Abstract
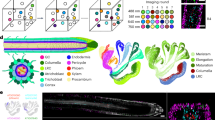
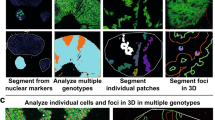
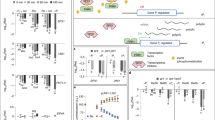
We present the coupled use of specifically localized fluorescent gene markers and image processing for automated quantitative analysis of cell growth and genetic activity across living plant tissues. We used fluorescent protein markers to identify cells, create seeds and boundaries for the automatic segmentation of cell geometries and ratiometrically measure gene expression cell by cell in Arabidopsis thaliana.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Caselles, V., Kimmel, R. & Sapiro, G. Int. J. Comput. Vis. 22, 61–79 (1997).
Otsu, N. IEEE Trans. Syst. Man Cybern. 9, 62–66 (1979).
Kapur, J.N., Sahoo, P.K. & Wong, A.K.C. Comput. Vis. Graph. Image Process 29, 273–285 (1985).
Rittscher, J. Ann. Rev. Biomed. Eng. 12, 315–344 (2010).
Makowski, P. et al. Comput. Med. Imaging Graph. 26, 9–17 (2002).
Beemster, G.T. & Baskin, T.I. Plant Physiol. 116, 1515–1526 (1998).
Laskowski, M. et al. PLoS Biol. 6, 307 (2008).
Ottenschläger, I. et al. Proc. Natl. Acad. Sci. USA 100, 2987–2991 (2003).
Jones, A.R. et al. Nat. Cell Biol. 11, 78–84 (2009).
Kass, M., Witkin, A. & Terzopoulos, D. Int. J. Computer Vision 1, 321–331 (1988).
Mace, D.L. et al. Bioinformatics 22, e323–e331 (2006).
Jönsson, H. et al. Bioinformatics 21 (suppl. 1), i232–i240 (2005).
Kelly, J.R. et al. J. Biol. Eng. 3, 4 (2009).
Boisnard-Lorig, C. et al. Plant Cell 13, 495–509 (2001).
Kurup, S. et al. Plant J. 42, 444–453 (2005).
Boresi, A.P. & Chong, K.P. Elasticity in Engineering Mechanics (Wiley-IEEE, New York, 2000).
Sage, D., Neumann, F.R., Hediger, F., Gasser, S.M. & Unser, M. IEEE Trans. Image Process. 14, 1372–1383 (2005).
Bao, Z. et al. Proc. Natl. Acad. Sci. USA 103, 2707–2712 (2006).
Acknowledgements
This research was supported by the Biotechnology and Biological Sciences Research Council, and Engineering and Physical Sciences Research Council research grants (BBS/B/16720 and BEP/17053 to J.H.), Institut de Recherche pour le Développement (L.L). F.F.'s PhD scholarship was supported by the Bill and Melinda Gates foundation (Gates Cambridge Trust). The Scottish Crop Research Institute receives grant-in-aid support from the Scottish Government Rural and Environment Research and Analysis Directorate (Workpackage 1.7). M.H. acknowledges the Australian Research Council for funding. We thank B. Scheres and H. Hofhuis (Utrecht University) for DR5rev∷3×Venus-N7 seeds and R.Y. Tsien (University of California, San Diego) for the mRFP1 cassette.
Author information
Authors and Affiliations
Contributions
F.F. generated the transgenic lines, analyzed the data and performed experiments. L.D. created the computational tools and plugin for ImageJ, and analyzed the data. J.H. provided instructions and supervision. M.H. performed confocal imaging of shoot apical meristem shown in Figure 2 and Supplementary Figure 1. L.L. generated the pBI 35S∷H2B-RFP binary vector and the Arabidopsis 35S∷H2B-RFP transgenic line.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 (PDF 14667 kb)
Supplementary Movie 1
Time-lapse movie of 35S::GFPLTI6b and 35S::mCherry-H2B in the root apical meristem. Images were acquired every 3 min. This video is shown with illustrative purposes, and it was not used for data analysis owing to its short duration (1 h 24 min). (MOV 726 kb)
Supplementary Movie 2
Time-lapse imaging of 35S::GFPLTI6b and 35S::mCherry-H2B to automate cell tracking in the root apical meristem. Images were acquired every 3 min. Nuclei tracking used SpotTracker to extract nuclei trajectories and to initiate balloon positions. This video is shown with illustrative purposes, and it was not used for data analysis owing to its short duration (1 h 24 min). (MOV 809 kb)
Rights and permissions
About this article
Cite this article
Federici, F., Dupuy, L., Laplaze, L. et al. Integrated genetic and computation methods for in planta cytometry. Nat Methods 9, 483–485 (2012). https://doi.org/10.1038/nmeth.1940
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1940
This article is cited by
-
Extending resolution within a single imaging frame
Nature Communications (2022)
-
Cell kinetics of auxin transport and activity in Arabidopsis root growth and skewing
Nature Communications (2021)
-
Week-long imaging of cell divisions in the Arabidopsis root meristem
Plant Methods (2019)
-
Semi-automatic Method for Ca2+ Imaging Data Analysis of Maturing Human Embryonic Stem Cells-Derived Retinal Pigment Epithelium
Annals of Biomedical Engineering (2016)
-
Reporters for sensitive and quantitative measurement of auxin response
Nature Methods (2015)