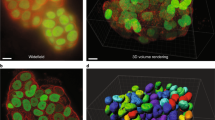
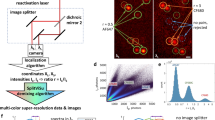
Abstract
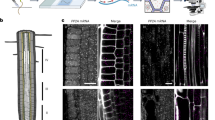
This protocol describes the steps needed to perform quantitative statistical colocalization on two-color confocal images, specifically of plant cells. The procedure includes a calibration test to check the chromatic alignment of the confocal microscope. A software tool is provided to calculate the Pearson and Spearman correlation coefficients ('Pearson–Spearman correlation colocalization' ImageJ plug-in) across regions of interest within the image. Steps are included to help the user practice using the software. The result is a quantitative estimate of the amount of colocalization in the images. Manual masking takes about 1–15 min per image, depending on the detail required, and calculating the correlation coefficients is almost instantaneous. Examples of suitable dyes for such two-color colocalization include Oregon Green or Alexa Fluor 488 dyes in the green range (excited with 488-nm laser line) and Alexa Fluor 555 dye in the red range (excited with 543-nm laser line).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
References
North, A.J. Seeing is believing? A beginner's guide to practical pitfalls in image acquisition. J. Cell Biol., 172, 9–18 (2006).
Bolte, S. & Cordelières, F.P. A guided tour into subcellular colocalization in light microscopy. J. Microsc., 224, 213–232 (2006).
Costes, S.V., Daelemans, D., Cho, E.H., Dobbin, Z. & Pavlakis, G. Automatic and quantitative measurement of protein–protein colocalization in live cells. Biophys. J., 86, 3993–4003 (2004).
Manders, E.M.M., Stap, J., Brakenhoff, G.J., Van Driel, R. & Aten, J.A. Dynamics of three-dimensional replication patterns during the S-phase, analysed by double labelling of DNA and confocal microscopy. J. Cell Sci., 103, 857–862 (1992).
Lachmanovich, E. et al. Co-localization analysis of complex formation among membrane proteins by computerized fluorescence microscopy: application to immunofluorescence co-patching studies. J. Microsc., 212, 122–131 (2003).
Glynn, M.W. & McAllister, A.K. Immunocytochemistry and quantification of protein colocalization in cultured neurons. Nat. Protoc., 1, 1287–1296 (2006).
Heintzmann, R. & Cremer, C. Axial tomographic confocal fluorescence microscopy. J. Microsc., 206, 7–23 (2002).
Dharmasiri, S. et al. AXR4 is required for localization of the auxin influx facilitator AUX1. Science, 312, 1218–1220 (2006).
Abramoff, M.D., Magelhaes, P.J. & Ram, S.J. Image processing with ImageJ. Biophotonics Int., 11, 36–42 (2004).
Zucker, R.M. & Price, O. Evaluation of confocal microscopy system performance. Cytometry, 44, 273–294 (2001).
Manders, E.M.M. Chromatic shift in multicolour confocal microscopy. J. Microsc., 185, 321–328 (1997).
Pawley, J. The 39 steps: a cautionary tale of quantitative 3-D fluorescence microscopy. Biotechniques, 28, 884–888 (2000).
Pawley, J. Handbook of Confocal Microscopy 3rd edn (Springer Science, New York, 2006).
Smallcombe, A. Multicolor imaging: the important question of co-localization. Biotechniques, 30, 1240–1245 (2001).
Kass, M., Witkin, A. & Terzopolous, D. Snakes: active contour models. Int. J. Comp. Vis., 1, 321–331 (1988).
Friml, J., Benkova, E., Mayer, U., Palme, K. & Muster, G. Automated whole mount localisation techniques for plant seedlings. Plant J., 34, 115–124 (2003).
Sauer, M., Paciorek, T., Benkova, E. & Friml, J. Immunocytochemical techniques for whole-mount in-situ protein localization in plants. Nat. Protoc., 1, 98–103 (2006).
Centonze, V. & Pawley, J. Tutorial on practical confocal microscopy and use of the confocal test specimen. In Handbook of Biological Confocal Microscopy 3rd edn. (ed. Pawley J.) 627–649 (Springer Science and Business Media, New York, 2006).
Swarup, R. et al. Structure–function analysis of the presumptive Arabidopsis auxin permease AUX1. Plant Cell, 16, 3069–3083 (2004).
Li, Q. et al. A Syntaxin 1, Gαo, and N-type calcium channel complex at a presynaptic nerve terminal: analyses by quantitative immunocolocalization. J. Neurosci., 24, 4070–4081 (2004).
Chinga, G. & Syverud, K. Quantification of paper mass distributions within local picking areas. Nordic Pulp Paper Res. J., 22, 441–446 (2007).
Acknowledgements
We acknowledge EPSRC and BBSRC CISB programme funding to CPIB and funding from the Belgian Federal IUAPVI programme to M.J.B. and R.S. as part of the BARN consortium.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
French, A., Mills, S., Swarup, R. et al. Colocalization of fluorescent markers in confocal microscope images of plant cells. Nat Protoc 3, 619–628 (2008). https://doi.org/10.1038/nprot.2008.31
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2008.31
This article is cited by
-
Lignin impairs Cel7A degradation of in vitro lignified cellulose by impeding enzyme movement and not by acting as a sink
Biotechnology for Biofuels and Bioproducts (2024)
-
Light-independent phytoplankton degradation and detoxification of methylmercury in water
Nature Water (2023)
-
A non-canonical role of ATG8 in Golgi recovery from heat stress in plants
Nature Plants (2023)
-
The K/HDEL receptor does not recycle but instead acts as a Golgi-gatekeeper
Nature Communications (2023)
-
Ex vivo observation of granulocyte activity during thrombus formation
BMC Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.