Abstract
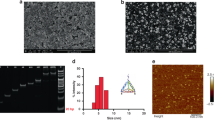
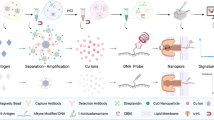
In resource-constrained countries, affordable methodologies for the detection of disease biomarkers at ultralow concentrations can potentially improve the standard of living1,2. However, current strategies for ultrasensitive detection often require sophisticated instruments that may not be available in laboratories with fewer resources3,4,5,6,7,8,9,10,11. Here, we circumvent this problem by introducing a signal generation mechanism for biosensing that enables the detection of a few molecules of analyte with the naked eye. The enzyme label of an enzyme-linked immunosorbent assay (ELISA) controls the growth of gold nanoparticles and generates coloured solutions with distinct tonality when the analyte is present. Prostate specific antigen (PSA) and HIV-1 capsid antigen p24 were detected in whole serum at the ultralow concentration of 1 × 10−18 g ml−1. p24 was also detected with the naked eye in the sera of HIV-infected patients showing viral loads undetectable by a gold standard nucleic acid-based test.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Patton, J. C., Coovadia, A. H., Meyers, T. M. & Sherman, G. G. Evaluation of the ultrasensitive human immunodeficiency virus type (HIV-1) p24 antigen assay performed on dried blood spots for diagnosis of HIV-1 infection in infants. Clin. Vaccine Immunol. 15, 388–391 (2008).
Tang, S. & Hewlett, I. Nanoparticle-based immunoassays for sensitive and early detection of HIV-1 capsid (p24) antigen. J. Infect. Dis. 201, S59–S64 (2010).
Rodriguez-Lorenzo, L., de la Rica, R., Alvarez-Puebla, R., Liz-Marzan, L. M. & Stevens, M. M. Plasmonic nanosensors with inverse sensitivity by means of enzyme-guided crystal growth. Nature Mater. 11, 604–607 (2012).
Nam, J. M., Thaxton, C. S. & Mirkin, C. A. Nanoparticle-based bio-bar codes for the ultrasensitive detection of proteins. Science 301, 1884–1886 (2003).
Rissin, D. M. et al. Single-molecule enzyme-linked immunosorbent assay detects serum proteins at subfemtomolar concentrations. Nature Biotechnol. 28, 595–599 (2010).
Tabakman, M. N. et al. Plasmonic substrates for multiplexed protein microarrays with femtomolar sensitivity and broad dynamic range. Nature Commun. 2, 466 (2011).
Laromaine, A., Koh, L. L., Murugesan, M., Ulijn, R. V. & Stevens, M. M. Protease-triggered dispersion of nanoparticle assemblies. J. Am. Chem. Soc. 129, 4156–4157 (2007).
De la Rica, R., Baldi, A., Fernandez-Sanchez, C. & Matsui, H. Single-cell pathogen detection with a reverse-phase immunoassay on impedimetric transducers. Anal. Chem. 81, 7732–7736 (2009).
De la Rica, R., Fratila, R. M., Szarpak, A., Huskens, J. & Velders, A. H. Multivalent nanoparticle networks as ultrasensitive enzyme sensors. Angew. Chem. Int. Ed. 50, 5703–5706 (2011).
Rodriguez-Lorenzo, L. et al. Zeptomol detection through controlled ultrasensitive surface-enhanced Raman scattering. J. Am. Chem. Soc. 131, 4616–4618 (2009).
Chen, S., Svedendahl, M., van Duyne, R. P. & Käll, M. Plasmon-enhanced colorimetric ELISA with single molecule sensitivity. Nano Lett. 11, 1826–1830 (2011).
Kowalczyk, B., Walker, D. A., Soh, S. & Grzybowski, B. A. Nanoparticle supracrystals and layered supracrystals as chemical amplifiers. Angew. Chem. Int. Ed. 49, 5737–5741 (2009).
Aili, D., Selegard, R., Baltzer, L., Enander, K. & Liedberg, B. Colorimetric protein sensing by controlled assembly of gold nanoparticles functionalized with synthetic receptors. Small 5, 2445–2452 (2009).
Qu, W., Liu, Y., Liu, D., Wang, Z. & Jiang, X. Copper-mediated amplification allows readout of immunoassays by the naked eye. Angew. Chem. Int. Ed. 50, 3442–3445 (2011).
Engelbrekt, C. et al. Green synthesis of gold nanoparticles with starch–glucose and application in bioelectrochemistry. J. Mater. Chem. 19, 7839–7847 (2009).
Thaxton, C. S. et al. Nanoparticle-based bio-barcode assay redefines ‘undetectable’ PSA and biochemical recurrence after radical prostatectomy. Proc. Natl Acad. Sci. USA 106, 18437–18442 (2009).
De la Rica, R. & Matsui, H. Urease as a nanoreactor for growing crystalline ZnO nanoshells at room temperature. Angew. Chem. Int. Ed. 47, 5415–5417 (2008).
Pejoux, C., de la Rica, R. & Matsui, H. Biomimetic crystallization of sulfide semiconductor nanoparticles in aqueous solution. Small 6, 999–1002 (2010).
De la Rica, R., Fabijanic, K. I., Baldi, A. & Matsui, H. Biomimetic crystallization nanolithography: simultaneous nanopatterning and crystallization. Angew. Chem. Int. Ed. 49, 1447–1450 (2010).
Fan, R. et al. Integrated barcode chips for rapid, multiplexed analysis of proteins in microliter quantities of blood. Nature Biotechnol. 26, 1373–1378 (2008).
Acknowledgements
The authors thank G. S. Cooke and S. Kaye (Faculty of Medicine, Imperial College London) for the kind gift of human samples containing different viral loads. M.M.S. thanks the Engineering and Physical Sciences Research Council (EPSRC) and the European Research Council (ERC) starting investigator grant ‘Naturale’ for funding. This research was supported by a Marie Curie Intra European Fellowship within the 7th European Community Framework Programme (R.R.). The monoclonal antibody to HIV-1 p24 from S. Zolla-Pazner was provided by the Centre for AIDS Reagents, National Institute for Biological Standards and Control (NIBSC, a centre of the UK Health Protection Agency).
Author information
Authors and Affiliations
Contributions
R.R. elaborated the concept, designed and performed experiments, and wrote the paper. M.M.S. supervised the project, participated in scientific discussions, and revised the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 14350 kb)
Rights and permissions
About this article
Cite this article
de la Rica, R., Stevens, M. Plasmonic ELISA for the ultrasensitive detection of disease biomarkers with the naked eye. Nature Nanotech 7, 821–824 (2012). https://doi.org/10.1038/nnano.2012.186
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2012.186
This article is cited by
-
Policy Guidelines for Smart Sanitation Technology as a Public Health Tool
Digital Society (2024)
-
Immobilization of azide-functionalized proteins to micro- and nanoparticles directly from cell lysate
Microchimica Acta (2024)
-
Naked-eye observation of water-forming reaction on palladium etalon: transduction of gas-matter reaction into light-matter interaction
PhotoniX (2023)
-
Molecular-electromechanical system for unamplified detection of trace analytes in biofluids
Nature Protocols (2023)
-
Direct digital sensing of protein biomarkers in solution
Nature Communications (2023)