Abstract
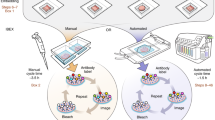
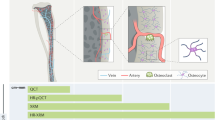
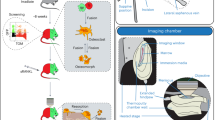
High-resolution confocal imaging is a vital tool for analyzing the 3D architecture and detailed spatial distribution of cells in situ. However, imaging of skeletal tissue has remained technically challenging because of its calcified nature. Here we describe a protocol that allows high-resolution imaging of skeletal tissue with preservation of cellular morphology and tissue architecture. The procedure involves tissue fixation, decalcification and cryosectioning of the mouse skeletal tissue to generate thick sections. The thick sections generated by this procedure are not only compatible with the analysis of genetically expressed fluorescent proteins but they also preserve antigenicity, thus enabling diverse combinations of antibody labeling. Further, this procedure also permits other fluorescence techniques such as TUNEL and ethynyl deoxyuridine (EdU) incorporation assays. Images resulting from the confocal imaging can be assessed qualitatively and quantitatively to analyze various parameters such as distribution and interrelationships of cell types. The technique is straightforward and robust, highly reproducible and can be completed in ∼11 d.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Lieberman, J.R. & Friedlaender, G.E. Bone Regeneration and Repair: Biology and Clinical Applications (Humana Press, 2005).
Favus, M.J., Christakos, S. & American Society for Bone and Mineral Research. Primer on the Metabolic Bone Diseases and Disorders of Mineral Metabolism 3rd edn. (Lippincott-Raven, 1996).
Wei, J. & Karsenty, G. An overview of the metabolic functions of osteocalcin. Rev. Endocr. Metab. Disord. 16, 93–98 (2015).
Ding, L., Saunders, T.L., Enikolopov, G. & Morrison, S.J. Endothelial and perivascular cells maintain haematopoietic stem cells. Nature 481, 457–462 (2012).
Kiel, M.J. et al. SLAM family receptors distinguish hematopoietic stem and progenitor cells and reveal endothelial niches for stem cells. Cell 121, 1109–1121 (2005).
Crisan, M. et al. A perivascular origin for mesenchymal stem cells in multiple human organs. Cell Stem Cell 3, 301–313 (2008).
Shi, S. & Gronthos, S. Perivascular niche of postnatal mesenchymal stem cells in human bone marrow and dental pulp. J. Bone Miner. Res. 18, 696–704 (2003).
Ding, L. & Morrison, S.J. Haematopoietic stem cells and early lymphoid progenitors occupy distinct bone marrow niches. Nature 495, 231–235 (2013).
Hooper, A.T. et al. Engraftment and reconstitution of hematopoiesis is dependent on VEGFR2-mediated regeneration of sinusoidal endothelial cells. Cell Stem Cell 4, 263–274 (2009).
Kusumbe, A.P., Ramasamy, S.K. & Adams, R.H. Coupling of angiogenesis and osteogenesis by a specific vessel subtype in bone. Nature 507, 323–328 (2014).
Ramasamy, S.K., Kusumbe, A.P., Wang, L. & Adams, R.H. Endothelial Notch activity promotes angiogenesis and osteogenesis in bone. Nature 507, 376–380 (2014).
Ramasamy, S.K., Kusumbe, A.P. & Adams, R.H. Regulation of tissue morphogenesis by endothelial cell-derived signals. Trends Cell Biol. 25, 148–157 (2015).
Snippert, H.J., Schepers, A.G., Delconte, G., Siersema, P.D. & Clevers, H. Slide preparation for single-cell-resolution imaging of fluorescent proteins in their three-dimensional near-native environment. Nat. Protoc. 6, 1221–1228 (2011).
Paddock, S.W. Confocal laser-scanning microscopy. Biotechniques 27, 992–996, 998–1002, 1004 (1999).
Idleburg, C., DeLassus, E.N. & Novack, D.V. Immunohistochemistry of skeletal tissues. Methods Mol. Biol. 1226, 87–95 (2015).
Claxton, S. et al. Efficient, inducible Cre-recombinase activation in vascular endothelium. Genesis 46, 74–80 (2008).
Kisanuki, Y.Y. et al. Tie2-Cre transgenic mice: a new model for endothelial cell-lineage analysis in vivo. Dev. Biol. 230, 230–242 (2001).
Wang, Y. et al. Ephrin-B2 controls VEGF-induced angiogenesis and lymphangiogenesis. Nature 465, 483–486 (2010).
Kim, J.E., Nakashima, K. & de Crombrugghe, B. Transgenic mice expressing a ligand-inducible Cre recombinase in osteoblasts and odontoblasts: a new tool to examine physiology and disease of postnatal bone and tooth. Am. J. Pathol. 165, 1875–1882 (2004).
Rodda, S.J. & McMahon, A.P. Distinct roles for Hedgehog and canonical Wnt signaling in specification, differentiation and maintenance of osteoblast progenitors. Development 133, 3231–3244 (2006).
Foo, S.S. et al. Ephrin-B2 controls cell motility and adhesion during blood-vessel-wall assembly. Cell 124, 161–173 (2006).
Kuhbandner, S. et al. Temporally controlled somatic mutagenesis in smooth muscle. Genesis 28, 15–22 (2000).
Mericskay, M. et al. An overlapping CArG/octamer element is required for regulation of desmin gene transcription in arterial smooth muscle cells. Dev. Biol. 226, 192–208 (2000).
Zhu, X., Bergles, D.E. & Nishiyama, A. NG2 cells generate both oligodendrocytes and gray matter astrocytes. Development 135, 145–157 (2008).
Harms, J.F., Budgeon, L.R., Christensen, N.D. & Welch, D.R. Maintaining GFP tissue fluorescence through bone decalcification and long-term storage. Biotechniques 33, 1197–1200 (2002).
Ashwood-Smith, M.J. & Warby, C. Studies on the molecular weight and cryoprotective properties of polyvinylpyrrolidone and dextran with bacteria and erythrocytes. Cryobiology 8, 453–464 (1971).
Bernhard, W. & Leduc, E.H. Ultrathin frozen sections. I. Methods and ultrastructural preservation. J. Cell Biol. 34, 757–771 (1967).
Gavrieli, Y., Sherman, Y. & Ben-Sasson, S.A. Identification of programmed cell death in situ via specific labeling of nuclear DNA fragmentation. J. Cell Biol. 119, 493–501 (1992).
Kapuscinski, J. DAPI: a DNA-specific fluorescent probe. Biotech. Histochem. 70, 220–233 (1995).
Bink, K. et al. TO-PRO-3 is an optimal fluorescent dye for nuclear counterstaining in dual-colour FISH on paraffin sections. Histochem. Cell Biol. 115, 293–299 (2001).
Kolb, H.C., Finn, M.G. & Sharpless, K.B. Click chemistry: diverse chemical function from a few good reactions. Angew. Chem. Int. Ed. Engl. 40, 2004–2021 (2001).
Salic, A. & Mitchison, T.J. chemical method for fast and sensitive detection of DNA synthesis in vivo. Proc. Natl. Acad. Sci. USA 105, 2415–2420 (2008).
Acknowledgements
We thank M. Schiller for technical assistance and S. Volkery for microscopy. Funding was provided by the Max Planck Society, the University of Münster, the Deutsche Forschungsgemeinschaft (DFG) cluster of excellence 'Cells in Motion', and the European Research Council (AdG 339409 AngioBone).
Author information
Authors and Affiliations
Contributions
A.P.K., S.K.R. and R.H.A. designed experiments, contributed to the development of methodology and wrote the paper. A.P.K. and S.K.R. directed A.S. and carried out the experiments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Kusumbe, A., Ramasamy, S., Starsichova, A. et al. Sample preparation for high-resolution 3D confocal imaging of mouse skeletal tissue. Nat Protoc 10, 1904–1914 (2015). https://doi.org/10.1038/nprot.2015.125
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2015.125
This article is cited by
-
Warning regarding hematological toxicity of tamoxifen activated CreERT2 in young Rosa26CreERT2 mice
Scientific Reports (2023)
-
Ebf3+ niche-derived CXCL12 is required for the localization and maintenance of hematopoietic stem cells
Nature Communications (2023)
-
BHLHE40 promotes osteoclastogenesis and abnormal bone resorption via c-Fos/NFATc1
Cell & Bioscience (2022)
-
Spred1 deficit promotes treatment resistance and transformation of chronic phase CML
Leukemia (2022)
-
Mesenchymal stromal cell-derived septoclasts resorb cartilage during developmental ossification and fracture healing
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.