Abstract
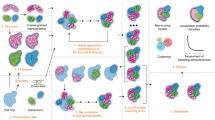
Although three-dimensional electron microscopy (3D-EM) permits structural characterization of macromolecular assemblies in distinct functional states, the inability to classify projections from structurally heterogeneous samples has severely limited its application. We present a maximum likelihood–based classification method that does not depend on prior knowledge about the structural variability, and demonstrate its effectiveness for two macromolecular assemblies with different types of conformational variability: the Escherichia coli ribosome and Simian virus 40 (SV40) large T-antigen.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Frank, J. Three-dimensional Electron Microscopy of Macromolecular Assemblies (Oxford University Press, New York, 2006).
Orlova, E.V. & Saibil, H.R. Curr. Opin. Struct. Biol. 14, 584–590 (2004).
Heymann, J.B., Conway, J.F. & Steven, A.C. J. Struct. Biol. 147, 291–301 (2004).
Gao, H., Valle, M., Ehrenberg, M. & Frank, J. J. Struct. Biol. 147, 283–290 (2004).
Brink, J. et al. Structure 12, 185–191 (2004).
Rice, J.A. Mathematical Statistics and Data Analysis (Duxbury Press, Belmost, 1995).
Sigworth, F.J. J. Struct. Biol. 122, 328–339 (1998).
Yin, Z., Zheng, Y., Doerschuk, P.C., Natarajan, P. & Johnson, J.E. J. Struct. Biol. 144, 24–50 (2003).
Scheres, S.H.W. et al. J. Mol. Biol. 348, 139–149 (2005).
Scheres, S.H.W., Valle, M. & Carazo, J.M. Bioinformatics 21 (Suppl. 2), ii243–ii244 (2005).
Dempster, A.P., Laird, N.M. & Rubin, D.B. J. Royal Statist. Soc. Ser. B 39, 1–38 (1977).
Valle, M. et al. EMBO J. 21, 3557–3567 (2002).
Sorzano, C.O.S. et al. J. Struct. Biol. 148, 194–204 (2004).
Gomez-Lorenzo, M.G. et al. EMBO J. 22, 6205–6213 (2003).
Valle, M. et al. Cell 114, 123–134 (2003).
Acknowledgements
We thank the Department of Computer Architecture and Electronics of the University of Almeria, and the Barcelona Supercomputing Center (Centro Nacional de Supercomputación) for providing computing resources, and T. Elfving for his help with weighted least-squares minimization. This work was supported by grants from the European Union (FP6-502828 and EGEE2-031688 to JMC), US National Institutes of Health (HL70472 to G.T.H. and J.M.C., and P41 RR01219 to J.F.), Howard Hughes Medical Institute (to J.F.) and the Spanish Comisión Interministerial de Ciencia y Tecnología (BFU2004-00217 to J.M.C.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Preliminary reconstruction of the ribosome dataset.
Supplementary Fig. 2
Likelihood optimization for the ribosome dataset.
Supplementary Fig. 3
Supervised classification of the ribosome dataset.
Supplementary Fig. 4
Likelihood optimization for the large T-antigen dataset.
Supplementary Note
Mathematical background and implementation details.
Rights and permissions
About this article
Cite this article
Scheres, S., Gao, H., Valle, M. et al. Disentangling conformational states of macromolecules in 3D-EM through likelihood optimization. Nat Methods 4, 27–29 (2007). https://doi.org/10.1038/nmeth992
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth992
This article is cited by
-
Prediction of hydrophilic and hydrophobic hydration structure of protein by neural network optimized using experimental data
Scientific Reports (2023)
-
3DFlex: determining structure and motion of flexible proteins from cryo-EM
Nature Methods (2023)
-
Methods and application of coherent X-ray diffraction imaging of noncrystalline particles
Biophysical Reviews (2020)