Abstract
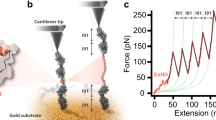
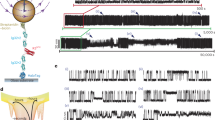
The conformational energy landscape of a protein out of equilibrium is poorly understood. We use single-molecule force-clamp spectroscopy to measure the kinetics of unfolding of the protein ubiquitin under a constant force. We discover a surprisingly broad distribution of unfolding rates that follows a power law with no characteristic mean. The structural fluctuations that give rise to this distribution reveal the architecture of the protein’s energy landscape. Following models of glassy dynamics, this complex kinetics implies large fluctuations in the energies of the folded protein, characterized by an exponential distribution with a width of 5–10kBT. Our results predict the existence of a ‘glass transition’ force below which the folded conformations interconvert between local minima on multiple timescales. These techniques offer a new tool to further test statistical energy landscape theories experimentally.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ansari, A. et al. Protein states and proteinquakes. Proc. Natl Acad. Sci. USA 82, 5000–5004 (1985).
Frauenfelder, H., Sligar, S. G. & Wolynes, P. The energy landscapes and motions of proteins. Science 254, 1598–1603 (1991).
Karplus, M., Gao, Y. Q., Ma, J., van der Vaart, A. & Yang, W. Protein structural transitions and their functional role. Phil. Trans. A 363, 331–356 (2005).
Bryngelson, J. D., Onuchic, J. N., Socci, N. D. & Wolynes, P. G. Funnels, pathways, and the energy landscape of protein folding: a synthesis. Proteins 21, 167195 (1995).
Dill, K. A. et al. Principles of protein folding — a perspective from simple exact models. Protein Sci. 4, 561–602 (1995).
Lazaridis, T. & Karplus, M. ‘New view’ of protein folding reconciled with the old through multiple unfolding simulations. Science 278, 1928–1931 (1997).
Hyeon, C. & Thirumalai, D. Can energy landscape roughness of proteins and RNA be measured by using mechanical unfolding experiments? Proc. Natl Acad. Sci. USA 100, 10249–10253 (2003).
Kneller, G. R. Quasielastic neutron scattering and relaxation processes in proteins: analytical and simulation-based models. Phys. Chem. Chem. Phys. 7, 2641–2655 (2005).
Lindorff-Larsen, K., Best, R. B., DePristo, M. A., Dobson, C. M. & Vendruscolo, M. Simultaneous determination of protein structure and dynamics. Nature 433, 128–132 (2005).
Xue, Q. & Yeung, E. S. Differences in the chemical reactivity of individual molecules of an enzyme. Nature 373, 681–683 (1995).
van Oijen, A. M. et al. Single-molecule kinetics of lambda exonuclease reveal base dependence and dynamic disorder. Science 301, 1235–1238 (2003).
Itoh, K. & Sasai, M. Dynamical transition and proteinquake in photoactive yellow protein. Proc. Natl Acad. Sci. USA 101, 14736–14741 (2004).
Yang, H. et al. Protein conformational dynamics probed by single-molecule electron transfer. Science 302, 262–266 (2003).
Min, W., Luo, G., Cherayil, B. J., Kou, S. C. & Xie, X. S. Observation of a power-law memory kernel for fluctuations within a single protein molecule. Phys. Rev. Lett. 94, 198302 (2005).
Fersht, A. R. Structure and Mechanism in Protein Science (Freeman, New York, 1992).
Carrion-Vazquez, M. et al. Protein nanomechanics — as studied by single-molecule force spectroscopy AFM. Biophys. J. 75, 662–671 (1998).
Li, H. B., Carrion-Vazquez, M., Oberhauser, A. F., Marszalek, P. E. & Fernandez, J. M. Point mutations alter the mechanical stability of immunoglobulin modules. Nature Struct. Biol. 7, 1117–1120 (2000).
Schlierf, M., Li, H. & Fernandez, J. M. The unfolding kinetics of ubiquitin captured with single-molecule force-clamp techniques. Proc. Natl Acad. Sci. USA 101, 7299–7304 (2004).
Ross, S. Stochastic Processes (Wiley, New York, 1996).
Johnston, D. & Wu, S. M. Foundations of Cellular Neurophysiology (MIT Press, Cambridge, Massachusetts, 1995).
Colquhoun, D., Hatton, C. J. & Hawkes, A. G. The quality of maximum likelihood estimates of ion channel rate constants. J. Physiol. 547, 699–728 (2003).
Chekmarev, S. F., Krivov, S. V. & Karplus, M. Folding time distributions as an approach to protein folding kinetics. J. Phys. Chem. B 109, 5312–5330 (2005).
Lee, C., Stell, G. & Wang, J. First passage time distribution and non-markovian diffusion dynamics of protein folding. J. Chem. Phys. 118, 959–968 (2003).
Angell, C. A. Perspective on the glass-transition. J. Phys. Chem. Solids 49, 863–871 (1988).
Lubchenko, V. & Wolynes, P. G. Theory of aging in structural glasses. J. Chem. Phys. 121, 2852–2865 (2004).
Marques, M. I. & Stanley, H. E. Relationship between fragility, diffusive directions and energy barriers in a supercooled liquid. Physica A 345, 395–403 (2005).
Derrida, B. Random-energy model — limit of a family of disordered models. Proc. Natl Acad. Sci. USA 45, 79–82 (1980).
Pande, V., Grosberg, A. & Tanaka, T. Statistical mechanics of simple models of protein folding and design. Biophys. J. 73, 3192–3210 (1997).
Bercu, V., Martinelli, M., Massa, C., Pardi, L. & Leporini, D. A study of the deep structure of the energy landscape of glassy polystyrene: the exponential distribution of the energy barriers revealed by high-field electron spin resonance spectroscopy. Biophys. J. 16, L479–L488 (2004).
Monthus, C. & Bouchaud, J. Models of traps and glass phenomenology. J. Phys. A 29, 3847–3869 (1996).
Lee, A. L. & Wand, A. J. Microscopic origins of entropy, heat capacity and the glass transition in proteins. Nature 411, 501–504 (2004).
Nevo, R., Brumfeld, V., Kapon, R., Hinterdorfer, P. & Reich, Z. Direct measurement of protein energy landscape roughness. EMBO Rep. 6, 482–486 (2005).
Rao, F. & Caflisch, A. The protein folding network. J. Mol. Biol. 342, 299–306 (2004).
Barabási, A.-L. & Oltvai, Z. N. Network biology: understanding the cell’s functional organization. Nature 5, 101–113 (2004).
Song, C., Havlin, S. & Makse, H. A. Self-similarity of complex networks. Nature 433, 392–395 (2005).
Acknowledgements
We would like to thank H. H. Huang for making the ubiquitin construct, S. Garcia-Manyes for data collection, and H. A. Makse and A. J. Tolley for enlightening discussions. This work has been supported by NIH grant R01 HL66030 to J.M.F.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Brujić, J., Hermans Z., R., Walther, K. et al. Single-molecule force spectroscopy reveals signatures of glassy dynamics in the energy landscape of ubiquitin. Nature Phys 2, 282–286 (2006). https://doi.org/10.1038/nphys269
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nphys269
This article is cited by
-
Direct observation of chaperone-modulated talin mechanics with single-molecule resolution
Communications Biology (2022)
-
Steering chemical reactions with force
Nature Reviews Chemistry (2017)
-
Protein folding trajectories can be described quantitatively by one-dimensional diffusion over measured energy landscapes
Nature Physics (2016)
-
Force dependency of biochemical reactions measured by single-molecule force-clamp spectroscopy
Nature Protocols (2013)
-
Similarities between protein folding and granular jamming
Nature Communications (2012)