Abstract
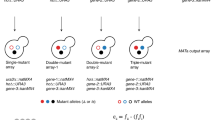
Epistasis analysis, which reports on the extent to which the function of one gene depends on the presence of a second, is a powerful tool for studying the functional organization of the cell. Systematic genome-wide studies of epistasis, however, have been limited, with the majority of data being collected in the budding yeast, Saccharomyces cerevisiae. Here we present two 'pombe epistasis mapper' strategies, PEM-1 and PEM-2, which allow for high-throughput double mutant generation in the fission yeast, S. pombe. These approaches take advantage of a previously undescribed, recessive, cycloheximide-resistance mutation. Both systems can be used for genome-wide screens or for the generation of high-density, quantitative epistatic miniarray profiles (E-MAPs). Since S. cerevisiae and S. pombe are evolutionary distant, this methodology will provide insight into conserved biological pathways that are present in S. pombe, but not S. cerevisiae, and will enable a comprehensive analysis of the conservation of genetic interaction networks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Tong, A.H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Tong, A.H. et al. Global mapping of the yeast genetic interaction network. Science 303, 808–813 (2004).
Pan, X. et al. A DNA integrity network in the yeast Saccharomyces cerevisiae. Cell 124, 1069–1081 (2006).
Pan, X. et al. A robust toolkit for functional profiling of the yeast genome. Mol. Cell 16, 487–496 (2004).
Kelley, R. & Ideker, T. Systematic interpretation of genetic interactions using protein networks. Nat. Biotechnol. 23, 561–566 (2005).
Schuldiner, M. et al. Exploration of the function and organization of the yeast early secretory pathway through an epistatic miniarray profile. Cell 123, 507–519 (2005).
Collins, S.R. et al. Functional dissection of protein complexes involved in yeast chromosome biology using a genetic interaction map. Nature 446, 806–810 (2007).
Collins, S.R., Schuldiner, M., Krogan, N.J. & Weissman, J.S. A strategy for extracting and analyzing large-scale quantitative epistatic interaction data. Genome Biol. 7, R63 (2006).
Schuldiner, M., Collins, S.R., Weissman, J.S. & Krogan, N.J. Quantitative genetic analysis in Saccharomyces cerevisiae using epistatic miniarray profiles (E-MAPs) and its application to chromatin functions. Methods 40, 344–352 (2006).
Sipiczki, M. Where does fission yeast sit on the tree of life? Genome Biol. 1, reviews1011.1–1011.4 (2000).
Pluta, A.F., Mackay, A.M., Ainsztein, A.M., Goldberg, I.G. & Earnshaw, W.C. The centromere: hub of chromosomal activities. Science 270, 1591–1594 (1995).
Grewal, S.I. & Jia, S. Heterochromatin revisited. Nat. Rev. Genet. 8, 35–46 (2007).
Hentges, P., Van Driessche, B., Tafforeau, L., Vandenhaute, J. & Carr, A.M. Three novel antibiotic marker cassettes for gene disruption and marker switching in Schizosaccharomyces pombe. Yeast 22, 1013–1019 (2005).
Ekwall, K. & Ruusala, T. Budding yeast CAN1 gene as a selection marker in fission yeast. Nucleic Acids Res. 19, 1150 (1991).
Kaufer, N.F., Fried, H.M., Schwindinger, W.F., Jasin, M. & Warner, J.R. Cycloheximide resistance in yeast: the gene and its protein. Nucleic Acids Res. 11, 3123–3135 (1983).
Ibrahim, M.A. & Coddington, A. Genetic studies on cycloheximide-resistant strains of Schizosaccharomyces pombe. Heredity 37, 179–191 (1976).
Spahn, C.M. et al. Domain movements of elongation factor eEF2 and the eukaryotic 80S ribosome facilitate tRNA translocation. EMBO J. 23, 1008–1019 (2004).
Stevens, D.R., Atteia, A., Franzen, L.G. & Purton, S. Cycloheximide resistance conferred by novel mutations in ribosomal protein L41 of Chlamydomonas reinhardtii. Mol. Gen. Genet. 264, 790–795 (2001).
Varma, A. & Kwon-Chung, K.J. Characterization of the L41 gene in Cryptococcus neoformans: its application as a selectable transformation marker for cycloheximide resistance. Yeast 16, 1397–1403 (2000).
Kawai, S. et al. Drastic alteration of cycloheximide sensitivity by substitution of one amino acid in the L41 ribosomal protein of yeasts. J. Bacteriol. 174, 254–262 (1992).
Tanaka, K., Davey, J., Imai, Y. & Yamamoto, M. Schizosaccharomyces pombe map3+ encodes the putative M-factor receptor. Mol. Cell. Biol. 13, 80–88 (1993).
Styrkarsdottir, U., Egel, R. & Nielsen, O. The smt-0 mutation which abolishes mating-type switching in fission yeast is a deletion. Curr. Genet. 23, 184–186 (1993).
Doe, C.L. & Whitby, M.C. The involvement of Srs2 in post-replication repair and homologous recombination in fission yeast. Nucleic Acids Res. 32, 1480–1491 (2004).
Edwards, R.J., Bentley, N.J. & Carr, A.M.A. Rad3-Rad26 complex responds to DNA damage independently of other checkpoint proteins. Nat. Cell Biol. 1, 393–398 (1999).
Torres, J.Z., Schnakenberg, S.L. & Zakian, V.A. Saccharomyces cerevisiae Rrm3p DNA helicase promotes genome integrity by preventing replication fork stalling: viability of rrm3 cells requires the intra-S-phase checkpoint and fork restart activities. Mol. Cell. Biol. 24, 3198–3212 (2004).
Moreno, S., Klar, A. & Nurse, P. Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol. 194, 795–823 (1991).
Acknowledgements
We thank A. Carr and O. Nielsen (pON177; University of Copenhagen) for providing reagents; S. Forsburg (FY1524 strain; University of Southern California) and K. Ekwall for reagents and discussion; C.J. Ingles, G. Cagney and D. Fiedler for critical reading of the manuscript and M. Shales for help with figures. J.S.W. is funded by the Howard Hughes Medical Institute, N.J.K. used funds from a Sandler Family Fellowship and A.R. was funded by a Howard Hughes Medical Institute postdoctoral fellowship. The work was also supported by funds from the California Institute of Quantitative Biology.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors plan on submitting a patent application for the genetic strategy described in this manuscript.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Table 1, Supplementary Protocols 1–2 (PDF 235 kb)
Rights and permissions
About this article
Cite this article
Roguev, A., Wiren, M., Weissman, J. et al. High-throughput genetic interaction mapping in the fission yeast Schizosaccharomyces pombe. Nat Methods 4, 861–866 (2007). https://doi.org/10.1038/nmeth1098
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth1098
This article is cited by
-
The SAGA histone acetyltransferase module targets SMC5/6 to specific genes
Epigenetics & Chromatin (2023)
-
Synthetic negative genome screen of the GPN-loop GTPase NPA3 in Saccharomyces cerevisiae
Current Genetics (2022)
-
PHENOS: a high-throughput and flexible tool for microorganism growth phenotyping on solid media
BMC Microbiology (2018)
-
A systematic genetic screen identifies new factors influencing centromeric heterochromatin integrity in fission yeast
Genome Biology (2014)
-
Evolutionarily conserved genetic interactions with budding and fission yeast MutS identify orthologous relationships in mismatch repair-deficient cancer cells
Genome Medicine (2014)