Abstract
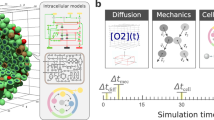
Cellular signaling processes depend on spatiotemporal distributions of molecular components. Multicolor, high-resolution microscopy permits detailed assessment of such distributions, providing input for fine-grained computational models that explore mechanisms governing dynamic assembly of multimolecular complexes and their role in shaping cellular behavior. However, it is challenging to incorporate into such models both complex molecular reaction cascades and the spatial localization of signaling components in dynamic cellular morphologies. Here we introduce an approach to address these challenges by automatically generating computational representations of complex reaction networks based on simple bimolecular interaction rules embedded into detailed, adaptive models of cellular morphology. Using examples of receptor-mediated cellular adhesion and signal-induced localized mitogen-activated protein kinase (MAPK) activation in yeast, we illustrate the capacity of this simulation technique to provide insights into cell biological processes. The modeling algorithms, implemented in a new version of the Simmune toolset, are accessible through intuitive graphical interfaces and programming libraries.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Lingwood, D. & Simons, K. Lipid rafts as a membrane-organizing principle. Science 327, 46–50 (2010).
Kholodenko, B.N. Four-dimensional organization of protein kinase signaling cascades: the roles of diffusion, endocytosis and molecular motors. J. Exp. Biol. 206, 2073–2082 (2003).
Delon, J. & Germain, R.N. Information transfer at the immunological synapse. Curr. Biol. 10, R923–R933 (2000).
Jones, R.B., Gordus, A., Krall, J.A. & MacBeath, G. A quantitative protein interaction network for the ErbB receptors using protein microarrays. Nature 439, 168–174 (2006).
Brown, M.D. & Sacks, D.B. Protein scaffolds in MAP kinase signalling. Cell. Signal. 21, 462–469 (2009).
Hlavacek, W.S., Faeder, J.R., Blinov, M.L., Perelson, A.S. & Goldstein, B. The complexity of complexes in signal transduction. Biotechnol. Bioeng. 84, 783–794 (2003).
Meier-Schellersheim, M. et al. Key role of local regulation in chemosensing revealed by a new molecular interaction-based modeling method. PLoS Comput. Biol. 2, e82 (2006).
Hlavacek, W.S. et al. Rules for modeling signal-transduction systems. Sci. STKE 2006, re6 (2006).
Lok, L. & Brent, R. Automatic generation of cellular reaction networks with Moleculizer 1.0. Nat. Biotechnol. 23, 131–136 (2005).
Feret, J., Danos, V., Krivine, J., Harmer, R. & Fontana, W. Internal coarse-graining of molecular systems. Proc. Natl. Acad. Sci. USA 106, 6453–6458 (2009).
Koschorreck, M. & Gilles, E.D. ALC: automated reduction of rule-based models. BMC Syst. Biol. 2, 91 (2008).
Mallavarapu, A., Thomson, M., Ullian, B. & Gunawardena, J. Programming with models: modularity and abstraction provide powerful capabilities for systems biology. J. R. Soc. Interface 6, 257–270 (2009).
Varma, R., Campi, G., Yokosuka, T., Saito, T. & Dustin, M.L. T cell receptor-proximal signals are sustained in peripheral microclusters and terminated in the central supramolecular activation cluster. Immunity 25, 117–127 (2006).
van Zon, J.S. & ten Wolde, P.R. Green's-function reaction dynamics: a particle-based approach for simulating biochemical networks in time and space. J. Chem. Phys. 123, 234910 (2005).
Andrews, S.S., Addy, N.J., Brent, R. & Arkin, A.P. Detailed simulations of cell biology with Smoldyn 2.1. PLoS Comput. Biol. 6, e1000705 (2010).
Gumbiner, B.M. Regulation of cadherin-mediated adhesion in morphogenesis. Nat. Rev. Mol. Cell Biol. 6, 622–634 (2005).
Sivasankar, S., Zhang, Y., Nelson, W.J. & Chu, S. Characterizing the initial encounter complex in cadherin adhesion. Structure 17, 1075–1081 (2009).
Cavey, M. & Lecuit, T. Molecular bases of cell-cell junctions stability and dynamics. Cold Spring Harb. Perspect. Biol. 1, a002998 (2009).
Zhang, Y., Sivasankar, S., Nelson, W.J. & Chu, S. Resolving cadherin interactions and binding cooperativity at the single-molecule level. Proc. Natl. Acad. Sci. USA 106, 109–114 (2009).
Troyanovsky, S. Cadherin dimers in cell-cell adhesion. Eur. J. Cell Biol. 84, 225–233 (2005).
Adams, C.L., Chen, Y.T., Smith, S.J. & Nelson, W.J. Mechanisms of epithelial cell-cell adhesion and cell compaction revealed by high-resolution tracking of E-cadherin-green fluorescent protein. J. Cell Biol. 142, 1105–1119 (1998).
Perez, T.D., Tamada, M., Sheetz, M.P. & Nelson, W.J. Immediate-early signaling induced by E-cadherin engagement and adhesion. J. Biol. Chem. 283, 5014–5022 (2008).
Glazier, J.A. & Graner, F. Simulation of the differential adhesion driven rearrangement of biological cells. Phys. Rev. E Stat. Phys. Plasmas Fluids Relat. Interdiscip. Topics 47, 2128–2154 (1993).
Beltman, J.B., Maree, A.F., Lynch, J.N., Miller, M.J. & de Boer, R.J. Lymph node topology dictates T cell migration behavior. J. Exp. Med. 204, 771–780 (2007).
Perez-Moreno, M., Jamora, C. & Fuchs, E. Sticky business: orchestrating cellular signals at adherens junctions. Cell 112, 535–548 (2003).
Hong, S., Troyanovsky, R.B. & Troyanovsky, S.M. Spontaneous assembly and active disassembly balance adherens junction homeostasis. Proc. Natl. Acad. Sci. USA 107, 3528–3533 (2010).
Dohlman, H.G. & Thorner, J.W. Regulation of G protein-initiated signal transduction in yeast: paradigms and principles. Annu. Rev. Biochem. 70, 703–754 (2001).
Kofahl, B. & Klipp, E. Modelling the dynamics of the yeast pheromone pathway. Yeast 21, 831–850 (2004).
Shao, D., Zheng, W., Qiu, W., Ouyang, Q. & Tang, C. Dynamic studies of scaffold-dependent mating pathway in yeast. Biophys. J. 91, 3986–4001 (2006).
Maeder, C.I. et al. Spatial regulation of Fus3 MAP kinase activity through a reaction-diffusion mechanism in yeast pheromone signalling. Nat. Cell Biol. 9, 1319–1326 (2007).
Good, M., Tang, G., Singleton, J., Remenyi, A. & Lim, W.A. The Ste5 scaffold directs mating signaling by catalytically unlocking the Fus3 MAP kinase for activation. Cell 136, 1085–1097 (2009).
Bhattacharyya, R.P. et al. The Ste5 scaffold allosterically modulates signaling output of the yeast mating pathway. Science 311, 822–826 (2006).
Evans, R. et al. Integrins in immunity. J. Cell Sci. 122, 215–225 (2009).
Eymard, R., Gallouet, T. & Herbin, R. Finite volume methods. in Handbook of Numerical Analysis (eds. Ciarlet, P.G. & Lions, J.L.) 7, 713–1020 (2000).
Novak, I.L. et al. Diffusion on a curved surface coupled to diffusion in the volume: application to cell biology. J. Comput. Phys. 226, 1271–1290 (2007).
Acknowledgements
We thank R. Schwartz, R. Varma, A. Nita-Lazar, I. Fraser, J. Tsang and D. Cioffi for helpful comments, members of the Laboratory for Systems Biology at the US National Institute of Allergy and Infectious Diseases for insightful discussions, G. Mack for advice during the early stages of Simmune development and A. Meier-Schellersheim for creating Figure 2 and Supplementary Figure 1. This work was supported by the Intramural Research Program of the US National Institute of Allergy and Infectious Diseases of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
B.R.A. wrote the application for designing cellular morphologies. A.D.G. wrote the simulator graphical user interface. M.M.-S. and F.K. developed the volume and membrane spatial discretization. B.R.A. developed the adaptive membrane diffusion discretization. B.R.A. and T.P. performed numerical tests. F.Z. wrote the application for defining molecular interactions. M.M.-S. developed the spatially adaptive network generator. B.R.A. and M.M.-S. developed the integrator drivers. B.R.A., F.K., A.D.G., T.P., F.Z., R.N.G. and M.M.S. discussed the project and wrote the manuscript. M.M.S. conceived and supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Notes 1–6 (PDF 12328 kb)
Supplementary Movie 1
E-cadherin–driven contact formation between two cells. (MOV 1145 kb)
Supplementary Movie 2
E-cadherin–driven contact formation in a segment of a multicellular layer. (MOV 1431 kb)
Rights and permissions
About this article
Cite this article
Angermann, B., Klauschen, F., Garcia, A. et al. Computational modeling of cellular signaling processes embedded into dynamic spatial contexts. Nat Methods 9, 283–289 (2012). https://doi.org/10.1038/nmeth.1861
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1861
This article is cited by
-
Absolute protein quantitation of the mouse macrophage Toll-like receptor and chemotaxis pathways
Scientific Data (2022)
-
The Perseus computational platform for comprehensive analysis of (prote)omics data
Nature Methods (2016)
-
Simulating tissue mechanics with agent-based models: concepts, perspectives and some novel results
Computational Particle Mechanics (2015)
-
Strategies for Efficient Numerical Implementation of Hybrid Multi-scale Agent-Based Models to Describe Biological Systems
Cellular and Molecular Bioengineering (2015)