Abstract
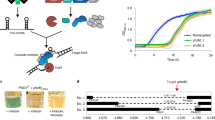
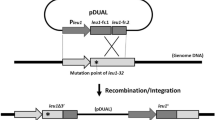
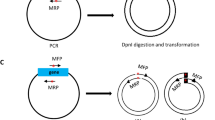
We describe a rapid method for the production of fusion PCR products that can be used, generally without band purification, to transform Aspergillus nidulans. This technique can be used to replace genes; tag genes with fluorescent moeties or epitope tags; or replace endogenous promoters with regulatable promoters, by introducing an appropriate selective cassette (e.g., fluorescent protein + selectable marker). The relevant genomic fragments and cassette are first amplified separately by PCR using primers that produce overlapping ends. A second PCR using 'nested' primers fuses the fragments into a single molecule with all sequences in the desired order. This procedure allows a cassette to be amplified once, frozen and used subsequently in many fusion PCRs. Transformation of nonhomologous recombination deficient (nkuAΔ) strains of A. nidulans with fusion PCR products results in high frequencies of accurate gene targeting. Fusion PCR takes less than 2 d. Protoplast formation and transformation takes less than 1 d.
NOTE:The version of this article initially published online contained the following errors: In the list of authors’ names, Berl Oakley should have been listed as Berl R Oakley. Figure 2: “Fusion PCR product”, “Chromosomal target” and “Recombined chromosome” labels were missing from each of the three panels, and an unnecessary “Upstream sequence” label appeared between panels b and c. Figure 3: “Histone H1” and “3' UTR” labels and corresponding bars were missing from panel a. p. 3113, under REAGENT SETUP: In the description of 1 M Tris–HCl solution, the last sentence should read “Store at room temperature (20 oC–27 oC) or at 4 oC." p. 3114, left column: In the third full paragraph, "tag4" should read "tag 4". p. 3314, Step 2: The last sentence should read: "We normally CsCl purify a large quantity of genomic DNA10 and use it for many experiments, but DNA purified in other ways will also work." The text above the table should read: "PCR reaction mix". The errors have been corrected in all versions of the article.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
08 March 2007
see note in abstract
References
Dunne, P.W. & Oakley, B.R. Mitotic gene conversion, reciprocal recombination and gene replacement at the benA, β-tubulin locus of Aspergillus nidulans. Mol. Gen. Genet. 213, 339–345 (1988).
Toews, M.W. et al. Establishment of mRFP1 as a fluorescent marker in Aspergillus nidulans and construction of expression vectors for high-throughput protein tagging using recombination in vitro (Gateway). Curr. Genet. 45, 383–389 (2004).
Kuwayama, H. et al. PCR-mediated generation of a gene disruption construct without the use of DNA ligase and plasmid vectors. Nucleic Acids Res. 30, E2 (2002).
Yang, L. et al. Rapid production of gene replacement constructs and generation of a green fluorescent protein-tagged centromeric marker in Aspergillus nidulans. Eukaryotic Cell 3, 1359–1362 (2004).
Yu, J.-H. et al. Double-joint PCR: a PCR-based molecular tool for gene manipulations in filamentous fungi. Fungal Genet. Biol. 41, 973–981 (2004).
Nayak, T. et al. A versatile and efficient gene-targeting system for Aspergillus nidulans. Genetics 172, 1557–1566 (2006).
Zarrin, M., Leeder, A.C. & Turner, G. A rapid method for promoter exchange in Aspergillus nidulans using recombinant PCR. Fungal Genet. Biol. 42, 1–8 (2005).
Galagan, J.E. et al. Sequencing of Aspergillus nidulans and comparative analysis with A. fumigatus and A. oryzae. Nature 438, 1105–1115 (2005).
Chaveroche, M.K., Ghigo, J.M. & d'Enfert, C. A rapid method for efficient gene replacement in the filamentous fungus Aspergillus nidulans. Nucleic Acids Res. 28, E97 (2000).
Oakley, B.R. et al. Cloning, mapping and molecular analysis of the pyrG (orotidine-5′-phosphate decarboxylase) gene of Aspergillus nidulans. Gene 61, 385–399 (1987).
Osmani, A.H., Oakley, B.R. & Osmani, S.A. Identification and analysis of essential genes using the heterokaryon rescue technique in Aspergillus nidulans. Nature Protocols (in the press). DOI: 10.1038/nprot.2006.406.
Vishniac, W. & Santer, M. The Thiobacilli. Bacteriol. Rev. 21, 195–213 (1957).
Innis, M.A. & Gelfand, D.H. Optimization of PCRs. In PCR Protocols (eds. Innis, M.A., Gelfand, D.H., Sninsky, J.J. & White, T.J.) 3–12 (Academic Press, San Diego, 1990).
Labib, K., Diffley, J.F. & Kearsey, S.E. G1-phase and B-type cyclins exclude the DNA replication factor Mcm4 from the nucleus. Nat. Cell Biol. 1, 415–422 (1999).
Acknowledgements
This work was supported by grants GM31837 and GM042564 from the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Szewczyk, E., Nayak, T., Oakley, C. et al. Fusion PCR and gene targeting in Aspergillus nidulans. Nat Protoc 1, 3111–3120 (2006). https://doi.org/10.1038/nprot.2006.405
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.405
This article is cited by
-
Streptomyces polyketides mediate bacteria–fungi interactions across soil environments
Nature Microbiology (2023)
-
Species-specific effects of the introduction of Aspergillus nidulans gfdB in osmophilic aspergilli
Applied Microbiology and Biotechnology (2023)
-
Construction of a food-grade gene editing system based on CRISPR-Cas9 and its application in Lactococcus lactis NZ9000
Biotechnology Letters (2023)
-
The C-22 sterol desaturase Erg5 is responsible for ergosterol biosynthesis and conidiation in Aspergillus fumigatus
Journal of Microbiology (2022)
-
Development of versatile and efficient genetic tools for the marine-derived fungus Aspergillus terreus RA2905
Current Genetics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.