Abstract
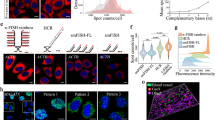
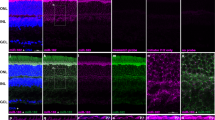
The ability to determine spatial and temporal microRNA (miRNA) accumulation at the tissue, cell and subcellular levels is essential for understanding the biological roles of miRNAs and miRNA-associated gene regulatory networks. This protocol describes a method for fast and effective detection of miRNAs in frozen tissue sections using fluorescence in situ hybridization (FISH). The method combines the unique miRNA recognition properties of locked nucleic acid (LNA)-modified oligonucleotide probes with FISH using the tyramide signal amplification (TSA) technology. Although both approaches have previously been shown to increase detection sensitivity in FISH, combining these techniques into one protocol significantly decreases the time needed for miRNA detection in cryosections, while simultaneously retaining high detection sensitivity. Starting with fixation of the tissue sections, this miRNA FISH protocol can be completed within approximately 6 h and allows miRNA detection in a wide variety of animal tissue cryosections as well as in human tumor biopsies at high cellular resolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bartel, D.P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Wienholds, E. & Plasterk, R.H. MicroRNA function in animal development. FEBS Lett. 579, 5911–5922 (2005).
Kloosterman, W.P. & Plasterk, R.H. The diverse functions of microRNAs in animal development and disease. Dev. Cell. 11, 441–450 (2006).
Brennecke, J., Hipfner, D.R., Stark, A., Russel, R.B. & Cohen, S.M. Bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in Drosophila. Cell 113, 25–36 (2003).
Mansfield, J.H. et al. MicroRNA-responsive 'sensor' transgenes uncover Hox-like and other developmentally regulated patterns of vertebrate microRNA expression. Nat. Genet. 36, 1079–1083 (2004).
Koshkin, A.A. et al. LNA (locked nucleic acids): Synthesis of the adenine, cytosine, guanine, 5-methylcytosine, thymine and uracil bicyclonucleoside monomers, oligomerisation, and unprecedented nucleic acid recognition. Tetrahedron 54, 3607–3630 (1998).
Braasch, D.A. & Corey, D.R. Locked nucleic acid (LNA): fine-tuning the recognition of DNA and RNA. Chem. Biol. 8, 1–7 (2001).
Kurreck, J., Wyszko, E., Gillen, C. & Erdmann, V.A. Design of antisense oligonucleotides stabilized by locked nucleic acids. Nucleic Acids Res. 30, 1911–1918 (2002).
Jacobsen, N. et al. Direct isolation of poly(A)+ RNA from 4 M guanidine thiocyanate-lysed cell extracts using locked nucleic acid-oligo(T) capture. Nucleic Acids Res. 32, e64 (2004).
Kauppinen, S., Vester, B. & Wengel, J. Locked nucleic acid: high-affinity targeting of complementary RNA for RNomics. Handb. Exp. Pharmacol. 173, 405–422 (2006).
Wienholds, E. et al. MicroRNA expression in zebrafish embryonic development. Science 309, 310–311 (2005).
Ason, B. et al. Differences in vertebrate microRNA expression. Proc. Natl. Acad. Sci. USA. 103, 14385–14389 (2006).
Darnell, D.K. et al. MicroRNA expression during chick embryo development. Dev. Dyn. 235, 3156–3165 (2006).
Kloosterman, W.P., Wienholds, E., de Bruijn, E., Kauppinen, S. & Plasterk, R.H. In situ detection of miRNAs in animal embryos using LNA-modified oligonucleotide probes. Nat. Methods 3, 27–29 (2006).
Wheeler, G., Ntounia-Fousara, S., Granda, B., Rathjen, T. & Dalmay, T. Identification of new central nervous system specific mouse microRNAs. FEBS Lett. 580, 2195–2200 (2006).
Nelson, P.T. et al. RAKE and LNA-ISH reveal microRNA expression and localization in archival human brain. RNA 12, 187–191 (2006).
Obernosterer, G., Leuschner, P.J., Alenius, M. & Martinez, J. Post-transcriptional regulation of microRNA expression. RNA 12, 1161–1167 (2006).
Obernosterer, G., Martinez, J. & Alenius, M. Locked nucleic acid-based in situ detection of microRNAs in mouse tissue sections. Nat. Protoc. 2, 1508–1514 (2007).
Silahtaroglu, A.N., Tommerup, N. & Vissing, H. FISHing with locked nucleic acids (LNA): evaluation of different LNA/DNA mixmers. Mol. Cell. Probes 17, 165–169 (2003).
Silahtaroglu, A., Pfundheller, H., Koshkin, A., Tommerup, N. & Kauppinen, S. LNA-modified oligonucleotides are highly efficient as FISH probes. Cytogenet. Genome Res. 107, 32–37 (2004).
Christoffersen, N.R., Silahtaroglu, A., Orom, U.A., Kauppinen, S. & Lund, A.H. miR-200b mediates post-transcriptional repression of ZFHX1B. RNA 13, 1172–1178 (2007).
Kerstens, H.M., Poddighe, P.J. & Hanselaar, A.G. A novel in situ hybridization signal amplification method based on the deposition of biotinylated tyramide. J. Histochem. Cytochem. 43, 347–352 (1995).
Acknowledgements
The authors wish to thank Anna Catharina Lassen, Ha Nguyen, Ursula Rentzmann, Lene Løgstrup and Eniser Zekirovska for their excellent technical assistance. This study was supported by grants from the Danish National Advanced Technology Foundation, Danish Medical Research Council to S.K., Danish Research Agency to A.N.S. and the Lundbeck Foundation to S.K. and A.N.S. Wilhelm Johannsen Centre for Functional Genome Research is established by the Danish National Research Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Silahtaroglu, A., Nolting, D., Dyrskjøt, L. et al. Detection of microRNAs in frozen tissue sections by fluorescence in situ hybridization using locked nucleic acid probes and tyramide signal amplification. Nat Protoc 2, 2520–2528 (2007). https://doi.org/10.1038/nprot.2007.313
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.313
This article is cited by
-
3D in vitro morphogenesis of human intestinal epithelium in a gut-on-a-chip or a hybrid chip with a cell culture insert
Nature Protocols (2022)
-
SQSTM1/p62 promotes miR-198 loading into extracellular vesicles and its autophagy-related secretion
Human Cell (2022)
-
Long transcripts minus touchdown qPCR (LTMT-qPCR): a simplified and convenient method for the screening and quantification of microRNA profile
Laboratory Investigation (2021)
-
miR-182-5p and miR-378a-3p regulate ferroptosis in I/R-induced renal injury
Cell Death & Disease (2020)
-
Fluorescence Signal Amplification Strategies Based on DNA Nanotechnology for miRNA Detection
Chemical Research in Chinese Universities (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.