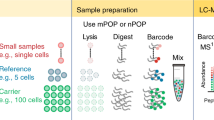
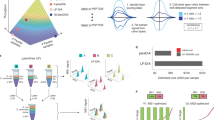
Abstract
Targeted quantification of proteins is a daily task in biological research but often relies on techniques such as western blotting that are only barely quantitative. Here we present a broadly applicable workflow for protein quantification from unpurified whole-cell extracts that can be completed in less than 3 d. Without prefractionation or affinity enrichment, a whole-cell extract is trypsin-digested in an acetonitrile-containing ammonium carbonate buffer and high-molecular-weight compounds are removed by filtration. A normalization strategy, which involves endogenous reference proteins, facilitates the determination of relative changes in protein expression without requiring isotope labeling or standard addition. On a triple-quadrupole mass spectrometer, we demonstrate standard-free quantification of yeast proteins present over five orders of magnitude and present at ≥500 copies per cell. Liquid chromatography/multiple reaction monitoring (LC-MRM)–based proteomics is therefore a next-generation alternative to western blotting, as it allows simultaneous and reliable quantification of multiple endogenous proteins without the need for enrichment, isotope labeling or use of antibodies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Walsh, C.T., Garneau-Tsodikova, S. & Gatto, G.J. Jr. Protein posttranslational modifications: the chemistry of proteome diversifications. Angew. Chem. Int. Ed 44, 7342–7372 (2005).
Towbin, H., Staehelin, T. & Gordon, J. Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc. Natl Acad. Sci. USA 76, 4350–4354 (1979).
Engvall, E. & Perlmann, P. Enzyme-linked immunosorbent assay (ELISA). Quantitative assay of immunoglobulin G. Immunochemistry 8, 871–874 (1971).
Westermeier, R. & Gronau, S. Electrophoresis in Practice: A Guide to Methods and Applications of DNA and Protein Separations 3rd edn. (Wiley-VCH, 2001).
Voshol, H., Ehrat, M., Traenkle, J., Bertrand, E. & van Oostrum, J. Antibody-based proteomics: analysis of signaling networks using reverse protein arrays. FEBS J. 276, 6871–6879 (2009).
Mann, M. Can proteomics retire the western blot? J. Proteome Res. 7, 3065 (2008).
Kiyonami, R. et al. Increased selectivity, analytical precision, and throughput in targeted proteomics. Mol. Cell Proteomics 10, M110.002931 (2011).
Wamelink, M.M. et al. The difference between rare and exceptionally rare: molecular characterization of ribose 5-phosphate isomerase deficiency. J. Mol. Med. 88, 931–939 (2010).
Timmermann, B. et al. A new dominant peroxiredoxin allele identified by whole-genome resequencing of random mutagenized yeast causes oxidant-resistance and premature aging. Aging 2, 475–486 (2010).
Addona, T.A. et al. Multi-site assessment of the precision and reproducibility of multiple reaction monitoring-based measurements of proteins in plasma. Nat. Biotechnol. 27, 633–641 (2009).
Gaspari, M., Abbonante, V. & Cuda, G. Gel-free sample preparation for the nanoscale LC-MS/MS analysis and identification of low-nanogram protein samples. J. Sep. Sci. 30, 2210–2216 (2007).
Yeung, Y.G., Nieves, E., Angeletti, R.H. & Stanley, E.R. Removal of detergents from protein digests for mass spectrometry analysis. Anal. Biochem. 382, 135–137 (2008).
Strader, M.B., Tabb, D.L., Hervey, W.J., Pan, C. & Hurst, G.B. Efficient and specific trypsin digestion of microgram to nanogram quantities of proteins in organic-aqueous solvent systems. Anal. Chem. 78, 125–134 (2006).
Russell, W.K., Park, Z.Y. & Russell, D.H. Proteolysis in mixed organic-aqueous solvent systems: applications for peptide mass mapping using mass spectrometry. Anal. Chem. 73, 2682–2685 (2001).
Slysz, G.W. & Schriemer, D.C. On-column digestion of proteins in aqueous-organic solvents. Rapid Commun. Mass Spectrom. 17, 1044–1050 (2003).
Simon, L.M., László, K., Vértesi, A., Bagi, K. & Szajáni, B. Stability of hydrolytic enzymes in water-organic solvent systems. J. Mol. Catal. B Enzym. 4, 41–45 (1998).
van Midwoud, P.M., Rieux, L., Bischoff, R., Verpoorte, E. & Niederlander, H.A. Improvement of recovery and repeatability in liquid chromatography-mass spectrometry analysis of peptides. J. Proteome Res. 6, 781–791 (2007).
Wisniewski, J.R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Lange, V., Picotti, P., Domon, B. & Aebersold, R. Selected reaction monitoring for quantitative proteomics: a tutorial. Mol. Syst. Biol. 4, 222 (2008).
Kettenbach, A.N., Rush, J. & Gerber, S.A. Absolute quantification of protein and post-translational modification abundance with stable isotope-labeled synthetic peptides. Nat. Protoc. 6, 175–186 (2011).
Chen, Y. et al. Quantification of beta-catenin signaling components in colon cancer cell lines, tissue sections, and microdissected tumor cells using reaction monitoring mass spectrometry. J. Proteome Res. 9, 4215–4227 (2010).
Wolf-Yadlin, A., Hautaniemi, S., Lauffenburger, D.A. & White, F.M. Multiple reaction monitoring for robust quantitative proteomic analysis of cellular signaling networks. Proc. Natl Acad. Sci. USA 104, 5860–5865 (2007).
Ciccimaro, E., Hanks, S.K., Yu, K.H. & Blair, I.A. Absolute quantification of phosphorylation on the kinase activation loop of cellular focal adhesion kinase by stable isotope dilution liquid chromatography/mass spectrometry. Anal. Chem. 81, 3304–3313 (2009).
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003).
Picotti, P., Bodenmiller, B., Mueller, L.N., Domon, B. & Aebersold, R. Full dynamic range proteome analysis of S. cerevisiae by targeted proteomics. Cell 138, 795–806 (2009).
Warringer, J. & Blomberg, A. Evolutionary constraints on yeast protein size. BMC Evol. Biol. 6, 61 (2006).
Cagney, G., Amiri, S., Premawaradena, T., Lindo, M. & Emili, A. In silico proteome analysis to facilitate proteomics experiments using mass spectrometry. Proteome Sci. 1, 5 (2003).
Ong, S.E. et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell Proteomics 1, 376–386 (2002).
Gerber, S.A., Rush, J., Stemman, O., Kirschner, M.W. & Gygi, S.P. Absolute quantification of proteins and phosphoproteins from cell lysates by tandem MS. Proc. Natl Acad. Sci. USA 100, 6940–6945 (2003).
Hopfgartner, G. et al. Triple quadrupole linear ion trap mass spectrometer for the analysis of small molecules and macromolecules. J. Mass Spectrom. 39, 845–855 (2004).
Kirkpatrick, D.S., Gerber, S.A. & Gygi, S.P. The absolute quantification strategy: a general procedure for the quantification of proteins and post-translational modifications. Methods 35, 265–273 (2005).
Gerber, S.A., Kettenbach, A.N., Rush, J. & Gygi, S.P. The absolute quantification strategy: application to phosphorylation profiling of human separase serine 1126. Methods Mol. Biol. 359, 71–86 (2007).
Lohr, D., Venkov, P. & Zlatanova, J. Transcriptional regulation in the yeast GAL gene family: a complex genetic network. FASEB J. 9, 777–787 (1995).
Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, e45 (2001).
Anderson, N.L. et al. Mass spectrometric quantitation of peptides and proteins using Stable Isotope Standards and Capture by Anti-Peptide Antibodies (SISCAPA). J. Proteome Res. 3, 235–244 (2004).
Unwin, R.D., Griffiths, J.R. & Whetton, A.D. A sensitive mass spectrometric method for hypothesis-driven detection of peptide post-translational modifications: multiple reaction monitoring-initiated detection and sequencing (MIDAS). Nat. Protoc. 4, 870–877 (2009).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Prakash, A. et al. Expediting the development of targeted SRM assays: using data from shotgun proteomics to automate method development. J. Proteome Res. 8, 2733–2739 (2009).
Reinert, K. & Kohlbacher, O. OpenMS and TOPP: open source software for LC-MS data analysis. Methods Mol. Biol. 604, 201–211 (2010).
Deutsch, E.W., Lam, H. & Aebersold, R. PeptideAtlas: a resource for target selection for emerging targeted proteomics workflows. EMBO Rep. 9, 429–434 (2008).
Picotti, P. et al. A database of mass spectrometric assays for the yeast proteome. Nat. Methods 5, 913–914 (2008).
Fusaro, V.A., Mani, D.R., Mesirov, J.P. & Carr, S.A. Prediction of high-responding peptides for targeted protein assays by mass spectrometry. Nat. Biotechnol. 27, 190–198 (2009).
Page, J.S., Kelly, R.T., Tang, K. & Smith, R.D. Ionization and transmission efficiency in an electrospray ionization-mass spectrometry interface. J. Am. Soc. Mass Spectrom. 18, 1582–1590 (2007).
Acknowledgements
We thank B. Lukaszewska-McGreal for help with sample preparation, our lab members for critical discussions and the Max Planck Institute for Molecular Genetics for funding. M.R. is a Wellcome Trust Research Career Development and Wellcome-Beit prize Fellow.
Author information
Authors and Affiliations
Contributions
K.B. designed and conducted experiments and contributed to the writing of the paper. M.R. designed and conducted experiments and contributed to the writing of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures 1 and 2
Supplementary Figure 1: Relative protein abundance in incomplete tryptic digests (PDF 644 kb)
Supplementary Figure 2: Detection of Cps1p in high concentrated protein extract.
Rights and permissions
About this article
Cite this article
Bluemlein, K., Ralser, M. Monitoring protein expression in whole-cell extracts by targeted label- and standard-free LC-MS/MS. Nat Protoc 6, 859–869 (2011). https://doi.org/10.1038/nprot.2011.333
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2011.333
This article is cited by
-
Alzheimer’s disease progression characterized by alterations in the molecular profiles and biogenesis of brain extracellular vesicles
Alzheimer's Research & Therapy (2020)
-
Cadmium stress dictates central carbon flux and alters membrane composition in Streptococcus pneumoniae
Communications Biology (2020)
-
Identification of aldolase A as a potential diagnostic biomarker for colorectal cancer based on proteomic analysis using formalin-fixed paraffin-embedded tissue
Tumor Biology (2016)
-
Tpo1‐mediated spermine and spermidine export controls cell cycle delay and times antioxidant protein expression during the oxidative stress response
EMBO reports (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.