Abstract
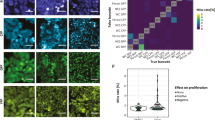
We present a protocol to tag proteins expressed from their endogenous chromosomal locations in individual mammalian cells using central dogma tagging. The protocol can be used to build libraries of cell clones, each expressing one endogenous protein tagged with a fluorophore such as the yellow fluorescent protein. Each round of library generation produces 100–200 cell clones and takes about 1 month. The protocol integrates procedures for high-throughput single-cell cloning using flow cytometry, high-throughput cDNA generation and 3′ rapid amplification of cDNA ends, semi-automatic protein localization screening using fluorescent microscopy and freezing cells in 96-well format.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
20 September 2007
In the version of this article initially published, the authors omitted the following acknowledgment: “We thank the Kahn Family Foundation and the Israel Science Foundation for support.” The acknowledgment has been added to the PDF version of the article.
References
Alon, U. An Introduction to Systems Biology: Design Principles of Biological Circuits (Chapman & Hall/CRC press, Boca Raton, FL, 2006).
Andersen, J.S. et al. Nucleolar proteome dynamics. Nature 433, 77–83 (2005).
Zhu, H. et al. Global analysis of protein activities using proteome chips. Science 293, 2101–2105 (2001).
Ho, Y. et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature 415, 180–183 (2002).
Gavin, A.C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 415, 141–147 (2002).
Butland, G. et al. Interaction network containing conserved and essential protein complexes in Escherichia coli . Nature 433, 531–537 (2005).
Yi, E.C. et al. Increased quantitative proteome coverage with (13)C/(12)C-based, acid-cleavable isotope-coded affinity tag reagent and modified data acquisition scheme. Proteomics 5, 380–387 (2005).
Whitney, A.R. et al. Individuality and variation in gene expression patterns in human blood. Proc. Natl. Acad. Sci. USA 100, 1896–1901 (2003).
Perlman, Z.E. et al. Multidimensional drug profiling by automated microscopy. Science 306, 1194–1198 (2004).
Mayer, T.U. et al. Small molecule inhibitor of mitotic spindle bipolarity identified in a phenotype-based screen. Science 286, 971–974 (1999).
Chen, D. & Huang, S. Nucleolar components involved in ribosome biogenesis cycle between the nucleolus and nucleoplasm in interphase cells. J. Cell Biol. 153, 169–176 (2001).
Bannasch, D. et al. LIFEdb: a database for functional genomics experiments integrating information from external sources, and serving as a sample tracking system. Nucleic Acids Res. 32 Database issue, D505–D508 (2004).
Simpson, J.C., Wellenreuther, R., Poustka, A., Pepperkok, R. & Wiemann, S. Systematic subcellular localization of novel proteins identified by large-scale cDNA sequencing. EMBO Rep. 1, 287–292 (2000).
Huh, W.K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Jarvik, J.W., Adler, S.A., Telmer, C.A., Subramaniam, V. & Lopez, A.J. CD-tagging: a new approach to gene and protein discovery and analysis. Biotechniques 20, 896–904 (1996).
Jarvik, J.W. et al. In vivo functional proteomics: mammalian genome annotation using CD-tagging. Biotechniques 33, 852–854 856, 858–860 passim (2002).
Jarvik, J.W. & Telmer, C.A. Epitope tagging. Annu. Rev. Genet. 32, 601–618 (1998).
Sigal, A. et al. Dynamic proteomics in individual human cells uncovers widespread cell-cycle dependence of nuclear proteins. Nat. Methods 3, 525–531 (2006).
Friedrich, G. & Soriano, P. Promoter traps in embryonic stem cells: a genetic screen to identify and mutate developmental genes in mice. Genes Dev 5, 1513–1523 (1991).
Gossler, A., Joyner, A.L., Rossant, J. & Skarnes, W.C. Mouse embryonic stem cells and reporter constructs to detect developmentally regulated genes. Science 244, 463–465 (1989).
Stanford, W.L. et al. Expression trapping: identification of novel genes expressed in hematopoietic and endothelial lineages by gene trapping in ES cells. Blood 92, 4622–4631 (1998).
Morin, X., Daneman, R., Zavortink, M. & Chia, W. A protein trap strategy to detect GFP-tagged proteins expressed from their endogenous loci in Drosophila . Proc. Natl. Acad. Sci. USA 98, 15050–15055 (2001).
Clyne, P.J., Brotman, J.S., Sweeney, S.T. & Davis, G. Green fluorescent protein tagging Drosophila proteins at their native genomic loci with small P elements. Genetics 165, 1433–1441 (2003).
Shaner, N.C. et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat. Biotechnol. 22, 1567–1572 (2004).
Sigal, A. et al. Variability and memory of protein levels in human cells. Nature 444, 643–646 (2006).
Wu, X., Li, Y., Crise, B. & Burgess, S.M. Transcription start regions in the human genome are favored targets for MLV integration. Science 300, 1749–1751 (2003).
Kent, W.J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Lander, E.S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Mitchell, R.S. et al. Retroviral DNA integration: ASLV, HIV, and MLV show distinct target site preferences. PLoS Biol. 2, E234 (2004).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Sigal, A., Danon, T., Cohen, A. et al. Generation of a fluorescently labeled endogenous protein library in living human cells. Nat Protoc 2, 1515–1527 (2007). https://doi.org/10.1038/nprot.2007.197
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.197
This article is cited by
-
Spatial proteomics: a powerful discovery tool for cell biology
Nature Reviews Molecular Cell Biology (2019)
-
Dynamic proteomics reveals bimodal protein dynamics of cancer cells in response to HSP90 inhibitor
BMC Systems Biology (2017)
-
Enhanced genome editing in mammalian cells with a modified dual-fluorescent surrogate system
Cellular and Molecular Life Sciences (2016)
-
NetworkPainter: dynamic intracellular pathway animation in Cytobank
BMC Bioinformatics (2015)
-
Studying and modelling dynamic biological processes using time-series gene expression data
Nature Reviews Genetics (2012)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.