Abstract
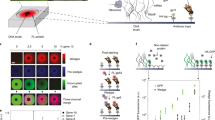
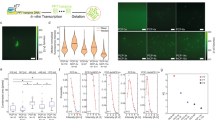
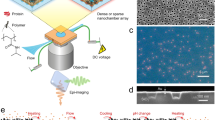
Biologically active complexes such as ribosomes and bacteriophages are formed through the self-assembly of proteins and nucleic acids1,2. Recapitulating these biological self-assembly processes in a cell-free environment offers a way to develop synthetic biodevices3,4,5,6. To visualize and understand the assembly process, a platform is required that enables simultaneous synthesis, assembly and imaging at the nanoscale. Here, we show that a silicon dioxide grid, used to support samples in transmission electron microscopy, can be modified into a biochip to combine in situ protein synthesis, assembly and imaging. Light is used to pattern the biochip surface with genes that encode specific proteins, and antibody traps that bind and assemble the nascent proteins. Using transmission electron microscopy imaging we show that protein nanotubes synthesized on the biochip surface in the presence of antibody traps efficiently assembled on these traps, but pre-assembled nanotubes were not effectively captured. Moreover, synthesis of green fluorescent protein from its immobilized gene generated a gradient of captured proteins decreasing in concentration away from the gene source. This biochip could be used to create spatial patterns of proteins assembled on surfaces.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Mulder, A. M. et al. Visualizing ribosome biogenesis: parallel assembly pathways for the 30S subunit. Science 330, 673–677 (2010).
Leiman, P. G. et al. Morphogenesis of the T4 tail and tail fibers. Virol. J. 7, 355 (2010).
Noireaux, V., Bar-Ziv, R. & Libchaber, A. Principles of cell-free genetic circuit assembly. Proc. Natl Acad. Sci. USA 100, 12672–12677 (2003).
Seeman, N. C. Nanomaterials based on DNA. Annu. Rev. Biochem. 79, 65–87 (2010).
Krishna, Y. & Simmel, F. Nucleic acid based molecular devices. Angew. Chem. 50, 3124–3156 (2011).
Franco, E. et al. Timing molecular motion and production with a synthetic transcriptional clock. Proc. Natl Acad. Sci. USA 108, E784–E793 (2011).
Jewett, M. C., Calhoun, K. A., Voloshin, A., Wuu, J. J. & Swartz, J. R. An integrated cell-free metabolic platform for protein production and synthetic biology. Mol. Sys. Biol. 4, 220 (2008).
Karzbrun, E., Shin, J., Bar-Ziv, R. H. & Noireaux, V. Coarse-grained dynamics of protein synthesis in a cell-free system. Phys. Rev. Lett. 106, 048104 (2011).
Shin, J. & Noireaux, V. An E. coli cell-free expression toolbox: application to synthetic gene circuits and artificial cells. ACS Synth. Biol. 1, 29–41 (2012).
Shimizu, Y. et al. Cell-free translation reconstituted with purified components. Nature Biotechnol. 19, 751–755 (2001).
Daube, S. S., Arad, T. & Bar-Ziv, R. Cell-free co-synthesis of protein nanoassemblies: tubes, rings, and doughnuts. Nano. Lett. 7, 638–641 (2007).
Jensen, G. J. Cryo-EM, Part B: 3-D reconstruction. Preface. Methods Enzymol. 482, xv–xvi (2010).
Frank, J. Single-particle reconstruction of biological macromolecules in electron microscopy—30 years. Q. Rev. Biophys. 42, 139–158 (2009).
Kelly, D. F., Abeyrathne, P. D., Dukovski, D. & Walz, T. The affinity grid: a pre-fabricated EM grid for monolayer purification. J. Mol. Biol. 382, 423–433 (2008).
Buxboim, A. et al. A single-step photolithographic interface for cell-free gene expression and active biochips. Small 3, 500–510 (2007).
Bar, M. & Bar-Ziv, R. H. Spatially resolved DNA brushes on a chip: gene activation by enzymatic cascade. Nano. Lett. 9, 4462–4466 (2009).
Buxboim, A., Daube, S. S. & Bar-Ziv, R. Synthetic gene brushes: a structure–function relationship. Mol. Sys. Biol. 4, 181 (2008).
Kearns, G. J., Foster, E. W. & Hutchison, J. E. Substrates for direct imaging of chemically functionalized SiO2 surfaces by transmission electron microscopy. Anal. Chem. 78, 298–303 (2006).
Aksyuk, A. A. et al. The tail sheath structure of bacteriophage T4: a molecular machine for infecting bacteria. EMBO J. 28, 821–829 (2009).
Cox, M. M. The bacterial RecA protein as a motor protein. Annu. Rev. Microbiol. 57, 551–577 (2003).
Buxboim, A., Daube, S. S. & Bar-Ziv, R. Ultradense synthetic gene brushes on a chip. Nano. Lett. 9, 909–913 (2009).
Hayat, M. A. & Miller, S. E. Negative Staining (McGraw-Hill, 1990).
Acknowledgements
This work was supported by the Israel Science Foundation (Converging Technology programme) and the Minerva Foundation. The electron microscopy studies were conducted at the Irving and Cherna Moskowitz Center for Nano and Bio-Nano Imaging at the Weizmann Institute of Science. The authors thank T. Arad and V. Shinder for TEM assistance.
Author information
Authors and Affiliations
Contributions
Y.H., S.S.D. and R.H.B-Z. conceived and designed the experiments. Y.H., A.B. and S.S.D. performed the experiments. S.G.W. contributed TEM expertise and materials/analysis tools. Y.H., S.S.D. and R.H.B-Z. co-wrote the paper. All authors analysed the data, discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 1059 kb)
Rights and permissions
About this article
Cite this article
Heyman, Y., Buxboim, A., Wolf, S. et al. Cell-free protein synthesis and assembly on a biochip. Nature Nanotech 7, 374–378 (2012). https://doi.org/10.1038/nnano.2012.65
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2012.65
This article is cited by
-
Programming multi-protein assembly by gene-brush patterns and two-dimensional compartment geometry
Nature Nanotechnology (2020)
-
Cell-free gene expression: an expanded repertoire of applications
Nature Reviews Genetics (2020)
-
DNA condensation in one dimension
Nature Nanotechnology (2016)
-
Assemble-on-a-chip
Nature Methods (2012)