Abstract
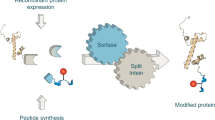
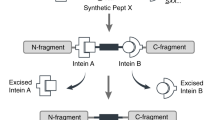
Biophysical techniques such as fluorescence spectroscopy and nuclear magnetic resonance (NMR) spectroscopy provide a window into the inner workings of proteins. These approaches make use of probes that can either be naturally present within the protein or introduced through a labeling procedure. In general, the more control one has over the type, location and number of probes in a protein, then the more information one can extract from a given biophysical analysis. Recently, two related approaches have emerged that allow proteins to be labeled with a broad range of physical probes. Expressed protein ligation (EPL) and protein trans-splicing (PTS) are both intein-based approaches that permit the assembly of a protein from smaller synthetic and/or recombinant pieces. Here we provide some guidelines for the use of EPL and PTS, and highlight how the dovetailing of these new protein chemistry methods with standard biophysical techniques has improved our ability to interrogate protein function, structure and folding.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hahn, M.E. & Muir, T.W. Manipulating proteins with chemistry: a cross-section of chemical biology. Trends Biochem. Sci. 30, 26–34 (2005).
Dawson, P.E., Muir, T.W., Clarklewis, I. & Kent, S.B.H. Synthesis of proteins by native chemical ligation. Science 266, 776–779 (1994).
Muir, T.W. Semisynthesis of proteins by expressed protein ligation. Annu. Rev. Biochem. 72, 249–289 (2003).
Paulus, H. Protein splicing and related forms of protein autoprocessing. Annu. Rev. Biochem. 69, 447–496 (2000).
Otomo, T., Ito, N., Kyogoku, Y. & Yamazaki, T. NMR observation of selected segments in a larger protein: central-segment isotope labeling through intein-mediated ligation. Biochemistry 38, 16040–16044 (1999).
Cotton, G.J., Ayers, B., Xu, R. & Muir, T.W. Insertion of a synthetic peptide into a recombinant protein framework: a protein biosensor. J. Am. Chem. Soc. 121, 1100–1101 (1999).
Blaschke, U.K., Cotton, G.J. & Muir, T.W. Synthesis of multi-domain proteins using expressed protein ligation: Strategies for segmental isotopic labeling of internal regions. Tetrahedron 56, 9461–9470 (2000).
Schwarzer, D. & Cole, P.A. Protein semisynthesis and expressed protein ligation: chasing a protein's tail. Curr. Opin. Chem. Biol. 9, 561–569 (2005).
Evans-Jr, T.C., Benner, J. & Xu, M.Q. Semisynthesis of cytotoxic proteins using a modified protein splicing element. Protein Sci. 7, 2256–2264 (1998).
Wu, W., Wood, D.W., Belfort, G., Derbyshire, V. & Belfort, M. Intein-mediated purification of cytotoxic endonuclease I-TevI by insertional inactivation and pH-controllable splicing. Nucleic Acids Res. 30, 4864–4871 (2002).
Yamazaki, T. et al. Segmental isotope labeling for protein NMR using peptide splicing. J. Am. Chem. Soc. 120, 5591–5592 (1998).
Xu, R., Ayers, B., Cowburn, D. & Muir, T.W. Chemical ligation of folded recombinant proteins: segmental isotopic labeling of domains for NMR studies. Proc. Natl. Acad. Sci. USA 96, 388–393 (1999).
Evans, T.C., Jr, Benner, J. & Xu, M-Q. The cyclization and polymerization of bacterially expressed proteins using modified self-splicing inteins. J. Biol. Chem. 274, 18359–18363 (1999).
Iwai, H. & Pluckthun, A. Circular β-lactamase: stability enhancement by cyclizing the backbone. FEBS Lett. 459, 166–172 (1999).
Scott, C.P., Abel-Santos, E., Wall, M., Wahnon, D.C. & Benkovic, S.J. Production of cyclic peptides and proteins in vivo. Proc. Natl. Acad. Sci. USA 96, 13638–13643 (1999).
Camarero, J.A. & Muir, T.W. Biosynthesis of a head-to-tail cyclized protein with improved biological activity. J. Am. Chem. Soc. 121, 5597–5598 (1999).
Evans, T.C., Jr et al. Protein trans-splicing and cyclization by a naturally split intein from the dnaE gene of Synechocystis Species PCC6803. J. Biol. Chem. 275, 9091–9094 (2000).
Camarero, J.A., Fushman, D., Cowburn, D. & Muir, T.W. Peptide chemical ligation inside living cells: in vivo generation of a circular protein domain. Bioorg. Med. Chem. 9, 2479–2484 (2001).
Camarero, J.A. et al. Rescuing a destabilized protein fold through backbone cyclization. J. Mol. Biol. 308, 1045–1062 (2001).
Iwai, H., Lingel, A. & Pluckthun, A. Cyclic green fluorescent protein produced in vivo using an artificially split PI-PfuI intein from Pyrococcus furiosus. J. Biol. Chem. 276, 16548–16554 (2001).
Scott, C.P., Abel-Santos, E., Jones, A.D. & Benkovic, S.J. Structural requirements for the biosynthesis of backbone cyclic peptide libraries. Chem. Biol. 8, 801–815 (2001).
Kinsella, T.M. et al. Retrovirally delivered random cyclic peptide libraries yield inhibitors of Interleukin-4 signaling in human B cells. J. Biol. Chem. 277, 37512–37518 (2002).
Williams, N.K. et al. Stabilization of native protein fold by intein-mediated covalent cyclization. J. Mol. Biol. 346, 1095–1108 (2005).
Kent, S.B.H. Chemical synthesis of peptides and proteins. Annu. Rev. Biochem. 57, 957–989 (1988).
Villain, M., Vizzavona, J. & Rose, K. Covalent capture: a new tool for the purification of synthetic and recombinant polypeptides. Chem. Biol. 8, 673–679 (2001).
Bang, D. & Kent, S.B.H. A one-pot total synthesis of crambin. Angew. Chem. Int. Edn. Engl. 43, 2534–2538 (2004).
McCaldon, P. & Argos, P. Oligopeptide biases in protein sequences and their use in predicting protein coding regions in nucleotide sequences. Proteins 4, 99–122 (1988).
Valiyaveetil, F.I., MacKinnon, R. & Muir, T.W. Semisynthesis and folding of the potassium channel KcsA. J. Am. Chem. Soc. 124, 9113–9120 (2002).
Yan, L.Z. & Dawson, P.E. Synthesis of peptides and proteins without cysteine residues by native chemical ligation combined with desulfurization. J. Am. Chem. Soc. 123, 526–533 (2001).
Canne, L.E., Bark, S.J. & Kent, S.B.H. Extending the applicability of native chemical ligation. J. Am. Chem. Soc. 118, 5891–5896 (1996).
Offer, J., Boddy, C.N.C. & Dawson, P.E. Extending synthetic access to proteins with a removable acyl transfer auxiliary. J. Am. Chem. Soc. 124, 4642–4646 (2002).
Saxon, E., Armstrong, J.I. & Bertozzi, C.R.A. “Traceless” Staudinger ligation for the chemoselective synthesis of amide bonds. Org. Lett. 2, 2141–2143 (2000).
Nilsson, B.L., Kiessling, L.L. & Raines, R.T. Staudinger ligation: a peptide from a thioester and azide. Org. Lett. 2, 1939–1941 (2000).
Chong, S., Williams, K.S., Wotkowicz, C. & Xu, M.Q. Modulation of protein splicing of the Saccharomyces cerevisiae vacuolar membrane ATPase intein. J. Biol. Chem. 273, 10567–10577 (1998).
Southworth, M.W., Amaya, K., Evans, T.C., Xu, M.Q. & Perler, F.B. Purification of proteins fused to either the amino or carboxy terminus of the Mycobacterium xenopi gyrase A intein. Biotechniques 27, 110–120 (1999).
Xu, M.Q., Paulus, H. & Chong, S. Fusions to self-splicing inteins for protein purification. Methods Enzymol. 326, 376–418 (2000).
Hackeng, T.M., Griffin, J.H. & Dawson, P.E. Protein synthesis by native chemical ligation: expanded scope by using straightforward methodology. Proc. Natl. Acad. Sci. USA 96, 10068–10073 (1999).
Botti, P., Villain, M., Manganiello, S. & Gaertner, H. Native chemical ligation through in situ O to S acyl shift. Org. Lett. 6, 4861–4864 (2004).
Ohta, Y. et al. Cysteine-derived S-protected oxazolidinones: potential chemical devices for the preparation of peptide thioesters. Org. Lett. 8, 467–470 (2006).
Muir, T.W., Sondhi, D. & Cole, P.A. Expressed protein ligation: a general method for protein engineering. Proc. Natl. Acad. Sci. USA 95, 6705–6710 (1998).
Camarero, J.A. et al. Autoregulation of a bacterial sigma factor explored by using segmental isotopic labeling and NMR. Proc. Natl. Acad. Sci. USA 99, 8536–8541 (2002).
Erlanson, D.A., Chytil, M. & Verdine, G.L. The leucine zipper domain controls the orientation of AP-1 in the NFAT center dot AP-1 center dot DNA complex. Chem. Biol. 3, 981–991 (1996).
Tolbert, T.J. & Wong, C.H. New methods for proteomic research: preparation of proteins with N-terminal cysteines for labeling and conjugation. Angew. Chem. Int. Edn. Engl. 41, 2171–2174 (2002).
Derbyshire, V. et al. Genetic definition of a protein-splicing domain: functional mini-inteins support structure predictions and a model for intein evolution. Proc. Natl. Acad. Sci. USA 94, 11466–11471 (1997).
Mathys, S. et al. Characterization of a self-splicing mini-intein and its conversion into autocatalytic N- and C-terminal cleavage elements: facile production of protein building blocks for protein ligation. Gene 231, 1–13 (1999).
Wood, D.W., Wu, W., Belfort, G., Derbyshire, V. & Belfort, M. A genetic system yields self-cleaving inteins for bioseparations. Nat. Biotechnol. 17, 889–892 (1999).
Romanelli, A., Shekhtman, A., Cowburn, D. & Muir, T.W. Semisynthesis of a segmental isotopically labeled protein splicing precursor: NMR evidence for an unusual peptide bond at the N-extein-intein junction. Proc. Natl. Acad. Sci. USA 101, 6397–6402 (2004).
Muralidharan, V. et al. Domain-specific incorporation of noninvasive optical probes into recombinant proteins. J. Am. Chem. Soc. 126, 14004–14012 (2004).
Beligere, G.S. & Dawson, P.E. Conformationally assisted protein ligation using C-terminal thioester peptides. J. Am. Chem. Soc. 121, 6332–6333 (1999).
Gentle, I.E., De Souza, D.P. & Baca, M. Direct production of proteins with N-terminal cysteine for site-specific conjugation. Bioconjug. Chem. 15, 658–663 (2004).
Shi, J.X. & Muir, T.W. Development of a tandem protein trans-splicing system based on native and engineered split inteins. J. Am. Chem. Soc. 127, 6198–6206 (2005).
Southworth, M.W. et al. Control of protein splicing by intein fragment reassembly. EMBO J. 17, 918–926 (1998).
Mills, K.V., Lew, B.M., Jiang, S.-q. & Paulus, H. Protein splicing in trans by purified N- and C-terminal fragments of the Mycobacterium tuberculosis RecA intein. Proc. Natl. Acad. Sci. USA 95, 3543–3548 (1998).
Sun, W., Yang, J., Liu, X-Q. Synthetic two-piece and three-piece split inteins for protein trans-splicing. J. Biol. Chem. 279, 35281–35286 (2004).
Wu, H., Hu, Z. & Liu, X-Q. Protein trans-splicing by a split intein encoded in a split DnaE gene of Synechocystis sp. PCC6803. Proc. Natl. Acad. Sci. USA 95, 9226–9231 (1998).
Zuger, S. & Iwai, H. Intein-based biosynthetic incorporation of unlabeled protein tags into isotopically labeled proteins for NMR studies. Nat. Biotechnol. 23, 736–740 (2005).
Mootz, H.D. & Muir, T.W. Protein splicing triggered by a small molecule. J. Am. Chem. Soc. 124, 9044–9045 (2002).
Ozawa, T., Kaihara, A., Sato, M., Tachihara, K. & Umezawa, Y. Split luciferase as an optical probe for detecting protein-protein interactions in mammalian cells based on protein splicing. Anal. Chem. 73, 2516–2521 (2001).
Giriat, I. & Muir, T.W. Protein semisynthesis in living cells. J. Am. Chem. Soc. 125, 7180–7181 (2003).
Chin, H.G. et al. Protein trans-splicing in transgenic plant chloroplast: reconstruction of herbicide resistance from split genes. Proc. Natl. Acad. Sci. USA 100, 4510–4515 (2003).
Yang, J., Fox, G.C., Jr & Henry-Smith, T.V. Intein-mediated assembly of a functional β-glucuronidase in transgenic plants. Proc. Natl. Acad. Sci. USA 100, 3513–3518 (2003).
Algire, M.A., Maag, D. & Lorsch, J.R. Pi release from eIF2, not GTP hydrolysis, is the step controlled by start-site selection during eukaryotic translation initiation. Mol. Cell 20, 251–262 (2005).
Maag, D., Fekete, C.A., Gryczynski, Z. & Lorsch, J.R. A conformational change in the eukaryotic translation preinitiation complex and release of eIF1 signal recognition of the start codon. Mol. Cell 17, 265–275 (2005).
Anderson, L.L., Marshall, G.R., Crocker, E., Smith, S.O. & Baranski, T.J. Motion of carboxyl terminus of Gα is restricted upon G protein activation. J. Biol. Chem. 280, 31019–31026 (2005).
Yagi, H., Tsujimoto, T., Yamazaki, T., Yoshida, M. & Akutsu, H. Conformational change of H+-ATPase β-monomer revealed on segmental isotope labeling NMR spectroscopy. J. Am. Chem. Soc. 126, 16632–16638 (2004).
Tatulian, S.A., Qin, S., Pande, A.H. & He, X.M. Positioning membrane proteins by novel protein engineering and biophysical approaches. J. Mol. Biol. 351, 939–947 (2005).
Wu, J-W., et al. Crystal Structure of a Phosphorylated Smad2: Recognition of phosphoserine by the MH2 domain and insights on Smad function in TGF-β signaling. Mol. Cell 8, 1277–1289 (2001).
Qin, B.Y., Lam, S.S., Correia, J.J. & Lin, K. Smad3 allostery links TGF-β receptor kinase activation to transcriptional control. Genes Dev. 16, 1950–1963 (2002).
Chacko, B.M. et al. Structural basis of heteromeric Smad protein assembly in TGF-β signaling. Mol. Cell 15, 813–823 (2004).
Rak, A. et al. Structure of Rab GDP-dissociation inhibitor in complex with prenylated YPT1 GTPase. Science 302, 646–650 (2003).
Clayton, D. et al. Total chemical synthesis and electrophysiological characterization of mechanosensitive channels from Escherichia coli and Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 101, 4764–4769 (2004).
Valiyaveetil, F.I., Sekedat, M., MacKinnon, R. & Muir, T.W. Glycine as a D-amino acid surrogate in the K+-selectivity filter. Proc. Natl. Acad. Sci. USA 101, 17045–17049 (2004).
Doyle, D.A. et al. The structure of the potassium channel: molecular basis of K+ conduction and selectivity. Science 280, 69–77 (1998).
Burbulis, I., Yamaguchi, K., Gordon, A., Carlson, R. & Brent, R. Using protein-DNA chimeras to detect and count small numbers of molecules. Nat. Methods 2, 31–37 (2005).
Lovrinovic, M. et al. Synthesis of protein-nucleic acid conjugates by expressed protein ligation. Chem. Commun. 822–823 (2003).
Chong, S. et al. Single-column purification of free recombinant proteins using a self-cleavable affinity tag derived from a protein splicing element. Gene 192, 271–281 (1997).
Shingledecker, K., Jiang, S.Q. & Paulus, H. Molecular dissection of the Mycobacterium tuberculosis RecA intein: design of a minimal intein and of a trans-splicing system involving two intein fragments. Gene 207, 187–195 (1998).
Evans, T.C., Jr, Benner, J. & Xu, M-Q. The in vitro ligation of bacterially expressed proteins using an intein from Methanobacterium thermoautotrophicum. J. Biol. Chem. 274, 3923–3926 (1999).
Camarero, J.A., Cotton, G.J., Adeva, A. & Muir, T.W. Chemical ligation of unprotected peptides directly from a solid support. J. Pept. Res. 51, 303–316 (1998).
Shin, Y. et al. Fmoc-based synthesis of peptide-(α)thioesters: Application to the total chemical synthesis of a glycoprotein by native chemical ligation. J. Am. Chem. Soc. 121, 11684–11689 (1999).
Ingenito, R., Bianchi, E., Fattori, D. & Pessi, A. Solid phase synthesis of peptide C-terminal thioesters by Fmoc/t-Bu chemistry. J. Am. Chem. Soc. 121, 11369–11374 (1999).
Alsina, J., Yokum, T.S., Albericio, F. & Barany, G. Backbone amide linker (BAL) strategy for N(α)-9-fluorenylmethoxycarbonyl (Fmoc) solid-phase synthesis of unprotected peptide p-nitroanilides and thioesters(1). J. Org. Chem. 64, 8761–8769 (1999).
Swinnen, D. & Hilvert, D. Facile, Fmoc-compatible solid-phase synthesis of peptide C-terminal thioesters. Org. Lett. 2, 2439–2442 (2000).
Camarero, J.A., Hackel, B.J., deYoreo, J.J. & Mitchell, A.R. Fmoc-based synthesis of peptide α-Thioesters using an aryl hydrazine support. J. Org. Chem. 69, 4145–4151 (2004).
Perler, F.B. InBase, the intein database. Nucleic Acids. Res. 30, 383–384 (2002).
Xu, M.Q. & Perler, F.B. The mechanism of protein splicing and its modulation by mutation. EMBO J. 15, 5146–5153 (1996).
Chong, S. et al. Utilizing the C-terminal cleavage activity of a protein splicing element to purify recombinant proteins in a single chromatographic step. Nucleic Acids Res. 26, 5109–5115 (1998).
Banki, M.R., Feng, L. & Wood, D.W. Simple bioseparations using self-cleaving elastin-like polypeptide tags. Nat. Methods 2, 659–662 (2005).
Ge, X. et al. Self-cleavable stimulus responsive tags for protein purification without chromatography. J. Am. Chem. Soc. 127, 11228–11229 (2005).
Liu, X-Q. & Yang, J. Split dnaE genes encoding multiple novel inteins in Trichodesmium erythraeum. J. Biol. Chem. 278, 26315–26318 (2003).
Nichols, N.M. & Evans, T.C., Jr . Mutational analysis of protein splicing, cleavage, and self-association reactions mediated by the naturally split Ssp DnaE intein. Biochemistry 43, 10265–10276 (2004).
Schnolzer, M. & Kent, S.B. Constructing proteins by dovetailing unprotected synthetic peptides: backbone-engineered HIV protease. Science 256, 221–225 (1992).
Gaertner, H.F. et al. Construction of protein analogs by site-specific Condensation of unprotected fragments. Bioconjug. Chem. 3, 262–268 (1992).
Hang, H.C. & Bertozzi, C.R. Chemoselective approaches to glycoprotein assembly. Acc. Chem. Res. 34, 727–736 (2001).
Kolb, H.C. & Sharpless, K.B. The growing impact of click chemistry on drug discovery. Drug Discov. Today 8, 1128 (2003).
Bianchi, E., Ingenito, R., Simon, R.J. & Pessi, A. Engineering and chemical synthesis of a transmembrane protein: the HCV protease cofactor protein NS4A. J. Am. Chem. Soc. 121, 7698–7699 (1999).
Hunter, C.L. & Kochendoerfer, G.G. Native chemical ligation of hydropobic peptides in lipid bilayer systems. Bioconjug. Chem. 15, 437–440 (2004).
Kochendoerfer, G.G. et al. Total chemical synthesis of the integral membrane protein influenza A virus M2: role of its C-terminal domain in tetramer assembly. Biochemistry 38, 11905–11913 (1999).
Yeo, D.S. et al. Cell-permeable small molecule probes for site-specific labeling of proteins. Chem. Commun. 2870–2871 (2003).
Dawson, P.E., Churchill, M.J., Ghadiri, M.R. & Kent, S.B.H. Modulation of reactivity in native chemical ligation through the use of thiol additives. J. Am. Chem. Soc. 119, 4325–4329 (1997).
Acknowledgements
Some of the work discussed in this review was performed by the authors and was supported by the US National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Muralidharan, V., Muir, T. Protein ligation: an enabling technology for the biophysical analysis of proteins. Nat Methods 3, 429–438 (2006). https://doi.org/10.1038/nmeth886
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth886
This article is cited by
-
Unexpected dynamics in femtomolar complexes of binding proteins with peptides
Nature Communications (2023)
-
mRNA trans-splicing dual AAV vectors for (epi)genome editing and gene therapy
Nature Communications (2023)
-
Maybe you can turn me on: CRISPRa-based strategies for therapeutic applications
Cellular and Molecular Life Sciences (2022)
-
Chemical modification of proteins by insertion of synthetic peptides using tandem protein trans-splicing
Nature Communications (2020)
-
Chemical Digestion of the -Asp-Cys- Sequence for Preparation of Post-translationally Modified Proteins
The Protein Journal (2020)