Abstract
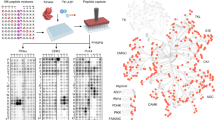
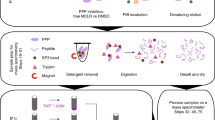
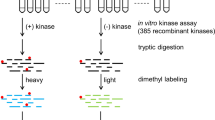
Protein phosphorylation and dephosphorylation play an important role in regulation of intracellular signal transduction pathways in the biological system. A key step in the biological characterization of phosphatases and their use as drug targets is the identification of their cellular partners and suitable substrates for potential inhibitor development. Herein we describe a microarray-based protocol to map the substrate specificity of protein Ser/Thr phosphatases. This protocol uses Pro-Q dye to sensitively and quantitatively detect the amount of dephosphorylation that occurs from many putative peptide substrates in parallel, and therefore could be used to generate the so-called peptide substrate fingerprints as well as detailed kinetic information of a target phosphatase. Excluding the synthesis of the peptide substrates, the whole protocol takes a total of 11 h to complete and in future can be readily extended to the study of other classes of phosphatases, i.e., protein tyrosine phosphatases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Manning, G., Whyte, D.B., Martinez, R., Hunter, T. & Sudarsanam, S. The protein kinase complement of the human genome. Science 298, 1912–1934 (2002).
Zhang, Z.Y. Protein-tyrosine phosphatases: biological function, structural characteristics, and mechanism of catalysis. CRC Crit. Rev. Biochem. Mol. Biol. 33, 1–52 (1998).
Johnson, T.O., Ermolieff, J. & Jirousek, M.R. Protein tyrosine phosphatase 1B inhibitors for diabetes. Nat. Rev. Drug Disc. 1, 696–709 (2002).
Bialy, L. & Waldmann, H. Inhibitors of protein tyrosine phosphatases: next-generation drugs? Angew. Chem. Int. Ed. 44, 3814–3839 (2005).
Pawson, T. & Scott, J.D. Signaling through scaffold, anchoring, and adaptor proteins. Science 278, 2075–2080 (1997).
Schutkowski, M. et al. High-content peptide microarrays for deciphering kinase specificity and biology. Angew. Chem. Int. Ed. 43, 2671–2674 (2004).
Garaud, M. & Pei, D. Substrate profiling of protein tyrosine phosphatase PTP1B by screening a combinatorial peptide library. J. Am. Chem. Soc. 129, 5366–5367 (2007).
Cho, J. et al. Substrate specificities of catalytic fragments of protein-tyrosine phosphatases (Hptp-Beta, Lar, and Cd45) toward phosphotyrosylpeptide substrates and thiophosphotyrosylated peptides as inhibitors. Protein Sci. 2, 977–984 (1993).
Zhang, Z.-Y., Maclean, D., McNamara, D.J., Sawyer, T.K. & Dixon, J.E. Protein tyrosine phosphatase substrate specificity: size and phosphotyrosine positioning requirements in peptide substrates. Biochemistry 33, 2285–2290 (1994).
Pellegrini, M.C. et al. Mapping the subsite preferences of protein tyrosine phosphatase PTP-1B using combinatorial chemistry approaches. Biochemistry 37, 15598–15606 (1998).
Walchli, S., Espanel, X., Harrenga, A., Rossi, M., Cesareni, G. & van Huijsduiinen, R.H. Probing protein-tyrosine phosphatase substrate specificity using a phosphotyrosine-containing phage library. J. Biol. Chem. 279, 311–318 (2004).
Vetter, S.W., Keng, Y.-F., Lawrence, D.S. & Zhang, Z.-Y. Assessment of protein-tyrosine phosphatase 1B substrate specificity using 'inverse alanine scanning'. J. Biol. Chem. 275, 2265–2268 (2000).
Tomizaki, K.Y., Usui, K. & Mihara, H. Protein-detecting microarrays: current accomplishments and requirements. ChemBioChem 6, 782–799 (2005).
Reineke, U., Volkmer-Engert, R. & Schneider-Mergener, J. Applications of peptide arrays prepared by the SPOT-technology. Curr. Opin. Biotechnol. 12, 59–64 (2001).
Houseman, B.T., Huh, J.H., Kron, S.J. & Mrksich, M. Peptide chips for the quantitative evaluation of protein kinase activity. Nat. Biotechnol. 20, 270–274 (2002).
Falsey, J.R., Renil, M., Park, S., Li, S. & Lam, K.S. Peptide and small molecule microarray for high throughput cell adhesion and functional assays. Bioconj. Chem. 12, 346–353 (2001).
Lesaicherre, M.L., Uttamchandani, M., Chen, G.Y. & Yao, S.Q. Antibody-based fluorescence detection of kinase activity on a peptide array. Bioorg. Med. Chem. Lett. 12, 2085–2088 (2002).
Salisbury, C.M., Maly, D.J. & Ellman, J.A. Peptide microarrays for the determination of protease substrate specificity. J. Am. Chem. Soc. 124, 14868–14870 (2002).
Köhn, M. et al. A microarray strategy for mapping the substrate specificity of protein tyrosine phosphatase. Angew. Chem. Int. Ed. 46, 7700–7703 (2007).
Sun, H., Lu, C.H.S., Uttamchandani, M., Xia, Y., Liou, Y.-C. & Yao, S.Q. Peptide microarray for high-throughput determination of phosphatase specificity and biology. Angew. Chem. Int. Ed. 47, 1698–1702 (2008).
Martin, K., Steinberg, T.H., Cooley, L.A., Gee, K.R., Beechem, J.M. & Patton, W.F. Quantitative analysis of protein phosphorylation status and protein kinase activity on microarrays using a novel fluorescent phosphorylation sensor dye. Proteomics 3, 1244–1255 (2003).
Lu, K.P., Liou, Y.C. & Zhou, X.Z. Pinning down proline-directed phosphorylation signaling. Trends Cell Biol. 12, 164–172 (2002).
Chattopadhaya, S., Tan, L.P. & Yao, S.Q. Strategies for site-specific protein biotinylation using in vitro, in vivo and cell-free systems: toward functional protein arrays. Nat. Protoc. 1, 2386–2398 (2006).
Lee, W.L., Li, J., Uttamchandani, M., Sun, H. & Yao, S.Q. Inhibitor fingerprinting of metalloproteases using microplate and microarray platforms—an enabling technology in catalomics. Nat. Protoc. 2, 2126–2138 (2007).
Uttamchandani, M., Lee, W.L., Wang, J. & Yao, S.Q. Quantitative inhibitor fingerprinting of metalloproteases using small molecule microarrays. J. Am. Chem. Soc. 129, 13110–13117 (2007).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sun, H., Lu, C., Shi, H. et al. Peptide microarrays for high-throughput studies of Ser/Thr phosphatases. Nat Protoc 3, 1485–1493 (2008). https://doi.org/10.1038/nprot.2008.139
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2008.139
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.