Abstract
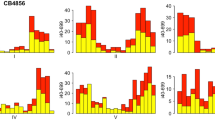
Positional cloning of chemically induced mutations is the rate-limiting step in forward genetic screens in Drosophila. Single-nucleotide polymorphisms (SNPs) are useful markers to locate a mutated region in the genome. Here, we provide a protocol for high-throughput, high-resolution SNP mapping that enables rapid and cost-effective positional cloning in Drosophila. In stage 1 of the protocol, we use highly multiplexed tag-array mini-sequencing assays to map mutations to an interval of 1–2 Mb. In these assays, SNPs are genotyped by primer extension using fluorescently labeled dideoxy-nucleotides. Fluorescent primers are captured and detected on a microarray. In stage 2, we selectively isolate recombinants within the identified 1–2 Mb interval for fine mapping of mutations to about 50 kb. We have previously demonstrated the applicability of this protocol by mapping 14 muscle morphogenesis mutants within 4 months, which represents a significant acceleration compared with other commonly used mapping strategies that may take years.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Jorgensen, E.M. & Mango, S.E. The art and design of genetic screens: Caenorhabditis elegans . Nat. Rev. Genet. 3, 356–369 (2002).
St Johnston, D. The art and design of genetic screens: Drosophila melanogaster . Nat. Rev. Genet. 3, 176–188 (2002).
van Eeden, F.J., Granato, M., Odenthal, J. & Haffter, P. Developmental mutant screens in the zebrafish. Methods Cell Biol. 60, 21–41 (1999).
Davis, M.W. et al. Rapid single nucleotide polymorphism mapping in C. elegans . BMC Genomics 6, 118 (2005).
Berger, J. et al. Genetic mapping with SNP markers in Drosophila . Nat. Genet. 29, 475–481 (2001).
Stickney, H.L. et al. Rapid mapping of zebrafish mutations with SNPs and oligonucleotide microarrays. Genome Res. 12, 1929–1934 (2002).
Hoskins, R.A. et al. Single nucleotide polymorphism markers for genetic mapping in Drosophila melanogaster . Genome Res. 11, 1100–1113 (2001).
Chen, D. et al. High-resolution, high-throughput SNP mapping in Drosophila melanogaster . Nat Methods 5, 323–328 (2008).
Baner, J. et al. Parallel gene analysis with allele-specific padlock probes and tag microarrays. Nucleic Acids Res. 31, e103 (2003).
Shen, R. et al. High-throughput SNP genotyping on universal bead arrays. Mutat. Res. 573, 70–82 (2005).
Macdonald, S.J., Pastinen, T., Genissel, A., Cornforth, T.W. & Long, A.D. A low-cost open-source SNP genotyping platform for association mapping applications. Genome Biol. 6, R105 (2005).
Thibault, S.T. et al. A complementary transposon tool kit for Drosophila melanogaster using P and piggyBac. Nat. Genet. 36, 283–287 (2004).
Parks, A.L. et al. Systematic generation of high-resolution deletion coverage of the Drosophila melanogaster genome. Nat. Genet. 36, 288–292 (2004).
Ryder, E. et al. The DrosDel deletion collection: a Drosophila genomewide chromosomal deficiency resource. Genetics 177, 615–629 (2007).
Lindroos, K., Sigurdsson, S., Johansson, K., Ronnblom, L. & Syvanen, A.C. Multiplex SNP genotyping in pooled DNA samples by a four-colour microarray system. Nucleic Acids Res. 30, e70 (2002).
Lovmar, L., Fredriksson, M., Liljedahl, U., Sigurdsson, S. & Syvanen, A.C. Quantitative evaluation by minisequencing and microarrays reveals accurate multiplexed SNP genotyping of whole genome amplified DNA. Nucleic Acids Res. 31, e129 (2003).
Fuger, P., Behrends, L.B., Mertel, S., Sigrist, S.J. & Rasse, T.M. Live imaging of synapse development and measuring protein dynamics using two-color fluorescence recovery after photo-bleaching at Drosophila synapses. Nat. Protoc. 2, 3285–3298 (2007).
Wieschaus, E. & Nüsslein-Volhard, C. In Drosophila, a Practical Approach (ed. Roberts, D.B.) (Oxford Science Publications, Oxford University Press, 1998).
Casso, D., Ramirez-Weber, F.A. & Kornberg, T.B. GFP-tagged balancer chromosomes for Drosophila melanogaster . Mech. Dev. 88, 229–232 (1999).
Schnorrer, F., Kalchhauser, I. & Dickson, B.J. The transmembrane protein Kon-tiki couples to Dgrip to mediate myotube targeting in Drosophila . Dev. Cell 12, 751–766 (2007).
Xu, T. & Rubin, G.M. Analysis of genetic mosaics in developing and adult Drosophila tissues. Development 117, 1223–1237 (1993).
Spradling, A.C. et al. The Berkeley Drosophila genome project gene disruption project: single P-element insertions mutating 25% of vital Drosophila genes. Genetics 153, 135–177 (1999).
Vallone, P.M. & Butler, J.M. AutoDimer: a screening tool for primer-dimer and hairpin structures. Biotechniques 37, 226–231 (2004).
Kaderali, L., Deshpande, A., Nolan, J.P. & White, P.S. Primer-design for multiplexed genotyping. Nucleic Acids Res. 31, 1796–1802 (2003).
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Data
Supplementary Data | An Excel file with example SNPmapper input files. Explanatory comments of the format are provided. (XLS 58 kb)
Rights and permissions
About this article
Cite this article
Schnorrer, F., Ahlford, A., Chen, D. et al. Positional cloning by fast-track SNP-mapping in Drosophila melanogaster. Nat Protoc 3, 1751–1765 (2008). https://doi.org/10.1038/nprot.2008.175
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2008.175
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.