Abstract
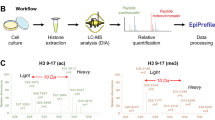
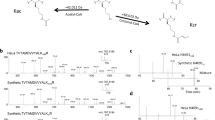
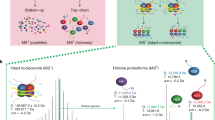
We developed a platform using hydrophilic interaction chromatography and high-resolution tandem mass spectrometry (MS) for analyses of histone H3 that allows comprehensive characterization of 'histone codes' at the molecular level. We identified over 150 differentially modified forms of histone H3.2 in asynchronously grown and butyrate-treated HeLa cells, revealing pervasive combinatorial modification previously unaccounted for by other techniques and providing a clarified estimate of the molecular diversity of histone H3 in mammals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Margueron, R., Trojer, P. & Reinberg, D. Curr. Opin. Genet. Dev. 15, 163–176 (2005).
Jenuwein, T. & Allis, C.D. Science 293, 1074–1080 (2001).
Coon, J.J. et al. Proc. Natl. Acad. Sci. USA 102, 9463–9468 (2005).
Pesavento, J.J., Kim, Y.B., Taylor, G.K. & Kelleher, N.L. J. Am. Chem. Soc. 126, 3386–3387 (2004).
Thomas, C.E., Kelleher, N.L. & Mizzen, C.A. J. Proteome Res. 5, 240–247 (2006).
Lindner, H., Sarg, B., Meraner, C. & Helliger, W. J. Chromatogr. A. 743, 137–144 (1996).
Mizzen, C.A., Alpert, A.J., Levesque, L., Kruck, T.P. & McLachlan, D.R. J. Chromatogr. B Biomed. Sci. Appl. 744, 33–46 (2000).
Pesavento, J.J., Mizzen, C.A. & Kelleher, N.L. Anal. Chem. 78, 4271–4280 (2006).
Loyola, A., Bonaldi, T., Roche, D., Imhof, A. & Almouzni, G. Mol. Cell 24, 309–316 (2006).
Zhang, K. et al. Mol. Cell. Proteomics 1, 500–508 (2002).
Sims, R.J., III & Reinberg, D. Genes Dev. 20, 2779–2786 (2006).
Bernstein, B.E. et al. Cell 120, 169–181 (2005).
Wysocka, J. et al. Nature 442, 86–90 (2006).
Shi, X. et al. Nature 442, 96–99 (2006).
Syka, J.E., Coon, J.J., Schroeder, M.J., Shabanowitz, J. & Hunt, D.F. Proc. Natl. Acad. Sci. USA 101, 9528–9533 (2004).
Acknowledgements
Funding from the US National Institutes of Health (NIH; GM 067193), the Packard Foundation, the Alfred P. Sloan Foundation and a 2004 Camille Dreyfus Teacher-Scholar Award from the Dreyfus Foundation to N.L.K., from the Roy J. Carver Charitable Trust (04-76) and the March of Dimes (Basil O'Connor Scholar Award 5-FY05-1232) to C.A.M., and from the Institute of Genomic Biology at the University of Illinois and the NIH (1 F32 GM 078942-01) to B.A.G. are gratefully acknowledged. J.J.P. is a recipient of an NIH Institutional National Research Service award in Molecular Biophysics (5T32 GM 08276).
Author information
Authors and Affiliations
Contributions
B.A.G. conducted experiments, interpreted data and wrote paper; J.J.P. conducted experiments; C.A.M. designed experiments and wrote paper; N.L.K. designed experiments, interpreted data and wrote paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
HILIC fractionation and mass spectrometry resolves isomeric differently modified histone H3 codes. (PDF 51 kb)
Supplementary Fig. 2
HILIC separation and mass spectrometry reveals interdependence of histone H3K4 methylation with acetylation. (PDF 59 kb)
Supplementary Fig. 3
Semi-quantitative determination of the relative abundances for the species at 5408 m/z from asynchronously grown and butyrate treated cells. (PDF 79 kb)
Supplementary Table 1
Histone H3.2 species separated by HILIC and characterized by ECD. (PDF 47 kb)
Supplementary Table 2
ECD fragment ions observed from histone H3.2 species from selected fractions 1-10. (PDF 122 kb)
Supplementary Table 3
Semi-quantitative relative abundance values for selected histone H3.2 species from asynchronously grown and butyrate treated HeLa cells separated by HILIC and characterized by ECD. (PDF 10 kb)
Rights and permissions
About this article
Cite this article
Garcia, B., Pesavento, J., Mizzen, C. et al. Pervasive combinatorial modification of histone H3 in human cells. Nat Methods 4, 487–489 (2007). https://doi.org/10.1038/nmeth1052
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth1052
This article is cited by
-
Decoding chromatin states by proteomic profiling of nucleosome readers
Nature (2024)
-
Epigenetic modifications of histones in cancer
Genome Biology (2019)
-
TagGraph reveals vast protein modification landscapes from large tandem mass spectrometry datasets
Nature Biotechnology (2019)
-
One-Pot Quantitative Top- and Middle-Down Analysis of GluC-Digested Histone H4
Journal of the American Society for Mass Spectrometry (2019)
-
How many human proteoforms are there?
Nature Chemical Biology (2018)