Abstract
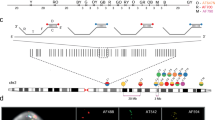
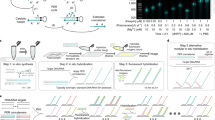
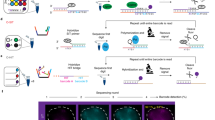
Combined binary ratio labeling (COBRA) is designed to increase the multiplicity of fluorescence in situ hybridization (FISH)—i.e., the number of targets that can be distinguished simultaneously. In principle, chemical (ULS), enzymatic (nick translation or random priming) or PCR-based labeling procedures of probes can be used. The method was originally designed to label chromosome-painting probes, but has also been used for probe sets specific for subtelomeric regions. COBRA imaging requires a digital fluorescence microscope equipped for sequential excitation and recording of color images. Staining of all 24 human chromosomes is accomplished with only four fluorochromes, compared with five for methods based on combinatorial labeling. The COBRA procedure takes ∼6 h laboratory work, 2–3 d incubation and a few hours imaging. The method is routinely applied in research (cultured cells from human or mouse origin) or to support clinical diagnosis, such as postnatal and perinatal genetic testing and in solid tumors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Nederlof, P.M. et al. Multiple fluorescence in situ hybridization. Cytometry 11, 126–131 (1990).
Nederlof, P.M., van der Flier, S., Vrolijk, J., Tanke, H.J. & Raap, A.K. Fluorescence ratio measurements of double-labeled probes for multiple in situ hybridization by digital imaging microscopy. Cytometry 13, 839–845 (1992).
Speicher, M.R., Gwyn, B.S. & Ward, D.C. Karyotyping human chromosomes by combinatorial multi-fluor FISH. Nat. Genet. 12, 368–375 (1996).
Schrock, E. et al. Multicolor spectral karyotyping of human chromosomes. Science 273, 494–497 (1996).
Tanke, H.J. et al. New strategy for multi-colour fluorescence in situ hybridisation: COBRA: COmbined Binary RAtio labelling. Eur. J. Hum. Genet. 7, 2–11 (1999).
Szuhai, K. et al. Simultaneous molecular karyotyping and mapping of viral DNA integration sites by 25-color COBRA–FISH. Genes Chromosomes Cancer 28, 92–97 (2000).
Wiegant, J. et al. Differentially painting human chromosome arms with combined binary ratio-labeling fluorescence in situ hybridization. Genome Res. 10, 861–865 (2000).
Bezrookove, V. et al. Individuals with abnormal phenotype and normal G-banding karyotype: improvement and limitations in the diagnosis by the use of 24-colour FISH. Hum. Genet. 106, 392–398 (2000).
Szuhai, K. et al. Multicolor fluorescence in situ hybridization analysis of a synovial sarcoma of the larynx with a t(X;18)(p11.2;q11.2) and trisomies 2 and 8. Cancer Genet. Cytogenet. 153, 48–52 (2004).
Engels, H. et al. Comprehensive analysis of human subtelomeres with combined binary ratio labelling fluorescence in situ hybridisation. Eur. J. Hum. Genet. 11, 643–651 (2003).
Fodde, R. et al. Mutations in the APC tumour suppressor gene cause chromosomal instability. Nat. Cell Biol. 3, 433–438 (2001).
Hazelbag, H.M., Szuhai, K., Tanke, H.J., Rosenberg, C. & Hogendoorn, P.C. Primary synovial sarcoma of the heart: a cytogenetic and molecular genetic analysis combining RT–PCR and COBRA–FISH of a case with a complex karyotype. Mod. Pathol. 17, 1434–1439 (2004).
Szuhai, K., Ijszenga, M., Tanke, H.J., Rosenberg, C. & Hogendoorn, P.C. Molecular cytogenetic characterization of four previously established and two newly established Ewing sarcoma cell lines. Cancer Genet. Cytogenet. 166, 173–179 (2006).
Telenius, H. et al. Degenerate oligonucleotide-primed PCR: general amplification of target DNA by a single degenerate primer. Genomics 13, 718–725 (1992).
Telenius, H. et al. Cytogenetic analysis by chromosome painting using DOP–PCR amplified flow-sorted chromosomes. Genes Chromosomes Cancer 4, 257–263 (1992).
Fiegler, H. et al. DNA microarrays for comparative genomic hybridization based on DOP–PCR amplification of BAC and PAC clones. Genes Chromosomes Cancer 36, 361–374 (2003).
Rabbitts, P. et al. Chromosome specific paints from a high resolution flow karyotype of the mouse. Nat. Genet. 9, 369–375 (1995).
Wiegant, J. C. et al. ULS: a versatile method of labeling nucleic acids for FISH based on a monofunctional reaction of cisplatin derivatives with guanine moieties. Cytogenet. Cell Genet. 87, 47–52 (1999).
van de Rijke, F.M., Heetebrij, R.J., Talman, E.G., Tanke, H.J. & Raap, A.K. Fluorescence properties, thermal duplex stability, and kinetics of formation of cyanin platinum DNAs. Anal. Biochem. 321, 71–78 (2003).
Wiegant, J. & Raap, A.K. in Current Protocols in Cytometry (eds. Robinson, J.P et al.) 8.3.1–8.3.21 (Wiley & Sons, New York, 1998).
Koopman, L.A. et al. Recurrent integration of human papillomaviruses 16, 45, and 67 near translocation breakpoints in new cervical cancer cell lines. Cancer Res. 59, 5615–5624 (1999).
Brink, A.A. et al. Simultaneous mapping of human papillomavirus integration sites and molecular karyotyping in short-term cultures of cervical carcinomas by using 49-color combined binary ratio labeling fluorescence in situ hybridization. Cancer Genet. Cytogenet. 134, 145–150 (2002).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Szuhai, K., Tanke, H. COBRA: combined binary ratio labeling of nucleic-acid probes for multi-color fluorescence in situ hybridization karyotyping. Nat Protoc 1, 264–275 (2006). https://doi.org/10.1038/nprot.2006.41
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.41
This article is cited by
-
Breast cancer dormancy is associated with a 4NG1 state and not senescence
npj Breast Cancer (2021)
-
Higher cMET dependence of sacral compared to clival chordoma cells: contributing to a better understanding of cMET in chordoma
Scientific Reports (2021)
-
Human melanoma brain metastases cell line MUG-Mel1, isolated clones and their detailed characterization
Scientific Reports (2019)
-
In trans paired nicking triggers seamless genome editing without double-stranded DNA cutting
Nature Communications (2017)
-
p120-catenin prevents multinucleation through control of MKLP1-dependent RhoA activity during cytokinesis
Nature Communications (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.