Abstract
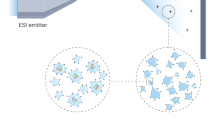
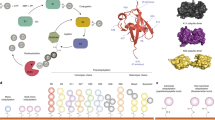
Among the main objectives of biomedical and proteomic research is to identify non-covalent interactions involving proteins. Here we provide a detailed protocol to apply matrix-assisted laser desorption ionization (MALDI) time-of-flight (TOF) mass spectrometry for such a purpose using proteases and protease inhibitors in complex biological samples. Our methodology is based on monitoring the reduction in intensity of inhibitors' mass spectrometric signals when their protease target is added to the MALDI sample. The versatility of the protocol permits the target to be added in a soluble form (direct protocol) or immobilized form (indirect protocol). The 'intensity fading' phenomenon is greatly favored when the binding assay is carried out in the sub-micromolar range and the interacting partners occur in mixtures of non-binding compounds. This protocol can be completed in 10 h, taking 20 or 30 min per sample to perform the mass spectrometric data acquisition, depending on whether a soluble or an immobilized target is used.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Hopkins, A.L. & Groom, C.R. The druggable genome. Nat. Rev. Drug Discov. 1, 727–730 (2002).
Villanueva, J., Yanes, O., Querol, E., Serrano, L. & Aviles, F.X. Identification of protein ligands in complex biological samples using intensity-fading MALDI-TOF mass spectrometry. Anal. Chem. 75, 3385–3395 (2003).
Yanes, O., Aviles, F.X., Roepstorff, P. & Jorgensen, T.J. Exploring the “intensity fading” phenomenon in the study of noncovalent interactions by MALDI-TOF mass spectrometry. J. Am. Soc. Mass Spectrom. in the press (2006).
Yanes, O., Villanueva, J., Querol, E. & Aviles, F.X. Functional screening of serine protease inhibitors in the medical leech Hirudo medicinalis monitored by intensity fading MALDI-TOF MS. Mol. Cell Proteomics 4, 1602–1613 (2005).
Yanes, O., Nazabal, A., Wenzel, R., Zenobi, R. & Aviles, F.X. Detection of noncovalent complexes in biological samples by intensity fading and high-mass detection MALDI-TOF mass spectrometry. J. Proteome Res. 5, 2711–2719 (2006).
Wei, J., Buriak, J.M. & Siuzdak, G. Desorption-ionization mass spectrometry on porous silicon. Nature 399, 243–246 (1999).
Go, E.P. et al. Desorption/ionization on silicon nanowires. Anal. Chem. 77, 1641–1646 (2005).
Berkenkamp, S., Karas, M. & Hillenkamp, F. Ice as a matrix for IR-matrix-assisted laser desorption/ionization: mass spectra from a protein single crystal. Proc. Natl. Acad. Sci. USA 93, 7003–7007 (1996).
Von Seggern, C.E. & Cotter, R.J. Fragmentation studies of noncovalent sugar-sugar complexes by infrared atmospheric pressure MALDI. J. Am. Soc. Mass Spectrom. 14, 1158–1165 (2003).
Von Seggern, C.E. & Cotter, R.J. Study of peptide-sugar non-covalent complexes by infrared atmospheric pressure matrix-assisted laser desorption/ionization. J. Mass Spectrom. 39, 736–742 (2004).
Bolivar, J.M. et al. Improvement of the stability of alcohol dehydrogenase by covalent immobilization on glyoxyl-agarose. J. Biotechnol. 125, 85–94 (2006).
Duckworth, B.P., Xu, J., Taton, T.A., Guo, A. & Distefano, M.D. Site-specific, covalent attachment of proteins to a solid surface. Bioconjug. Chem. 17, 967–974 (2006).
Hodneland, C.D., Lee, Y.S., Min, D.H. & Mrksich, M. Selective immobilization of proteins to self-assembled monolayers presenting active site-directed capture ligands. Proc. Natl. Acad. Sci. USA 99, 5048–5052 (2002).
Ruzicka, J., Carroll, A.D. & Lahdesmaki, I. Immobilization of proteins on agarose beads, monitored in real time by bead injection spectroscopy. Analyst 131, 799–808 (2006).
Krem, M.M. & Di Cera, E. Dissecting substrate recognition by thrombin using the inactive mutant S195A. Biophys. Chem. 100, 315–323 (2003).
Pallares, I. et al. Structure of human carboxypeptidase A4 with its endogenous protein inhibitor, latexin. Proc. Natl. Acad. Sci. USA 102, 3978–3983 (2005).
Viguera, A.R., Arrondo, J.L., Musacchio, A., Saraste, M. & Serrano, L. Characterization of the interaction of natural proline-rich peptides with five different SH3 domains. Biochemistry 33, 10925–10933 (1994).
Villanueva, J. et al. Characterization of the wound-induced metallocarboxypeptidase inhibitor from potato. cDNA sequence, induction of gene expression, subcellular immunolocalization and potential roles of the C-terminal propeptide. FEBS Lett. 440, 175–182 (1998).
Reverter, D. et al. A carboxypeptidase inhibitor from the medical leech Hirudo medicinalis. Isolation, sequence analysis, cDNA cloning, recombinant expression, and characterization. J. Biol. Chem. 273, 32927–32933 (1998).
Barret A., Rawlings, N. & Woessner J. (eds.) Handbook of Proteolytic Enzymes (Academic Press, Elsevier, New York (2004)).
Karas, M. & Hillenkamp, F. Laser desorption ionization of proteins with molecular masses exceeding 10,000 daltons. Anal. Chem. 60, 2299–2301 (1988).
Frank, M. et al. High-efficiency detection of 66,000 Da protein molecules using a cryogenic detector in a matrix-assisted laser desorption/ionization time-of-flight mass spectrometer. Rapid Commun. Mass Spectrom. 10, 1946–1950 (1996).
Twerenbold, D. et al. Single molecule detector for mass spectrometry with mass independent detection efficiency. Proteomics 1, 66–69 (2001).
Acknowledgements
We are very grateful to Professors Luís Serrano (EMBL, Germany), Peter Roepstorff and Thomas J.D. Jørgensen (University of Southern Denmark), Renato Zenobi (ETH Zürich, Switzerland) and M. Angeles Chavez (University of La Habana, Cuba), and their research teams, for their invaluable advice, collaboration and interest in the research and development of different aspects of the intensity fading MS methodology. Also to PROTEORED (Spain). This work has been supported by grants GEN2003-20642-C09-05 and BIO2004-05879 (Ministerio de Ciencia y Tecnología, Spain), grant LSHG-2006-018830-CAMP (EC-Dir.F), and BIO2004-05879, and by the Centre de Referència en Biotecnologia (Generalitat de Catalunya, Spain). O.Y. acknowledges a fellowship from the Ministerio de Ciencia y Tecnología.
Author information
Authors and Affiliations
Contributions
O.Y. and J.V. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Yanes, O., Villanueva, J., Querol, E. et al. Detection of non-covalent protein interactions by 'intensity fading' MALDI-TOF mass spectrometry: applications to proteases and protease inhibitors. Nat Protoc 2, 119–130 (2007). https://doi.org/10.1038/nprot.2006.487
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.487
This article is cited by
-
Macromolecular properties and partial amino acid sequence of a Kunitz-type protease inhibitor from okra (Abelmoschus esculentus) seeds
Journal of Biosciences (2019)
-
Enzyme-Coupled Nanoparticles-Assisted Laser Desorption Ionization Mass Spectrometry for Searching for Low-Mass Inhibitors of Enzymes in Complex Mixtures
Journal of the American Society for Mass Spectrometry (2014)
-
Study of Papain–Cystatin Interaction by Intensity Fading MALDI-TOF-MS
The Protein Journal (2008)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.