Abstract
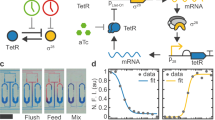
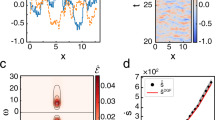
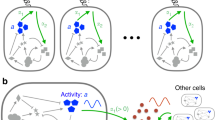
Oscillations within the cell regulate the timing of many important life cycles. However, in this noisy environment, oscillations can be highly inaccurate owing to phase fluctuations. It remains poorly understood how biochemical circuits suppress these phase fluctuations and what is the incurred thermodynamic cost. Here, we study three different types of biochemical oscillation, representing three basic oscillation motifs shared by all known oscillatory systems. In all the systems studied, we find that the phase diffusion constant depends on the free-energy dissipation per period, following the same inverse relation parameterized by system-specific constants. This relationship and its range of validity are shown analytically in a model of noisy oscillation. Microscopically, we find that the oscillation is driven by multiple irreversible cycles that hydrolyse fuel molecules such as ATP; the number of phase coherent periods is proportional to the free energy consumed per period. Experimental evidence in support of this general relationship and testable predictions are also presented.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Eisenberg, E. & Hill, T. L. Muscle contraction and free energy transduction in biological systems. Science 227, 999–1006 (1985).
Hill, T. L. Free Energy Transduction and Biochemical Cycle Kinetics (Academic, 1977).
Qian, H. & Beard, D. A. Chemical Biophysics: Quantitative Analysis of Cellular Systems (Cambridge Univ. Press, 2008).
Jülicher, F., Ajdari, A. & Prost, J. Modeling molecular motors. Rev. Mod. Phys. 69, 1269–1281 (1997).
Nelson, D. L., Lehninger, A. L. & Cox, M. M. Lehninger Principles of Biochemistry (Macmillan, 2008).
Bialek, W. & Setayeshgar, S. Physical limits to biochemical signaling. Proc. Natl Acad. Sci. USA 102, 10040–10045 (2005).
Hu, B., Chen, W., Rappel, W. J. & Levine, H. Physical limits on cellular sensing of spatial gradients. Phys. Rev. Lett. 105, 048104 (2010).
Lan, G., Sartori, P., Neumann, S., Sourjik, V. & Tu, Y. The energy–speed–accuracy tradeoff in sensory adaptation. Nature Phys. 8, 422–428 (2012).
Lan, G. & Tu, Y. The cost of sensitive response and accurate adaptation in networks with an incoherent type-1 feed-forward loop. J. R. Soc. Interface 10, 20130489 (2013).
Skoge, M., Naqvi, S., Meir, Y. & Wingreen, N. S. Chemical sensing by nonequilibrium cooperative receptors. Phys. Rev. Lett. 110, 248102 (2013).
Lang, A. H., Fisher, C. K., Mora, T. & Mehta, P. Thermodynamics of statistical inference by cells. Phys. Rev. Lett. 113, 148103 (2014).
Goldbeter, A. Biochemical Oscillations and Cellular Rhythms: The Molecular Bases of Periodic and Chaotic Behaviour (Cambridge Univ. Press, 1996).
Martiel, J. L. & Goldbeter, A. A model based on receptor desensitization for cyclic amp signaling in dictyostelium cells. Biophys. J. 52, 807–828 (1987).
Pomerening, J. R., Sontag, E. D. & Ferrell, J. E. Building a cell cycle oscillator: Hysteresis and bistability in the activation of cdc2. Nature Cell Biol. 5, 346–351 (2003).
Tsai, T. Y.-C. et al. Robust, tunable biological oscillations from interlinked positive and negative feedback loops. Science 321, 126–129 (2008).
Ferrell, J. J., Tsai, T. Y. & Yang, Q. Modeling the cell cycle: Why do certain circuits oscillate? Cell 144, 874–885 (2011).
Hogenesch, J. B. & Ueda, H. R. Understanding systems-level properties: Timely stories from the study of clocks. Nature Rev. Genet. 12, 407–416 (2011).
Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
Stricker, J. et al. A fast, robust and tunable synthetic gene oscillator. Nature 456, 516–519 (2008).
Novak, B. & Tyson, J. J. Design principles of biochemical oscillators. Nature Rev. Mol. Cell Biol. 9, 981–991 (2008).
Barkai, N. & Leibler, S. Circadian clocks limited by noise. Nature 403, 267–268 (2000).
van Dorp, M., Lannoo, B. & Carlon, E. Generation of oscillating gene regulatory network motifs. Phys. Rev. E 88, 012722 (2013).
Nakajima, M. et al. Reconstitution of circadian oscillation of cyanobacterial KaiC phosphorylation in vitro. Science 308, 414–415 (2005).
Rust, M. J., Markson, J. S., Lane, W. S., Fisher, D. S. & O’Shea, E. K. Ordered phosphorylation governs oscillation of a three-protein circadian clock. Science 318, 809–812 (2007).
Goldbeter, A. A minimal cascade model for the mitotic oscillator involving cyclin and cdc2 kinase. Proc. Natl Acad. Sci. USA 88, 9107–9111 (1991).
Pomerening, J. R., Kim, S. Y. & Ferrell, J. E. Jr. Systems-level dissection of the cell-cycle oscillator: Bypassing positive feedback produces damped oscillations. Cell 122, 565–578 (2005).
Danino, T., Mondragon-Palomino, O., Tsimring, L. & Hasty, J. A synchronized quorum of genetic clocks. Nature 463, 326–330 (2010).
Prindle, A. et al. A sensing array of radically coupled genetic biopixels. Nature 481, 39–44 (2012).
Krishna, S., Jensen, M. H. & Sneppen, K. Minimal model of spiky oscillations in nf-κb signaling. Proc. Natl Acad. Sci. USA 103, 10840–10845 (2006).
Geva-Zatorsky, N. et al. Oscillations and variability in the p53 system. Mol. Syst. Biol. 2, 2006.0033 (2006).
Szallasi, Z., Stelling, J. & Periwal, V. System Modeling in Cell Biology: From Concepts to Nuts and Bolts (MIT Press, 2006).
Goldbeter, A. & Lefever, R. Dissipative structures for an allosteric model. Application to glycolytic oscillations. Biophys. J. 12, 1302–1315 (1972).
Dupont, G., Berridge, M. & Goldbeter, A. Signal-induced Ca2+ oscillations: Properties of a model based on Ca2+-induced Ca2+ release. Cell Calcium 12, 73–85 (1991).
Qian, H. Phosphorylation energy hypothesis: Open chemical systems and their biological functions. Annu. Rev. Phys. Chem. 58, 113–142 (2007).
Rust, M. J., Golden, S. S. & O’Shea, E. K. Light-driven changes in energy metabolism directly entrain the cyanobacterial circadian oscillator. Science 331, 220–223 (2011).
Phong, C., Markson, J. S., Wilhoite, C. M. & Rust, M. J. Robust and tunable circadian rhythms from differentially sensitive catalytic domains. Proc. Natl Acad. Sci. USA 110, 1124–1129 (2013).
Tome, T. & de Oliveira, M. J. Entropy production in irreversible systems described by a Fokker–Planck equation. Phys. Rev. E 82, 021120 (2010).
Seifert, U. Entropy production along a stochastic trajectory and an integral fluctuation theorem. Phys. Rev. Lett. 95, 040602 (2005).
Berg, H. C. & Purcell, E. M. Physics of chemoreception. Biophys. J. 20, 193–219 (1977).
Terauchi, K. et al. Atpase activity of KaiC determines the basic timing for circadian clock of cyanobacteria. Proc. Natl Acad. Sci. USA 104, 16377–16381 (2007).
Barkai, N. & Leibler, S. Robustness in simple biochemical networks. Nature 387, 913–917 (1997).
Ma, W., Trusina, A., El-Samad, H., Lim, W. A. & Tang, C. Defining network topologies that can achieve biochemical adaptation. Cell 138, 760–773 (2009).
Gillespie, D. T. Exact stochastic simulation of coupled chemical reactions. J. Chem. Phys. 81, 2340–2361 (1977).
Author information
Authors and Affiliations
Contributions
Y.T. and Q.O. initiated the work; Y.C., H.W., Q.O. and Y.T. designed the research; Y.C. performed simulations; Y.C. and Y.T. contributed to the analytical results; Y.C., Q.O. and Y.T. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 647 kb)
Rights and permissions
About this article
Cite this article
Cao, Y., Wang, H., Ouyang, Q. et al. The free-energy cost of accurate biochemical oscillations. Nature Phys 11, 772–778 (2015). https://doi.org/10.1038/nphys3412
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nphys3412
This article is cited by
-
OxPhos defects cause hypermetabolism and reduce lifespan in cells and in patients with mitochondrial diseases
Communications Biology (2023)
-
Complex dynamics in a synchronized cell-free genetic clock
Nature Communications (2022)
-
Dynamics and Sensitivity of Signaling Pathways
Current Pathobiology Reports (2022)
-
The energy cost and optimal design for synchronization of coupled molecular oscillators
Nature Physics (2020)
-
Noise control and utility: From regulatory network to spatial patterning
Science China Mathematics (2020)