 M stock) and 257.5 μl of hFGF2 (10 ng μl−1 stock). hES medium should be stored at 4 °C and used within 3–4 weeks.
mTeSR1 medium for maintenance of iPS cells on Matrigel
-
Combine 400 ml of mTeSR1 basal medium and 100 ml of mTeSR1 5× supplement. Store the reconstituted mTeSR1 at 4 °C and use it within 2–3 weeks.
Procedure
Transformation of competent bacteria and isolation of required plasmids
-
1
Before starting the transformation, prewarm SOC medium and ampicillin plates to 37 °C and set the temperature of the water bath/heating block to 42 °C. Thaw One Shot TOP10 chemically competent E. coli vials from −80 °C on ice.
-
2
Add 10 pg–100 ng of plasmids containing reprogramming and reporter constructs separately to the competent cells and mix gently. Incubate the mixture for 30 min on ice.
-
3
Heat-shock the bacteria for 30 s at 42 °C in a heating block or water bath without shaking.
-
4
Immediately place them on ice after heat-shock treatment for 2 min.
-
5
Add 250 μl of prewarmed SOC medium to the cells. Incubate the cells at 37 °C for 45 min–1 h with constant shaking at 200–300 r.p.m. After incubation, plate them on prewarmed agar plates containing ampicillin (50–100 μg ml−1). Place the plates in a 37 °C incubator until colonies become visible (usually overnight).
-
6
Inoculate a single colony from each plate into 15 ml of LB medium containing 50–100 μg of ampicillin per ml, and incubate it at 37 °C overnight with constant shaking at 200 r.p.m. in a bacterial incubator.
-
7
Isolate the plasmid DNAs by standard mini-prep, according to the manufacturer's instructions. We use the QIAprep spin miniprep kit, which provides enough DNA for template synthesis. However, if multiple reprogramming experiments are planned, a maxi-prep should be done.
Pause point
Isolated plasmids can be stored at −20 °C for prolonged time.
Linearization of plasmids for tail PCR
-
8
Digest plasmids containing the reprogramming factors with restriction enzymes that cut once in the vector backbone (SpeI or SalI, Fig. 3a) to produce a linearized vector that can be used as the template for the poly-(A) tail-PCR. The purpose of linearization is to eliminate circular templates that could potentially generate run-on transcripts during the IVT reaction. Set up a 50-μl restriction digestion containing the following components:
Table 2
Component
Amount
Final concentration
Plasmid
5 μg
500 ng μl−1
Buffer 4, 10×
5 μl
1 ×
Suitable restriction enzyme (e.g., SpeI)
5 U
0.1 U μl−1
BSA, 100×
0.5 μl
1 ×
DNase-free water
Made up to 50 μl
-
9
Mix the reaction and incubate it for 2 h at 37 °C.
-
10
Analyze a small aliquot of digested mix by gel electrophoresis and check for complete digestion of plasmid.
Critical Step
The plasmid should be digested to completion. After incubation, take a small aliquot (10 μl) and run it on an agarose gel (1%, wt/vol). The undigested plasmid should be run in another lane as a control.
-
11
Heat-inactivate the enzyme by incubating it at 80 °C for 20 min.
-
12
Purify the reaction mix through a PCR purification column.
Pause point
The linearized plasmid can be stored at −20 °C for several months and can be used for template generation.
Addition of poly-(A) tail by PCR
-
13
Make the tail PCR master mix (total volume of 200 μl, eight reactions of 25 μl each) as detailed below:
Table 3
Component
Amount (μl)
Final concentration
KAPA PCR ready mix (2×)
100
1×
Xu-F1, 10 μM
6
0.3 μM
Xu-T120, 10 μM
6
0.3 μM
Water
80
Digested ORF plasmid, 1–10 ng μl−1
8
40–400 pg μl−1
-
14
Aliquot the tail PCR master mix into eight PCR tubes (25 μl each).
Critical Step
Carry out tail PCR in multiple tubes. A minimum of eight reactions is required for sufficient yield of the tailed product, which can be used in 5–10 IVT reactions.
-
15
Run the tail PCR using the following conditions:
Table 4
Cycle number
Denature
Anneal
Extend
1
95 °C, 2–3 min
2–31
98 °C, 20 s
60 °C, 15 s
72 °C, 60 s
32
72 °C, 3 min
Critical Step
Conditions may vary depending upon the DNA polymerase used.
-
16
Check the quality of PCR products by gel electrophoresis.
-
17
Use PCR purification kits to purify the reactions.
-
18
Adjust the final concentration of the tailed template to 100 ng μl−1.
Critical Step
Check the purity of the tail PCR products by running small aliquots on a 1% (wt/vol) agarose gel (Fig. 3c).
Pause point
The tailed template can be stored at −20 °C for a long time.
In vitro transcription (40-μl reaction volume)
-
19
Clean the working area, pipettes and so on with RNaseZap solution and thaw the required reagents (NTPs, cap analog, IVT reaction buffers and so on) on ice.
Critical Step
Clean the lab bench and pipettes with an RNase decontamination solution (e.g., Ambion RNaseZap solution) and use RNase-free pipette tips. Frequent glove changes are recommended.
-
20
Prepare the custom NTP mix as outlined below. Spin down one vial of cap analog before proceeding. Reconstitute the cap analog with 21.8 μl of DNase- and RNase-free water (concentration upon reconstitution = 60 mM).
Add the following in order, and then mix the contents thoroughly by vortexing and spin down briefly:
Table 5
Component
Stock solution (mM)
Volume (μl) per IVT reaction
Final concentration (mM)
3′-O-Me-m7G cap analog (NEB)
60
4.0
6.0
GTP (from MEGAscript T7 kit)
75
0.8
1.5
ATP (from MEGAscript T7 kit)
75
4.0
7.5
Me-CTP (from Trilink)
100
3.0
7.5
Pseudo-UTP (from Trilink)
100
3.0
7.5
-
21
Mix the reagents (in a PCR tube) in the order listed below (indicated volumes for one IVT reaction). Mix the contents thoroughly by vortexing and spin down briefly.
Table 6
Component
Amount (μl)
Final concentration
DNase/RNase-free water
1.2
Custom NTP mix (from Step 20)
14.8
Tailed PCR product, 100 ng μl−1
16.0
40 ng μl−1
T7 buffer, 10× (from MEGAscript T7 kit)
4.0
1×
T7 enzyme mix, 10× (from MEGAscript T7 kit)
4.0
1×
Critical Step
To carry out one reprogramming experiment, we usually set up 3–5 reactions for OCT4, 2–3 reactions each for KLF4, c-MYC, SOX2 and NDG and 1–2 reaction(s) for LIN28A. This can be scaled up depending upon the scale of the experiment.
-
22
Incubate the reaction at 37 °C for 3–6 h in a thermocycler or a dry air incubator.
Critical Step
Do not incubate the reaction in a water bath in order to avoid contamination with RNases.
-
23
Add 2 μl of Turbo DNase (from MEGAscript T7 kit) to each sample. Mix it gently and incubate it at 37 °C for 15 min.
-
24
Purify the DNase-treated reaction using the MEGAclear kit as per the manufacturer's instructions; elute the modified mRNA with a total of 100 μl of elution buffer (twice with 50 μl of elution buffer).
Critical Step
We follow RNA elution option 2 (preheating elution buffer to 95 °C) as described in MEGAclear kit manual.
Phosphatase treatment of purified modified mRNA
-
25
To each sample (∼100 μl), add 11 μl of 10× Antarctic phosphatase buffer and then add 2 μl of Antarctic phosphatase; gently mix the samples and incubate them at 37 °C for 30 min–1 h.
Critical Step
Phosphatase treatment of uncapped RNA removes 5′ triphosphates and prevents recognition by the RIG-I complex; therefore, the phosphatase treatment is critical.
-
26
Purify Antarctic phosphatase–treated reaction using MEGAclear kits as described in Step 20.
Pause point
Purified modified mRNA can be stored at −80 °C for several months.
Quality control of modified mRNA
-
27
Measure the concentration of the modified mRNA in a NanoDrop spectrophotometer after elution. The expected total yield should be ∼50 μg (range of 30–70 μg; 300–700 ng μl−1 in 100 μl elution volume from one 40 μl IVT reaction). Adjust the concentration to 100 ng μl−1 by adding elution buffer or TE buffer (pH 7.0).
Critical Step
Pay attention to the NanoDrop reading as one of the quality control guidelines (Fig. 3d). The ratio of absorbance at 260 nm/280 nm (A260/A280) should be more than 1.8 (usually between 1.8 and 2.0 or more). The ratio of A260/A230 should be approaching 2.0. Values closer to 2.0 indicate purity.
Critical Step
A yield below 200 ng μl−1 indicates a suboptimal yield, and thus the reaction should be done again.
-
28
Monitor modified-mRNA integrity. We recommend using the microfluidics-based Bioanalyzer platform using an Agilent RNA 6000 nano kit (if available) (Fig. 3e). In the absence of a Bioanalyzer, one should run a small aliquot of synthesized modified mRNA on a denaturing agarose gel to check integrity.
Critical Step
It is very important to verify the quality of in vitro–transcribed modified mRNA before starting a reprogramming experiment. We highly recommend transfecting the modified mRNA into cells and evaluating translation of the encoded protein by western blot analysis or immunocytochemistry (Fig. 1).
Critical Step
Although one can store the modified mRNA after IVT at −20 °C overnight, we recommend purification on the same day.
Preparing a reprogramming cocktail
-
29
Ensure that the concentration of the modified mRNAs generated from the tailed template by IVT as described in detail in Steps 1–27 are adjusted to 100 ng μl−1 for each factor.
-
30
Prepare a modified-mRNA cocktail by mixing each of the factors (OCT4, KLF4, c-MYC, SOX2, LIN28A and NDG) in a molar ratio of 3:1:1:1:1:1, respectively. To prepare the modified-mRNA cocktail, combine 420 μl of OCT4, 170 μl of KLF4, 160 μl of c-MYC, 130 μl of SOX2, 90 μl of LIN28A and 120 μl of NDG. Preadjusting the concentration of each modified mRNA to 100 ng μl−1 before making the cocktail and mixing each factor in the prescribed volumes results in a molar ratio of 3:1:1:1:1:1, respectively.
Critical Step
Use RNase-free sterile 1.5-ml microcentrifuge tubes and aerosol-barrier tips. The modified-mRNA cocktail should be aliquotted into single-use aliquots required for daily transfection (dependent upon the scale of the experiment) and stored at −80 °C. Multiple freeze/thaw cycles should be avoided. Always keep modified mRNA on ice during handling.
Pause point
Purified modified mRNA can be stored individually or as a mixed cocktail at −80 °C for couple of months before starting the reprogramming experiment.
Reprogramming human fibroblasts
-
31
Coat the tissue culture plate with 0.1% (wt/vol) gelatin (1 ml per well of a six-well plate) or 500 μl per well of a 12-well plate) for at least 1 h at room temperature (20–25 °C). Alternatively, the tissue culture plate can be coated with 0.1% (wt/vol) gelatin at 4 °C overnight, 1 d before plating human NuFF feeder cells. Remove the gelatin by aspiration and let the plate dry at room temperature.
-
32
Plate human NuFF feeder cells (day –2). Thaw one vial of mitotically inactivated NuFFs and plate the cells on gelatin-coated plates (at a density of 3–4 × 105 cells per well of six-well plate, or 1–1.5 × 105 cells per well of 12-well plate). Both plate types work well. The advantage of the second option is that the smaller well format conserves reagents. One vial of feeder cells is enough for plating two 6-well or four 12-well plates. Incubate the cells overnight at 37 °C in a 5% O2 incubator.
Critical Step
Ensure that the feeder cells are uniformly distributed and that they cover the entire plate after plating.
-
33
On day −1, plate the fibroblasts (to be reprogrammed) on the feeder cells, preferably in the morning. If you are using a six-well plate format, plate the cells at varying densities such as 5 × 103, 1 × 104, 2.5 × 104 and 5.0 × 104 per well on NuFF feeder cells. If you are using a 12-well format, plate the cells at varying densities such as 5 × 103, 1.5 × 104 and 4.5 × 104 per well on NuFF feeder cells.
Critical Step
We recommend including a control fibroblast line (Bj human fibroblasts) that is reliably reprogrammed to pluripotency by modified mRNA. We plate these cells at a density of 2.5 × 104 cells per well (six-well format).
Critical Step
We recommend carrying wells throughout the reprogramming experiment that are transfected only with modified mRNA encoding NDG. This will allow morphological changes observed in wells transfected with the reprogramming cocktail to be readily identified through comparison.
-
34
At 6–12 h after plating target fibroblasts, replace the fibroblast medium with Pluriton complete reprogramming medium (Pluriton medium supplemented with 2500× supplements and B18R (200 ng ml−1)). Use 2 ml per well for a six-well format or 1 ml per well for a 12-well format. If the cells were plated in the morning, change the medium in the evening, and then incubate the cells overnight at 37 °C in a 5% CO2, 5% O2 incubator. The cells should be preconditioned for at least 9–12 h in reprogramming medium before transfection.
Critical Step
When you are performing modified-mRNA transfection experiments, antibiotics are not added to the medium. Therefore, care must be taken to ensure that sterility is not breached while handling cells.
Critical Step
Although we always conduct reprogramming experiments using incubators that permit low (5%) O2 tension, we have previously successfully reprogrammed fibroblasts with somewhat reduced efficacy in standard incubators without control of O2 tension22.
-
35
Transfect target cells (day 0): After testing many transfection reagents, the reagents that are compatible with modified-mRNA reprogramming in our hands are Lipofectamine RNAiMax (option A) and Stemfect (option B). Unlike Lipofectamine RNAiMax, Stemfect transfection reagent allows longer incubation time (12–16 h) without affecting cell viability. If transfecting with Stemfect, we therefore transfect the cells in the evening (last thing before leaving the laboratory) and change the medium in the morning (first thing after arriving).
-
A
Transfection using Lipofectamine RNAiMax
-
i
Thaw 10 μl of the modified-mRNA cocktail (100 ng μl−1). To this, add 40 μl of OPTI-MEM. Mix the contents gently by repeated pipetting. Do not vortex the mixture.
-
ii
In a separate tube, add 45 μl of OPTI-MEM. To this, add 5 μl of Lipofectamine RNAiMax. Mix the contents gently by repeated pipetting.
-
iii
Add the diluted Lipofectamine RNAiMax to the diluted modified mRNA and mix the contents gently by repeated pipetting. Do not vortex the mixture.
-
iv
Incubate the mixture at room temperature for 15 min to allow the modified mRNA and transfection reagent to complex.
-
v
Transfect one well of a six-well plate with 100 μl of modified-mRNA/transfection reagent complex (use 50 μl per well in a 12-well plate) by adding the mix to the cells in a dropwise fashion in a circular pattern moving from the periphery to the center of the well. This ensures uniform distribution of the transfection complex throughout the well.
-
vi
Gently rock the plate sideways and back and forth to ensure uniform spreading of the transfection complex.
-
vii
Incubate the plate for 4 h at 37 °C, 5% CO2, 5% O2.
Critical Step
Incubation times longer than 4 h may result in cytotoxicity, adversely affecting the outcome of the experiment.
-
i
-
B
Transfection using the Stemfect RNA transfection kit
-
i
Aliquot 60 μl of Stemfect buffer into each of two 1.5-ml microfuge tubes. Mark them A and B.
-
ii
In tube A, add 10 μl of modified-mRNA cocktail (100 ng μl−1). Mix the contents by gentle pipetting.
-
iii
In tube B, add 4 μl of Stemfect transfection reagent. Mix the contents by gentle pipetting.
-
iv
Add the contents of tube B to tube A and mix by gentle pipetting.
-
v
Incubate the mixture at room temperature for 10–15 min.
-
vi
Add 134 μl of the mixture from Step 35B(v) to one well of a six-well plate (or 67 μl of mix to one well of 12-well plate) in a dropwise fashion in a circular pattern moving from the periphery to the center of the well. This ensures uniform distribution of the transfection complex throughout the well.
-
vii
Gently rock the plate sideways and back and forth to ensure uniform spreading of the transfection complex.
-
viii
Incubate the plate for 12–16 h at 37 °C, 5% CO2, 5% O2.
Critical Step
Ensure that the transfection complex is uniformly distributed; otherwise, it may kill the cells in places of high concentration. We do not incubate the cells with Stemfect longer than 16 h.
-
i
Critical Step
Before starting transfection, bring all the reagents including the modified-mRNA cocktail to room temperature. Clean the laminar flow hood and maintain sterility.
Critical Step
Volumes stated are the reagents required for 1 μg of modified mRNA, which is sufficient for transfection of one well of a 6-well plate or two wells of a 12-well plate. The volumes given in this step should be scaled up according to the number of wells to be transfected.
-
A
-
36
During transfection, make the reprogramming medium by mixing 2500× supplements with Pluriton basal medium and adding the required amount of B18R. Equilibrate the reprogramming medium in a 5% O2 incubator for 4 h at 37 °C.
-
37
After the indicated time of transfection, aspirate the medium containing the transfection mix, and wash the well with 2 ml of PBS without calcium and magnesium. Add 2 ml of complete reprogramming medium to the cells, and return the plate to the incubator. Incubate the cells until 24 h has passed since the cells were incubated with transfection medium.
Critical Step
While washing or changing the medium, do not add the solution forcefully to the cells, as this may lead to the detachment of cells. Gently tilt the plate and add the solution to the wall of the wells rather on the cell directly. When handling multiple six-well plates, handle one plate at a time and avoid drying while changing the medium.
Critical Step
When you are performing modified-mRNA transfection experiments, antibiotics should not be added to the medium. Therefore, care must be taken to ensure that sterility is not breached while handling cells.
-
38
Repeat Step 35 through 37 at 24-h intervals after the previous transfection six times; i.e., from day 1 to day 5 after the first transfection. By day 5 or 6, observable morphological changes in the cells transfected with the reprogramming factors should be evident in comparison with control wells transfected with NDG modified mRNA (Fig. 2b). Whereas NDG-transfected wells should contain cells with typical elongated fibroblast morphology (Fig. 2b, upper panel), KMOSL-transfected wells should contain patches of cells with epithelioid morphology indicative of the mesenchymal to epithelial transition (Fig. 2b).
-
39
On day 5 of the experiment, plate NuFF feeder cells at a density of 3–4 × 104 cells per well on six-well plates as indicated in Steps 31 and 32.
-
40
On day 6, split and transfect the cells as described in Box 1.
Critical Step
The morphological changes observed at day 6 are not evident on passaging, and it usually takes a couple of days to be able to see clusters of cells with morphological changes indicative of emergent colonies of reprogrammed cells (Fig. 2b).
-
41
Repeat the transfection protocol on each of the next 7–10 d. As colonies start to emerge (usually evident around day 14 and onward) (Fig. 2b), we suggest performing live staining for the pluripotency markers Tra-1-60/Tra-1-81 and SSEA-4 on one well of a given experiment (Fig. 2c), as described in Step 42. Positive staining for these markers on emerging colonies indicates that the experiment is likely to yield fully reprogrammed iPS cells. If live staining is not performed, proceed to Step 45.
Critical Step
We do not recommend performing live staining on all the wells of an experiment, because the procedure increases the chances of contamination, and it can have an adverse impact on the experiment if the cells are kept out of the incubator for too long.
-
42
For live staining, wash the wells once with PBS. Make up antibody dilutions in prewarmed Pluriton basal medium. Incubate cells with anti-human SSEA-4 antibody (1:100) and anti-human Tra-1-60 antibody (1:75) for 1 h in the incubator. Wash the plate once with PBS and replenish it with fresh medium.
-
43
Image the well at low magnification (×4–10). If colony picking is not the desired end point, DNA staining dye (Hoechst, 1:2,000 dilution) can be included.
-
44
Once colonies that stain positively for live pluripotency markers emerge (Fig. 2c) and have typical iPS cell morphology (tight clusters with sharply boundaries; this typically occurs on days 14–17), modified-mRNA transfection can be stopped. After this point, allow 2–3 d for the emerging colonies to grow, but continue to change the medium everyday during this phase (days 15–19). B18R supplementation is not required during this phase.
Critical Step
Continued transfection of wells for a few days after colonies emerge does not have an adverse effect on reprogramming and may be useful for completing reprogramming of partially reprogrammed colonies.
Critical Step
In the event that there are no emerging colonies by day 18, we have found that splitting the cells and plating them on fresh feeder cells followed by several more days of transfection can yield reprogrammed iPS cell colonies that can be picked and established.
-
45
Manually pick colonies showing tight boundaries and typical hES/iPS cell morphology (typically days 18–20). We recommend picking five to ten clones from each experimental setting. Most clones are stable and can be expanded easily. Typically, we maintain freshly picked clones on irradiated mouse embryonic fibroblasts (CF-1 MEFs) in hES cell medium.
-
46
Passage the cells on CF-1 feeders every 4–5 d, depending on the growth of cells, by using a passaging protocol for hES/iPS cells described elsewhere. During the first few passages, manually pick the best growing colonies from each clone for passaging and expansion.
-
47
Once clones are established, immunostaining for a panel of pluripotency markers is recommended (Fig. 2d; also see ref. 22).
-
48
Expand established clones to six-well plates and freeze them in mFreSR medium as per the manufacturer's recommended protocol. One subconfluent well from a six-well plate should be frozen in one cryovial (1.5–2.0 ml). Established clones can also be adapted to and cultivated on Matrigel-coated plates in mTeSR1 medium as per standard hES maintenance protocols35.
Troubleshooting
Troubleshooting advice can be found in Table 1.
Timing
As a guideline, an estimated time required for each steps is outlined below; steps at which one can stop and resume are noted in the PROCEDURE as Pause Points.
RNA synthesis and purification: 2–3 d
Steps 1–5, transformation of bacteria: 2 h
Step 6, propagation of bacteria: 12 h (overnight)
Step 7, isolation of plasmids: ∼2 h (depending on the method used and the number of samples)
Steps 8 and 9, digestion of plasmids: 2 h
Step 10, confirming linearization by gel electrophoresis: 1–2 h
Step 11, heat inactivation of restriction enzyme: 20 min
Step 12, purification of linearized plasmids: 30 min
Steps 13 and 14, setting up a tail PCR: 15–30 min
Step 15, tail PCR: 2 h
Step 16, checking the tailed product on gel: 1–2 h
Step 17, purification of the tailed template: 30 min–1 h
Step 18, measuring and adjusting the concentration: 30 min–1 h
Step 19, cleaning working area, pipette and so on and thawing the reagents for IVT: 15 min
Step 20, making NTPs master mix for IVT: 15 min
Step 21, setting up IVT: 30 min–1 h
Step 22, IVT reaction: 3–6 h
Step 23, DNase treatment, 15 min
Step 24, RNA purification (first purification): 30 min–2 h (depending on the number of reactions)
Step 25, phosphatase treatment: 30 min
Step 26, RNA purification (second purification), 30 min–2 h (depending on the number of reactions)
Step 27, measuring and adjusting modified-mRNA concentration: 30 min
Step 28, testing RNA quality: 2–3 d. Gel electrophoresis, Bioanalyzer, western blotting/immunostaining
Reprogramming of human fibroblasts to iPS cells: ∼3 weeks
Steps 29 and 30, making modified-mRNA reprogramming cocktail
Step 31, coating tissue culture plate with gelatin: 1 h (overnight unattended)
Step 32, day −2, plating of NuFF feeder cells: 30 min
Step 33, day −1, plating of target fibroblasts: 30 min–1 h
Step 34, preconditioning of cells with complete reprogramming medium: overnight
Step 35, day 0, setting up transfection and adding the transfection complex to the cells: 30 min–1 h
Transfection with modified mRNA (unattended): 4 h (option A) or overnight (option B)
Step 36, making complete reprogramming medium (10 min) and equilibrating it with low oxygen: 4 h
Step 37, medium change: 30 min–1 h
Step 38, through days 1–6, repeat Steps 35–37 at 24-h intervals
Step 39, plating of NuFF feeder cells on day 5: 30 min
Step 40, day 6, daily transfection, 30 min, and splitting of cells at day 6: 1–2 h
Step 41, days 7–14, repeat Steps 35 through 37 for 7–10 d: 30 min–1 h
Step 42, live staining for pluripotency markers (SSEA-4 and Tra-1-60) on emerging colonies: 3 h
Step 43, imaging of stained colonies: 1–2 h
Steps 44 and 45, days 15–18, manual picking of iPS cell colonies: 1–2 h
Step 46, expansion and maintenance of picked clones on CF-1 feeder cells: 1–2 h
Step 47, immunostaining for pluripotency markers on established clones: 3–4 h
Step 48, freezing the expanded clones: 30 min–1 h
Note: cells should be fed every day with fresh medium during the expansion phase
Anticipated results
The expanded clones should be positive for endogenous pluripotency markers (e.g., OCT4, Nanog, Tra-1-60, Tra-1-80/81) and should show typical ES/iPS cell morphology (Fig. 2c,d). The events encountered during the course of the reprogramming experiment are shown in Figure 4 for reference. The efficiency of reprogramming may vary depending upon the origin, proliferative potential and genetic background of different primary fibroblasts; however, a successful modified-mRNA reprogramming experiment typically yields several hundred colonies. We recommend picking five to ten colonies in order to ensure establishment of multiple independent clones for each primary fibroblast line reprogrammed.
(a–i) Images showing the most commonly encountered events during the course of a reprogramming experiment. Their causes and possible explanations are described in the Troubleshooting section. (a) Overgrowth on plate. (b) Rare feeder cell density. (c) Holes in cell monolayer. (d) Cell clumps after passaging or transformation event. (e) Emerging colonies growing under the feeder layer (yellow arrows). Note that, in this field, a normal emerging colony is marked with a red arrow. (f) Cluster of loosely packed cells. (g) Partially reprogrammed colony. (h) Emerging colony with a sharp boundary, but probably not fully reprogrammed. (i) Typical iPS cell colony. Scale bars, 100 μm.
Change history
20 March 2013
In the version of this article initially published, acknowledgment of the technical assistance of Andrew Ettenger was omitted. The error has been corrected in the HTML and PDF versions of the article.
References
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Park, I.H. et al. Disease-specific induced pluripotent stem cells. Cell 134, 877–886 (2008).
Grskovic, M., Javaherian, A., Strulovici, B. & Daley, G.Q. Induced pluripotent stem cells–opportunities for disease modelling and drug discovery. Nat. Rev. Drug Discov. 10, 915–929 (2011).
Robinton, D.A. & Daley, G.Q. The promise of induced pluripotent stem cells in research and therapy. Nature 481, 295–305 (2012).
Stadtfeld, M., Nagaya, M., Utikal, J., Weir, G. & Hochedlinger, K. Induced pluripotent stem cells generated without viral integration. Science 322, 945–949 (2008).
Yu, J. et al. Human induced pluripotent stem cells free of vector and transgene sequences. Science 324, 797–801 (2009).
Gonzalez, F. et al. Generation of mouse-induced pluripotent stem cells by transient expression of a single nonviral polycistronic vector. Proc. Natl. Acad. Sci. USA 106, 8918–8922 (2009).
Hu, K. et al. Efficient generation of transgene-free induced pluripotent stem cells from normal and neoplastic bone marrow and cord blood mononuclear cells. Blood 117, e109–e119 (2011).
Jia, F. et al. A nonviral minicircle vector for deriving human iPS cells. Nat. Methods 7, 197–199 (2010).
Narsinh, K.H. et al. Generation of adult human induced pluripotent stem cells using nonviral minicircle DNA vectors. Nat. Protoc. 6, 78–88 (2011).
Okita, K. et al. A more efficient method to generate integration-free human iPS cells. Nat. Methods 8, 409–412 (2011).
Okita, K., Nakagawa, M., Hyenjong, H., Ichisaka, T. & Yamanaka, S. Generation of mouse induced pluripotent stem cells without viral vectors. Science 322, 949–953 (2008).
Si-Tayeb, K. et al. Generation of human induced pluripotent stem cells by simple transient transfection of plasmid DNA encoding reprogramming factors. BMC Dev. Biol. 10, 81 (2010).
Woltjen, K. et al. piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 458, 766–770 (2009).
Yu, J., Chau, K.F., Vodyanik, M.A., Jiang, J. & Jiang, Y. Efficient feeder-free episomal reprogramming with small molecules. PLoS ONE 6, e17557 (2011).
Yusa, K., Rad, R., Takeda, J. & Bradley, A. Generation of transgene-free induced pluripotent mouse stem cells by the piggyBac transposon. Nat. Methods 6, 363–369 (2009).
Kim, D. et al. Generation of human induced pluripotent stem cells by direct delivery of reprogramming proteins. Cell Stem Cell 4, 472–476 (2009).
Zhou, H. et al. Generation of induced pluripotent stem cells using recombinant proteins. Cell Stem Cell 4, 381–384 (2009).
Fusaki, N., Ban, H., Nishiyama, A., Saeki, K. & Hasegawa, M. Efficient induction of transgene-free human pluripotent stem cells using a vector based on Sendai virus, an RNA virus that does not integrate into the host genome. Proc. Jpn Acad. Ser. B Phys. Biol. Sci. 85, 348–362 (2009).
Ban, H. et al. Efficient generation of transgene-free human induced pluripotent stem cells (iPSCs) by temperature-sensitive Sendai virus vectors. Proc. Natl. Acad. Sci. USA 108, 14234–14239 (2011).
Warren, L. et al. Highly efficient reprogramming to pluripotency and directed differentiation of human cells with synthetic modified mRNA. Cell Stem Cell 7, 618–630 (2010).
Diebold, S.S., Kaisho, T., Hemmi, H., Akira, S. & Reis e Sousa, C. Innate antiviral responses by means of TLR7-mediated recognition of single-stranded RNA. Science 303, 1529–1531 (2004).
Hornung, V. et al. 5′-Triphosphate RNA is the ligand for RIG-I. Science 314, 994–997 (2006).
Pichlmair, A. et al. RIG-I-mediated antiviral responses to single-stranded RNA bearing 5′-phosphates. Science 314, 997–1001 (2006).
Kariko, K., Buckstein, M., Ni, H. & Weissman, D. Suppression of RNA recognition by Toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA. Immunity 23, 165–175 (2005).
Angel, M. & Yanik, M.F. Innate immune suppression enables frequent transfection with RNA encoding reprogramming proteins. PLoS ONE 5, e11756 (2010).
Kariko, K. & Weissman, D. Naturally occurring nucleoside modifications suppress the immunostimulatory activity of RNA: implication for therapeutic RNA development. Curr. Opin Drug Discov. Devel. 10, 523–532 (2007).
Kariko, K. et al. Incorporation of pseudouridine into mRNA yields superior nonimmunogenic vector with increased translational capacity and biological stability. Mol. Ther. 16, 1833–1840 (2008).
Anderson, B.R. et al. Incorporation of pseudouridine into mRNA enhances translation by diminishing PKR activation. Nucleic Acids Res. 38, 5884–5892 (2010).
Liptakova, H., Kontsekova, E., Alcami, A., Smith, G.L. & Kontsek, P. Analysis of an interaction between the soluble vaccinia virus-coded type I interferon (IFN)-receptor and human IFN-α1 and IFN-α2. Virology 232, 86–90 (1997).
Kormann, M.S. et al. Expression of therapeutic proteins after delivery of chemically modified mRNA in mice. Nat. Biotechnol. 29, 154–157 (2011).
Kariko, K., Muramatsu, H., Keller, J.M. & Weissman, D. Increased erythropoiesis in mice injected with submicrogram quantities of pseudouridine-containing mRNA encoding erythropoietin. Mol. Ther. 20, 948–953 (2012).
Warren, L., Ni, Y., Wang, J. & Guo, X. Feeder-free derivation of human induced pluripotent stem cells with messenger RNA. Scientific Reports 2, 657 doi:10.1038/srep00657 (2012).
McElroy, S.L. & Reijo Pera, R.A. Culturing human embryonic stem cells in feeder-free conditions. Cold Spring Harb. Protoc. 2008, doi:10.1101/pdb.prot5044 (2008).
Acknowledgements
We thank T. Schlaeger, L. Daheron, W. Ebina and L. Zhangi for their valuable suggestions and discussion, and L. Warren and P. Manos for past contributions. We thank A. Ettenger for technical assistance. This work was funded in part by a grant from the Harvard Stem Cell Institute. D.J.R. is a New York Stem Cell Foundation Robertson Investigator.
Author information
Authors and Affiliations
Contributions
P.K.M. and D.J.R. designed the experiments. P.K.M. performed the experiments. P.K.M. and D.J.R. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
D.J.R. is a cofounder of ModeRNA Therapeutics, a Cambridge, Massachusetts–based biotechnology company that is exploring the therapeutic potential of modified mRNA.
Supplementary information
Supplementary Data
Annotated sequence files of reprogramming factors (KLF4, c-MYC, OCT4, SOX2 and LIN28A) and NDG (PDF 710 kb)
Rights and permissions
About this article
Cite this article
Mandal, P., Rossi, D. Reprogramming human fibroblasts to pluripotency using modified mRNA. Nat Protoc 8, 568–582 (2013). https://doi.org/10.1038/nprot.2013.019
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.019
This article is cited by
-
A TRIM21-based bioPROTAC highlights the therapeutic benefit of HuR degradation
Nature Communications (2023)
-
Rapid differentiation of hiPSCs into functional oligodendrocytes using an OLIG2 synthetic modified messenger RNA
Communications Biology (2022)
-
Loss of the fragile X syndrome protein FMRP results in misregulation of nonsense-mediated mRNA decay
Nature Cell Biology (2021)
-
TSC patient-derived isogenic neural progenitor cells reveal altered early neurodevelopmental phenotypes and rapamycin-induced MNK-eIF4E signaling
Molecular Autism (2020)
-
Human adipose-derived stem cells enriched with VEGF-modified mRNA promote angiogenesis and long-term graft survival in a fat graft transplantation model
Stem Cell Research & Therapy (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.