Abstract
Previous observations of the anatomical distribution and pharmacological profile of the dopamine D3 receptor (DRD3) have indicated its potential role in antipsychotic drug action. Risperidone, an effective first-line atypical antipsychotic agent, exhibits a relatively high affinity for this receptor. Recent studies have reported an association of the Ser9Gly polymorphism in the DRD3 gene with therapeutic response to risperidone, but the results were inconsistent. We therefore postulated that the Ser9Gly polymorphism might be in linkage disequilibrium with an undetected variant that exerts a direct influence on risperidone efficacy. The present study genotyped eight single nucleotide polymorphisms (SNPs) distributed throughout the DRD3 gene and examined five of these for association with treatment outcome, following an 8-week period of risperidone monotherapy in 130 schizophrenic patients from mainland China. Clinical symptoms were assessed before and after the treatment period, using the Brief Psychiatry Rating Scale (BPRS). The confounding effects of non-genetic factors were estimated and the baseline symptom score was included as a covariate for adjustment. Neither was any association observed between the five polymorphisms and improvement in total BPRS scores nor was any combined effect of these variants detected in the haplotype analysis. The current results indicate that genetic variations within the DRD3 gene may not contribute significantly to interindividual differences in the therapeutic efficacy of risperidone.
Similar content being viewed by others
INTRODUCTION
In the last three decades, the dopamine hypothesis of schizophrenia has been the dominant force in driving the development of antipsychotic drugs. Given the tight correlation between the clinical potency and the D2-binding affinity of the conventional neuroleptics (Creese et al, 1976; Seeman et al, 1976), the dopamine D2 receptor is assumed to serve as a principal target for antipsychotic action. In vivo imaging studies have further demonstrated the importance of dopamine D2 receptor occupancy in predicting antipsychotic response and side effects (Remington and Kapur, 1999). The advent of novel antipsychotics and the identification of additional dopamine receptor subtypes, however, have allowed for new insights into the wider diversity of pharmacological actions of antipsychotic drugs.
The dopamine D3 receptor (DRD3) was cloned and characterized as a D2-like receptor in 1990 (Sokoloff et al, 1990). Further observations on the preferential expression of this receptor in limbic and basal ganglia regions associated with cognitive, emotional, and motor functions (Murray et al, 1994; Suzuki et al, 1998), as well as other neurochemical and anatomical investigations (Schmauss et al, 1993; Gurevich et al, 1997), implicated a role for the D3 receptor in the etiology of schizophrenia. It is postulated that the D3 receptor plays an important role in the regulation of neurotransmission by inhibiting spontaneous secretion of the neurotransmitter (Kuzhikandathil and Oxford, 1999, 2000). Pharmacological studies have shown that many antipsychotic drugs also exhibit a high affinity for the D3 receptor (Schwartz et al, 2000); thus, raising the question of the role this protein may play as an important target for antipsychotic drugs.
Compared with conventional neuroleptics, atypical antipsychotics are characterized by a low rate of extrapyramidal side effects (EPS) and a broader therapeutic profile, including efficacy in treating both positive and negative symptoms of schizophrenia. Risperidone, a benzisoxazole derivative with atypical antipsychotic activity, is widely used as a first-line drug for schizophrenia treatment. Higher antagonistic activity of risperidone at 5-HT2A receptors than at D2 receptors is assumed to well contribute to its atypical property but may not be sufficient by itself (Meltzer, 1999; Kapur and Remington, 2001). Competition experiments vs [125I]iodosulpride showed that risperidone also displayed a high affinity for D3 receptors (Ki=5.4 nM) (Sokoloff et al, 1992). Recent studies have implicated a beneficial role for D3 receptor antagonism in ameliorating negative symptoms of schizophrenia (Joyce and Millan, 2005). For example, administration of selective D3 receptor antagonists enhances social interaction in isolation-reared rodents (Reavill et al, 2000; Brocco et al, 2004). Despite the potential role of D3 receptor antagonism in the treatment of schizophrenia, several studies have showed that D3 agonists rather than D3 antagonists could produce antipsychotic effects (Witkin et al, 1998; Fink-Jensen, 2000). Consequently, the contribution of D3 receptor antagonism to the antipsychotic activity of risperidone remains to be examined.
Among the single nucleotide polymorphisms (SNPs) identified in the DRD3 gene, the Ser9Gly polymorphism has been the most extensively investigated. This polymorphic site in the first exon of the DRD3 gene results in a serine to glycine substitution in the N-terminal extracellular domain of the receptor protein (Lannfelt et al, 1992). A series of reports including a recent meta-analysis have suggested an association in favor of the Gly-9 vs the Ser-9 variant with an increased risk of developing antipsychotic-induced tardive dyskinesia (Steen et al, 1997; Lerer et al, 2002; Bakker et al, 2006). Prior studies on the association between medication response and the Ser9Gly polymorphism were primarily a by-product of the research on the role of this polymorphism in schizophrenia. Two studies investigating a monotherapeutic strategy detected an excess of Gly-9 allele in responders to clozapine treatment (Shaikh et al, 1996; Scharfetter et al, 1999). However, a third report failed to replicate these results (Malhotra et al, 1998).
Recently, several studies have suggested an association between the D3 receptor Ser9Gly polymorphism and antipsychotic response to risperidone, but the results were inconsistent (Szekeres et al, 2004; Lane et al, 2005; Reynolds et al, 2005). To further evaluate the importance of the DRD3 in the therapeutic efficacy of risperidone, we performed an association study on polymorphisms from different regions of the DRD3 gene, in a relatively large sample of Chinese schizophrenic patients following risperidone treatment.
MATERIALS AND METHODS
Subjects
One hundred and forty-five Chinese Han schizophrenia patients participated in the study on admission, all of who were recruited from the Shanghai Mental Health Center. Diagnosis was confirmed by two independent psychiatrists conducting of clinical interviews based on the criteria of DSM-IV. Any subject suffering from a severe neurological or physical disease was excluded from the study. All patients were receiving atypical antipsychotic drug treatment for the first time. The study was conducted in compliance with all principles of the Helsinki Accord. A standard informed consent in the protocol, which was reviewed and approved by the Shanghai Ethical Committee of Human Genetic Resources, was obtained from the participants after the procedure had been fully explained.
Clinical Protocols
Before treatment with risperidone, all patients were subjected to at least a 4-week washout period. Risperidone was given at an initial dose of 2 mg/day, and dosage was gradually increased to 4 mg/day within the first week. The drug dose was maintained until day 14, and then adjusted according to individual tolerance. Medication compliance was closely monitored and confirmed by nursing staff. During the study period, no other drugs were given except flunitrazepam for acute insomnia and biperiden for moderate EPS. The Brief Psychiatric Rating Scale (BPRS; Overall and Gorham, 1962) was used to assess clinical status at the beginning and after 8 weeks of treatment. Medication efficacy was evaluated in terms of a percent change in BPRS total scores. Clinical response was defined as an improvement from baseline of 40% or more in BPRS total score.
Genotyping
High-molecular-weight genomic DNA was prepared from venous blood using standard phenol chloroform extraction. Six SNPs including rs324028, rs717668, rs324026, rs6280, rs3773678, rs2134655 were selected from dbSNP (http://www.ncbi.nlm.nih.gov), spanning around 36.4 kb in the DRD3 gene. SNP rs324028, SNP rs717668 in the 5′ noncoding region and SNP rs324026 in the 5′ promoter region of the DRD3 gene are located 3741, 1074, and 203 bp upstream from the start codon, respectively. SNP rs6280 is in the coding sequence of the DRD3 gene, resulting in an amino acid substitution of serine for glycine at position 9 (Ser9Gly). SNP rs3773678 and SNP rs2134655 are located in the introns. Other polymorphisms (Ala38Thr, −7685G>C) in the DRD3 gene (Ishiguro et al, 2000; Staddon et al, 2005) were excluded from further tests for association with drug response because they were present at very low frequency in our sample.
All SNPs were genotyped by direct DNA sequencing, using a BigDye Terminator Cycle Sequencing Kit and an ABI PRISM 3100 DNA sequencer (PE Applied Biosystems, Perkin-Elmer). PCR amplification was performed on a Gene Amp PCR system 9700 (PE Applied Biosystems, Foster City, CA) in a volume of 10 μl containing 10 ng genomic DNA, 1 U Taq polymerase, 0.5 μM of each primer, 10 × PCR buffer, 1.5 mM MgCl2, and 0.2 mM of each dNTP. PCR conditions consisted of an initial denaturation at 95°C for 3 min, 35 cycles of 30 s at 94°C, 30 s at annealing temperature, and 1 min at 72°C, followed by a terminal extension at 72°C for 7 min. Primer sequences and annealing temperatures are listed in Table 1. All genotypes were called blind to the clinical outcome of antipsychotic treatment.
Statistical Analysis
Deviations from Hardy–Weinberg equilibrium for each polymorphism were assessed using exact tests based on a Markov chain algorithm using GENEPOP 3.4 (Rousset and Raymond, 1995). Linkage disequilibrium (LD) of each pair of the six polymorphisms was measured by estimating standardized D′ values on software 2LD (Zapata et al, 2001).
SPSS for Windows (version 10.0) was used for further statistical analyses. A comparison of the mean values of demographic and clinical variables between genotypic subgroups was made using ANOVA followed by Tukey's HSD test. These variables were also examined for gender differences using the Student's t-test or the nonparametric Mann–Whitney U-test. To determine the appropriate method, each variable was checked for normal distribution using the Kolmogorov–Smirnov test and for homogeneity of variances using the Levene's test. The influences of potential confounding factors (age, age of onset, gender, drug concentrations, and baseline BPRS scores) on symptom improvement were assessed in a stepwise regression procedure. Variables significantly correlated with the percentage change in total BPRS scores were selected as covariates for the control of confounding effects. In the analysis of genetic effects on therapeutic efficacy, the general linear model was used with or without adjustment for covariates. The factor-by-covariate interaction was tested to check the homogeneity of within-group regression slopes. Pairwise comparisons between genotypic subgroups were carried out using UNIANOVA. To confirm the results, allele and genotype frequencies of each polymorphism were compared between responder and non-responder groups using the online software SHEsis (http://202.120.7.14/analysis/myAnalysis.php). This software was also used to estimate the significance of differences in haplotype distributions between the two groups. Power analysis of our sample was performed using the G*Power program (Erdfelder et al, 1996). All tests were two-tailed and statistical significance was assumed at p<0.05.
RESULTS
Clinical Profiles
One hundred and thirty patients were available for assessment at the end of the treatment. Of them, 67 had no prior antipsychotic exposure and the remainder had prior exposure to conventional neuroleptics. Fifteen patients dropped out of the study owing to concurrent somatic illness (n=3), poor response (n=6), or noncompliance (n=6). Clinical profiles of the patients eligible for analysis are shown in Table 2. The patients were divided into different groups on the basis of gender. There were no significant differences between gender groups in age, age of onset, duration of illness, steady-state plasma concentrations of risperidone and 9-hydroxyrisperidone, or in total BPRS scores at baseline and at the end of risperidone monotherapy. This indicates that our sample data were not significantly biased with respect to gender for these clinical parameters. In respect of treatment efficacy, the patients were classified as treatment responders (n=72) and non-responders (n=58), based on the percentage improvement in total BPRS scores before and after the 8-week medication. No significant differences in demographic characteristics or plasma levels of risperidone and 9-hydroxyrisperidone were observed between the two groups (p>0.05) (data not shown).
Effects of Individual Polymorphism on Clinical Improvement
All polymorphisms were successfully genotyped in the 130 schizophrenic patients. Genotype frequencies of these polymorphisms showed no significant deviations from Hardy–Weinberg equilibrium (data not shown). Two SNPs (rs324028 and rs324026) were found to be in complete LD (D′=1). This indicated that only one of these two SNPs could provide useful genetic information. Consequently, SNP rs324028 but not rs324026 was included in our association study. The genotypic distributions of five SNPs in the sample are shown in Table 3. There were no significant differences in the clinical parameters including age, age of onset, duration of illness, or drug concentrations between genotypes of analyzed polymorphisms (p>0.05).
The mean BPRS total score was reduced from 42.4±12.4 to 24.0±6.6 during the treatment period, with an improvement of 40.6% in general symptoms. The values of percentage improvement in total BPRS scores were normally distributed (p=0.53); therefore, the data were suitable for multiple regression analysis. The confounding effects of non-genetic factors on symptom improvement were then assessed by a stepwise regression analysis. No significant correlations were found between independent variables, such as age, age of onset, gender, or drug concentrations and the improvement of total BPRS scores, although the severity of illness (ie baseline BPRS scores) exerted a significant influence (p<0.001) on treatment efficacy (data not shown). Consequently, baseline BPRS score was identified as a relevant variable for treatment outcome and was included as a covariate in the analysis of association between DRD3 genotype and symptom improvement.
The mean values of the covariate were not significantly different between genotypic subgroups of each polymorphism (one-way ANOVA p>0.05). Table 3 illustrates the effects of DRD3 polymorphisms on the improvement in total BPRS scores, before and after adjusting for the covariate effects. No significant associations were found between the genotypes of the five polymorphisms and general symptom improvement. Pairwise comparisons between genotypic subgroups also failed to detect any significant differences.
Given the possible influence of the Ser9Gly mutation on the binding capacity of the D3 receptor, we compared patients with the Gly9Gly genotype with those with the Ser9Ser or Ser9Gly genotype. The Gly9Gly genotype had no significant influence on clinical improvement (p=0.418). Moreover, there were no significant differences in allele or genotype distribution in any of the five polymorphisms between responder and non-responder groups.
LD between SNPs and Haplotype Analysis
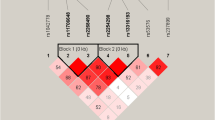
Pairwise LD between the five markers was estimated (expressed in term of D′). As shown in Table 4, strong LD was observed among the three SNPs near the 5′ end of the DRD3 gene. The SNP rs3773678 located in the downstream intron was in much weaker LD with these markers (D′≈0.20), but this was not the case for another intron, SNP rs2134655.
Four SNPs (rs324028, rs717668, rs6280, rs2134655) with significant LD (D′>0.75) were included in the haplotype analysis. Out of 16 possible combinations of alleles, only five haplotypes with a probability >3% represented the majority of the haplotype diversity (96%). The frequencies of these common haplotypes were compared between responder and non-responder groups. As shown in Table 5, there were no significant differences in haplotype distributions between the two groups (global χ2=2.603, p=0.626).
DISCUSSION
The DRD3 has been suggested as a potential drug target for antipsychotic drugs (Schwartz et al, 2000). Recent studies have focused on the Ser9Gly polymorphism of the DRD3 gene, but the results were inconclusive. Although it has been suggested that the Ser9Gly variant could induce a shift in intracellular signal transduction pathways, the molecular mechanisms and the clinical relevance of the switch remain unclear (Hellstrand et al, 2004). There might be other functional variants within the DRD3 gene that affect transcriptional regulation, mRNA stability or splicing efficiency. In our study, eight polymorphisms spanning the DRD3 gene were genotyped and five were investigated for their possible association with risperidone response in 130 unrelated schizophrenic patients from mainland China. However, no significant correlation was found between any variations and general symptom improvement, nor was a trend toward an excess of Ser9 variant observed in patients with better response. Furthermore, we did not detect significant discrepancy of allele, genotype or haplotype frequencies between responder and non-responder groups.
Our results indicate that the effect of variations in the DRD3 gene on therapeutic efficacy of risperidone might be weak or absent. However, this does not rule out a potential role for the D3 receptor in the action of risperidone. Pharmacological studies have suggested that D3 receptor antagonism may contribute to the efficacy of atypical antipsychotics in relieving negative symptoms of schizophrenia (Joyce and Millan, 2005). The overall pharmacological effects of risperidone, however, may be mediated by multireceptor antagonism. In addition to its preferentially high affinity for 5-HT2A and D2 receptors, risperidone also displays relatively high affinity for histamine H1 and alpha-adrenergic receptors (Leysen et al, 1994; Megens et al, 1994). Genetic variations of these receptors may produce confounding effects on assessing the predictive role of DRD3 polymorphisms for risperidone response.
The current study attempted to identify genetic contributors to the clinical variability in risperidone response. However, true factors underlying interindividual variation in medication response may also include demographic, clinical, and environmental variables, which probably confound most pharmacogenetic association studies. The design of this study has some advantages in minimizing the impacts of potential confounding factors. First, patients involved in the current study were receiving atypical antipsychotics for the first time. Prior drug treatment may produce confounding effects on clinical response to subsequent antipsychotic medications. It has been found that long-term effects of atypical antipsychotics on neurotransmitter receptors are somewhat different from those of conventional neuroleptics (White and Wang, 1983; Damask et al, 1996; Tarazi et al, 2002). In particular, long-term treatment with atypical antipsychotics but not with conventional neuroleptics significantly alters the abundance of 5-HT receptor subtypes in forebrain regions (Tarazi et al, 2002). Accordingly, the nature of our sample may reduce to some degree the effects of medical history on risperidone response. Second, the effects of other potential confounding factors (such as age, age of onset, gender, plasma drug concentrations, initial severity of illness) were estimated by regression analysis. We found that initial symptom severity of patients was closely related with their treatment outcome and this variable was controlled as a covariate for the primary variable of interest. Third, our patients were scheduled to receive a target dose of 4 mg/day of risperidone rather than a standard dose of 6 mg/day. A low risk of EPS was observed among these subjects, as was suggested by several other studies (Nyberg et al, 1999; Williams, 2001). Consequently, we were able to maintain a uniform dosage schedule for most patients. This may be useful for reducing individual differences with regard to drug concentration and the possible effect of this variable on medication response.
Power analysis revealed that the statistical power of our sample to detect a significant association (p<0.05) was about 90% in allele or genotype comparisons and over 80% for haplotype analysis when a medium effect size (w=0.3) was presumed. This indicates that the sample size in our study was sufficient to achieve a considerably low risk of a type II error.
In summary, we conducted a detailed association study of the DRD3 gene with risperidone response, but did not detect any significant effect of DRD3 polymorphisms. Our results indicate that variants in either coding or regulatory regions of the gene are not likely to greatly affect the therapeutic efficacy of risperidone for the mainland Chinese population. The assumed predictive role of markers within the DRD3 gene for risperidone treatment outcome should be further evaluated. It may be helpful to investigate a set of markers from different genes that might exert a combined or synergistic effect on drug action, as risperidone has also been found to interact with dopamine D2, D4, serotonin 5-HT2A, histamine H1, adrenergic α1, and α2 receptors. In addition, appropriate control of confounding factors would be useful for further pharmacogenetic studies to focus on the effect of genetic variation on antipsychotic response.
References
Bakker PR, van Harten PN, van Os J (2006). Antipsychotic-induced tardive dyskinesia and the Ser9Gly polymorphism in the DRD3 gene: a meta analysis. Schizophr Res 83: 185–192.
Brocco MC, Gobert A, Dekeyne A, Lavielle G, Millan MJ (2004). S33138, a novel and preferential antagonist at dopamine D3 receptors: functional profile in rodent models of the control of negative-cognitive symptoms of schizophrenia. Int J Neuropsychopharm 7: S265.
Creese I, Burt DR, Snyder SH (1976). Dopamine receptor binding predicts clinical and pharmacological potencies of antischizophrenic drugs. Science 192: 481–483.
Damask SP, Bovenkerk KA, de la Pena G, Hoversten KM, Peters DB, Valentine AM et al (1996). Differential effects of clozapine and haloperidol on dopamine receptor mRNA expression in rat striatum and cortex. Brain Res Mol Brain Res 41: 241–249.
Erdfelder E, Faul F, Buchner A (1996). G*Power: a general power analysis program. Behav Res Methods Instrum Comput 28: 1–11.
Fink-Jensen A (2000). Novel pharmacological approaches to the treatment of schizophrenia. Dan Med Bull 47: 151–167.
Gurevich EV, Bordelon Y, Shapiro RM, Arnold SE, Gur RE, Joyce JN (1997). Mesolimbic dopamine D3 receptors and use of antipsychotics in patients with schizophrenia. A postmortem study. Arch Gen Psychiatry 54: 225–232.
Hellstrand M, Danielsen EA, Steen VM, Ekman A, Eriksson E, Nilsson CL (2004). The ser9gly SNP in the dopamine D3 receptor causes a shift from cAMP related to PGE2 related signal transduction mechanisms in transfected CHO cells. J Med Genet 41: 867–871.
Ishiguro H, Okuyama Y, Toru M, Arinami T (2000). Mutation and association analysis of the 5′ region of the dopamine D3 receptor gene in schizophrenia patients: identification of the Ala38Thr polymorphism and suggested association between DRD3 haplotypes and schizophrenia. Mol Psychiatry 5: 433–438.
Joyce JN, Millan MJ (2005). Dopamine D3 receptor antagonists as therapeutic agents. Drug Discov Today 10: 917–925.
Kapur S, Remington G (2001). Dopamine D2 receptors and their role in atypical antipsychotic action: still necessary and may even be sufficient. Biol Psychiatry 50: 873–883.
Kuzhikandathil EV, Oxford GS (1999). Activation of human D3 dopamine receptor inhibits P/Q-type calcium channels and secretory activity in AtT-20 cells. J Neurosci 19: 1698–1707.
Kuzhikandathil EV, Oxford GS (2000). Dominant-negative mutants identify a role for GIRK channels in D3 dopamine receptor-mediated regulation of spontaneous secretory activity. J Gen Physiol 115: 697–706.
Lane HY, Hsu SK, Liu YC, Chang YC, Huang CH, Chang WH (2005). Dopamine D3 receptor Ser9Gly polymorphism and risperidone response. J Clin Psychopharmacol 25: 6–11.
Lannfelt L, Sokoloff P, Martres MP, Pilon C, Giros B, Jonsson E et al (1992). Amino acid substitution in the dopamine D3 receptor as a useful polymorphism for investigating psychiatric disorders. Psychiatr Genet 2: 249–256.
Lerer B, Segman RH, Fangerau H, Daly AK, Basile VS, Cavallaro R et al (2002). Pharmacogenetics of tardive dyskinesia: combined analysis of 780 patients supports association with dopamine D3 receptor gene Ser9Gly polymorphism. Neuropsychopharmacology 27: 105–119.
Leysen JE, Janssen PM, Megens AA, Schotte A (1994). Risperidone: a novel antipsychotic with balanced serotonin-dopamine antagonism, receptor occupancy profile, and pharmacologic activity. J Clin Psychiatry 55: 5–12.
Malhotra AK, Goldman D, Buchanan RW, Rooney W, Clifton A, Kosmidis MH et al (1998). The dopamine D3 receptor (DRD3) Ser9Gly polymorphism and schizophrenia: a haplotype relative risk study and association with clozapine response. Mol Psychiatry 3: 72–75.
Megens AA, Awouters FH, Schotte A, Meert TF, Dugovic C, Niemegeers CJ et al (1994). Survey on the pharmacodynamics of the new antipsychotic risperidone. Psychopharmacology (Berlin) 114: 9–23.
Meltzer HY (1999). The role of serotonin in antipsychotic drug action. Neuropsychopharmacology 21: 106S–115S.
Murray AM, Ryoo HL, Gurevich E, Joyce JN (1994). Localization of dopamine D3 receptors to mesolimbic and D2 receptors to mesostriatal regions of human forebrain. Proc Natl Acad Sci USA 91: 11271–11275.
Nyberg S, Eriksson B, Oxenstierna G, Halldin C, Farde L (1999). Suggested minimal effective dose of risperidone based on PET-measured D2 and 5-HT2A receptor occupancy in schizophrenic patients. Am J Psychiatry 156: 869–875.
Overall JE, Gorham DR (1962). The Brief Psychiatric Rating Scale. Psychol Rep 10: 799–812.
Reavill C, Taylor SG, Wood MD, Ashmeade T, Austin NE, Avenell KY et al (2000). Pharmacological actions of a novel, high-affinity, and selective human dopamine D3 receptor antagonist, SB-277011-A. J Pharmacol Exp Ther 294: 1154–1165.
Remington G, Kapur S (1999). D2 and 5-HT2 receptor effects of antipsychotics: bridging basic and clinical findings using PET. J Clin Psychiatry 60 (Suppl 10): 15–19.
Reynolds GP, Yao Z, Zhang X, Sun J, Zhang Z (2005). Pharmacogenetics of treatment in first-episode schizophrenia: D3 and 5-HT2C receptor polymorphisms separately associate with positive and negative symptom response. Eur Neuropsychopharmacol 15: 143–151.
Rousset F, Raymond M (1995). Testing heterozygote excess and deficiency. Genetics 140: 1413–1419.
Scharfetter J, Chaudhry HR, Hornik K, Fuchs K, Sieghart W, Kasper S et al (1999). Dopamine D3 receptor gene polymorphism and response to clozapine in schizophrenic Pakastani patients. Eur Neuropsychopharmacol 10: 17–20.
Schmauss C, Haroutunian V, Davis KL, Davidson M (1993). Selective loss of dopamine D3-type receptor mRNA expression in parietal and motor cortices of patients with chronic schizophrenia. Proc Natl Acad Sci USA 90: 8942–8946.
Schwartz JC, Diaz J, Pilon C, Sokoloff P (2000). Possible implications of the dopamine D3 receptor in schizophrenia and in antipsychotic drug actions. Brain Res Brain Res Rev 31: 277–287.
Seeman P, Lee T, Chau-Wong M, Wong K (1976). Antipsychotic drug doses and neuroleptic/dopamine receptors. Nature 261: 717–719.
Shaikh S, Collier DA, Sham PC, Ball D, Aitchison K, Vallada H et al (1996). Allelic association between a Ser-9-Gly polymorphism in the dopamine D3 receptor gene and schizophrenia. Hum Genet 97: 714–719.
Sokoloff P, Andrieux M, Besancon R, Pilon C, Martres MP, Giros B et al (1992). Pharmacology of human dopamine D3 receptor expressed in a mammalian cell line: comparison with D2 receptor. Eur J Pharmacol 225: 331–337.
Sokoloff P, Giros B, Martres MP, Bouthenet ML, Schwartz JC (1990). Molecular cloning and characterization of a novel dopamine receptor (D3) as a target for neuroleptics. Nature 347: 146–151.
Staddon S, Arranz MJ, Mancama D, Perez-Nievas F, Arrizabalaga I, Anney R et al (2005). Association between dopamine D3 receptor gene polymorphisms and schizophrenia in an isolate population. Schizophr Res 73: 49–54.
Steen VM, Lovlie R, MacEwan T, McCreadie RG (1997). Dopamine D3-receptor gene variant and susceptibility to tardive dyskinesia in schizophrenic patients. Mol Psychiatry 2: 139–145.
Suzuki M, Hurd YL, Sokoloff P, Schwartz JC, Sedvall G (1998). D3 dopamine receptor mRNA is widely expressed in the human brain. Brain Res 779: 58–74.
Szekeres G, Keri S, Juhasz A, Rimanoczy A, Szendi I, Czimmer C et al (2004). Role of dopamine D3 receptor (DRD3) and dopamine transporter (DAT) polymorphism in cognitive dysfunctions and therapeutic response to atypical antipsychotics in patients with schizophrenia. Am J Med Genet B Neuropsychiatr Genet 124: 1–5.
Tarazi FI, Zhang K, Baldessarini RJ (2002). Long-term effects of olanzapine, risperidone, and quetiapine on serotonin 1A, 2A and 2C receptors in rat forebrain regions. Psychopharmacology (Berlin) 161: 263–270.
White FJ, Wang RY (1983). Differential effects of classical and atypical antipsychotic drugs on A9 and A10 dopamine neurons. Science 221: 1054–1057.
Williams R (2001). Optimal dosing with risperidone: updated recommendations. J Clin Psychiatry 62: 282–289.
Witkin J, Gasior M, Acri J, Beekman M, Thurkauf A, Yuan J et al (1998). Atypical antipsychotic-like effects of the dopamine D3 receptor agonist, (+)-PD 128,907. Eur J Pharmacol 347: R1–R3.
Zapata C, Carollo C, Rodriguez S (2001). Sampling variance and distribution of the D′ measure of overall gametic disequilibrium between multiallelic loci. Ann Hum Genet 65: 395–406.
Acknowledgements
We are grateful to all the participants working on the project. This work was supported by the Key Grant Project of the Chinese Ministry of Education (No. 10414), the National 973 Programs of China, the National Natural Science Foundation of China, and the Shanghai Municipal Commission for Science and Technology.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Xuan, J., Zhao, X., He, G. et al. Effects of the Dopamine D3 Receptor (DRD3) Gene Polymorphisms on Risperidone Response: A Pharmacogenetic Study. Neuropsychopharmacol 33, 305–311 (2008). https://doi.org/10.1038/sj.npp.1301418
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.npp.1301418
Keywords
This article is cited by
-
Ethnic Differences in Antipsychotic Response: What Genetic Variation Does and Does Not Tell Us
Current Behavioral Neuroscience Reports (2023)
-
Clinical pharmacogenomics in action: design, assessment and implementation of a novel pharmacogenetic panel supporting drug selection for diseases of the central nervous system (CNS)
Journal of Translational Medicine (2021)
-
Combined study of genetic and epigenetic biomarker risperidone treatment efficacy in Chinese Han schizophrenia patients
Translational Psychiatry (2017)
-
Association studies of genomic variants with treatment response to risperidone, clozapine, quetiapine and chlorpromazine in the Chinese Han population
The Pharmacogenomics Journal (2016)
-
Association of common genetic variants with risperidone adverse events in a Spanish schizophrenic population
The Pharmacogenomics Journal (2013)