Abstract
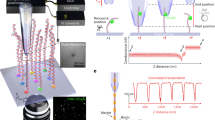
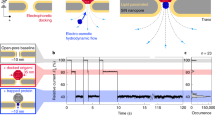
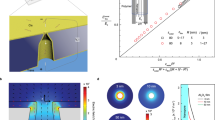
The development of solid-state nanopores1,2,3,4,5,6,7, inspired by their biological counterparts8,9,10,11,12,13,14,15, shows great potential for the study of single macromolecules16,17,18,19,20,21. Applications such as DNA sequencing6,22,23 and the exploration of protein folding6 require control of the dynamics of the molecule's interaction with the pore, but DNA capture by a solid-state nanopore is not well understood24,25,26. By recapturing individual molecules soon after they pass through a nanopore, we reveal the mechanism by which double-stranded DNA enters the pore. The observed recapture rates and times agree with solutions of a drift-diffusion model. Electric forces draw DNA to the pore over micrometer-scale distances, and upon arrival at the pore, molecules begin translocation almost immediately. Repeated translocation of the same molecule improves measurement accuracy, offers a way to probe the chemical transformations and internal dynamics of macromolecules on sub-millisecond time and sub-micrometre length scales, and demonstrates the ability to trap, study and manipulate individual macromolecules in solution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Li, J. et al. Ion-beam sculpting at nanometre length scales. Nature 412, 166–169 (2001).
Storm, A. J., Chen, J. H., Ling, X. S., Zandbergen, H. W. & Dekker, C. Electron-beam-induced deformations of SiO2 nanostructures. J. Appl. Phys. 98, 014307 (2005).
Wu, M. Y., Krapf, D., Zandbergen, M., Zandbergen, H. & Batson, P. E. Formation of nanopores in a SiN/SiO2 membrane with an electron beam. Appl. Phys. Lett. 87, 113106 (2005).
Chen, P. et al. Atomic layer deposition to fine-tune the surface properties and diameters of fabricated nanopores. Nano Lett. 4, 1333–1337 (2004).
Lo, C. J., Aref, T. & Bezryadin, A. Fabrication of symmetric sub-5 nm nanopores using focused ion and electron beams. Nanotechnology 17, 3264–3267 (2006).
Dekker, C. Solid-state nanopores. Nature Nanotech. 2, 209–215 (2007).
Martin, C. R. & Siwy, Z. S. Chemistry: Learning nature's way: Biosensing with synthetic nanopores. Science 317, 331–332 (2007).
Bezrukov, S. M., Vodyanoy, I. & Parsegian, V. A. Counting polymers moving through a single-ion channel. Nature 370, 279–281 (1994).
Kasianowicz, J. J., Brandin, E., Branton, D. & Deamer, D. W. Characterization of individual polynucleotide molecules using a membrane channel. Proc. Natl Acad. Sci. USA 93, 13770–13773 (1996).
Gu, L. Q., Braha, O., Conlan, S., Cheley, S. & Bayley, H. Stochastic sensing of organic analytes by a pore-forming protein containing a molecular adapter. Nature 398, 686–690 (1999).
Bayley, H. & Cremer, P. S. Stochastic sensors inspired by biology. Nature 413, 226–230 (2001).
Meller, A., Nivon, L. & Branton, D. Voltage-driven DNA translocations through a nanopore. Phys. Rev. Lett. 86, 3435–3438 (2001).
Bates, M., Burns, M. & Meller, A. Dynamics of DNA molecules in a membrane channel probed by active control techniques. Biophys. J. 84, 2366–2372 (2003).
Ambjornsson, T., Apell, S. P., Konkoli, Z., Di Marzio, E. A. & Kasianowicz, J. J. Charged polymer membrane translocation. J. Chem. Phys. 117, 4063–4073 (2002).
Henrickson, S. E., Misakian, M., Robertson, B. & Kasianowicz, J. J. Driven DNA transport into an asymmetric nanometer-scale pore. Phys. Rev. Lett. 85, 3057–3060 (2000).
Li, J. L., Gershow, M., Stein, D., Brandin, E. & Golovchenko, J. A. DNA molecules and configurations in a solid-state nanopore microscope. Nature Mater. 2, 611–615 (2003).
Han, A. P. et al. Sensing protein molecules using nanofabricated poresn. Appl. Phys. Lett. 88, 093901 (2006).
Fologea, D., Ledden, B., McNabb, D. S. & Li, J. Electrical characterization of protein molecules by a solid-state nanopore. Appl. Phys. Lett. 91, 053901 (2007).
Fologea, D. et al. Detecting single stranded DNA with a solid state nanopore. Nano Lett. 5, 1905–1909 (2005).
Fologea, D., Uplinger, J., Thomas, B., McNabb, D. S. & Li, J. L. Slowing DNA translocation in a solid-state nanopore. Nano Lett. 5, 1734–1737 (2005).
Storm, A. J. et al. Fast DNA translocation through a solid-state nanopore. Nano Lett. 5, 1193–1197 (2005).
Rhee, M. & Burns, M. A. Nanopore sequencing technology: research trends and applications. Trends Biotechnol. 24, 580–586 (2006).
Lagerqvist, J., Zwolak, M. & Di Ventra, M. Fast DNA sequencing via transverse electronic transport. Nano Lett. 6, 779–782 (2006).
Muthukumar, M. Polymer translocation through a hole. J. Chem. Phys. 111, 10371–10374 (1999).
Chen, P. et al. Probing single DNA molecule transport using fabricated nanopores. Nano Lett. 4, 2293–2298 (2004).
Keyser, U. F. et al. Direct force measurements on DNA in a solid-state nanopore. Nature Phys. 2, 473–477 (2006).
Storm, A. J., Chen, J. H., Zandbergen, H. W. & Dekker, C. Translocation of double-strand DNA through a silicon oxide nanopore. Phys. Rev. E 71, 051903 (2005).
Nkodo, A. E. et al. Diffusion coefficient of DNA molecules during free solution electrophoresis. Electrophoresis 22, 2424–2432 (2001).
Stellwagen, N. C., Gelfi, C. & Righetti, P. G. The free solution mobility of DNA. Biopolymers 42, 687–703 (1997).
Nakane, J., Akeson, M. & Marziali, A. Evaluation of nanopores as candidates for electronic analyte detection. Electrophoresis 23, 2592–2601 (2002).
Hall, J. E. Access resistance of a small circular pore. J. Gen. Physiol. 66, 531–532 (1975).
Panwar, A. S. & Kumar, S. Time scales in polymer electrophoresis through narrow constrictions: A Brownian dynamics study. Macromolecules 39, 1279–1289 (2006).
Grosberg, A. Y. & Khokhlov, A. R. Statistical Physics of Macromolecules (AIP Press, Woodbury, NY, 1994).
Acknowledgements
This work was supported by NIH/NGRI grant no. 5 R01 HG00370302. Some fabrication was carried out at Harvard University's Center for Nanoscale Systems, with the assistance of D. Bell, Yuan Lu and J.D. Deng. We thank E. Brandin for preparing the molecules for the trapping experiment, S. Coutreau and P. Testa for machining assistance, Jiali Li, D. Branton, S. Bezrukov, D. Hoogerheide and D. Vlassarev for useful discussions, and M. Biercuk for valuable suggestions regarding the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
Supplementary experimental methods and supplementary figures S1–S3 (PDF 409 kb)
Rights and permissions
About this article
Cite this article
Gershow, M., Golovchenko, J. Recapturing and trapping single molecules with a solid-state nanopore. Nature Nanotech 2, 775–779 (2007). https://doi.org/10.1038/nnano.2007.381
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2007.381
This article is cited by
-
Driven translocation of a semiflexible polymer through a conical channel in the presence of attractive surface interactions
Scientific Reports (2022)
-
Elucidating the dynamics of polymer transport through nanopores using asymmetric salt concentrations
Nano Research (2022)
-
On demand delivery and analysis of single molecules on a programmable nanopore-optofluidic device
Nature Communications (2019)
-
Label-free analysis of physiological hyaluronan size distribution with a solid-state nanopore sensor
Nature Communications (2018)
-
Conic shapes have higher sensitivity than cylindrical ones in nanopore DNA sequencing
Scientific Reports (2018)