Abstract
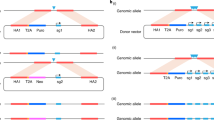
We present a new concept in DNA engineering based on a pipeline of serial recombineering steps in liquid culture. This approach is fast, straightforward and facilitates simultaneous processing of multiple samples in parallel. We validated the approach by generating green fluorescent protein (GFP)-tagged transgenes from Caenorhabditis briggsae genomic clones in a multistep pipeline that takes only 4 d. The transgenes were engineered with minimal disturbance to the natural genomic context so that the correct level and pattern of expression will be secured after transgenesis. An example transgene for the C. briggsae ortholog of lin-59 was used for ballistic transformation in Caenorhabditis elegans. We show that the cross-species transgene is correctly expressed and rescues RNA interference (RNAi)-mediated knockdown of the endogenous C. elegans gene. The strategy that we describe adapts the power of recombineering in Escherichia coli for fluent DNA engineering to a format that can be directly scaled up for genomic projects.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003).
Huh, W.K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Gavin, A.C. et al. Proteome survey reveals modularity of the yeast cell machinery. Nature 440, 631–636 (2006).
Krogan, N.J. et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 440, 637–643 (2006).
Ristevski, S. Making better transgenic models: conditional, temporal, and spatial approaches. Mol. Biotechnol. 29, 153–163 (2005).
Zhang, Y., Buchholz, F., Muyrers, J.P. & Stewart, A.F. A new logic for DNA engineering using recombination in Escherichia coli. Nat. Genet. 20, 123–128 (1998).
Muyrers, J.P., Zhang, Y., Testa, G. & Stewart, A.F. Rapid modification of bacterial artificial chromosomes by ET-recombination. Nucleic Acids Res. 27, 1555–1557 (1999).
Zhang, Y., Muyrers, J.P., Testa, G. & Stewart, A.F. DNA cloning by homologous recombination in Escherichia coli. Nat. Biotechnol. 18, 1314–1317 (2000).
Lee, E.C. et al. A highly efficient Escherichia coli-based chromosome engineering system adapted for recombinogenic targeting and subcloning of BAC DNA. Genomics 73, 56–65 (2001).
Muyrers, J.P., Zhang, Y. & Stewart, A.F. Techniques: recombinogenic engineering—new options for cloning and manipulating DNA. Trends Biochem. Sci. 26, 325–331 (2001).
Court, D.L., Sawitzke, J.A. & Thomason, L.C. Genetic engineering using homologous recombination. Annu. Rev. Genet. 36, 361–388 (2002).
Kittler, R. et al. RNA interference rescue by bacterial artificial chromosome transgenesis in mammalian tissue culture cells. Proc. Natl. Acad. Sci. USA 102, 2396–2401 (2005).
Sarov, M. & Stewart, A.F. The best control for the specificity of RNAi. Trends Biotechnol. 23, 446–448 (2005).
Bao, Z. et al. Automated cell lineage tracing in Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 103, 2707–2712 (2006).
Polanowska, J. et al. Tandem immunoaffinity purification of protein complexes from Caenorhabditis elegans. Biotechniques 36, 778–780, 782 (2004).
Stein, L.D. et al. The genome sequence of Caenorhabditis briggsae: a platform for comparative genomics. PLoS Biol. 1, E45 (2003).
Wang, J. et al. An improved recombineering approach by adding RecA to λ Red recombination. Mol. Biotechnol. 32, 43–54 (2006).
Cardona, S.T. & Valvano, M.A. An expression vector containing a rhamnose-inducible promoter provides tightly regulated gene expression in Burkholderia cenocepacia. Plasmid 54, 219–228 (2005).
Buchholz, F., Ringrose, L., Angrand, P.O., Rossi, F. & Stewart, A.F. Different thermostabilities of FLP and Cre recombinases: implications for applied site-specific recombination. Nucleic Acids Res. 24, 4256–4262 (1996).
Skerra, A. Use of the tetracycline promoter for the tightly regulated production of a murine antibody fragment in Escherichia coli. Gene 151, 131–135 (1994).
Hashimoto-Gotoh, T. & Sekiguchi, M. Mutations of temperature sensitivity in R plasmid pSC101. J. Bacteriol. 131, 405–412 (1977).
Penfold, R.J. & Pemberton, J.M. An improved suicide vector for construction of chromosomal insertion mutations in bacteria. Gene 118, 145–146 (1992).
Praitis, V., Casey, E., Collar, D. & Austin, J. Creation of low-copy integrated transgenic lines in Caenorhabditis elegans. Genetics 157, 1217–1226 (2001).
Maduro, M. & Pilgrim, D. Conservation of function and expression of unc-119 from two Caenorhabditis species despite divergence of non-coding DNA. Gene 183, 77–85 (1996).
Chamberlin, H.M. & Thomas, J.H. The bromodomain protein LIN-49 and trithorax-related protein LIN-59 affect development and gene expression in Caenorhabditis elegans. Development 127, 713–723 (2000).
Simmer, F. et al. Genome-wide RNAi of C. elegans using the hypersensitive rrf-3 strain reveals novel gene functions. PLoS Biol. 1, E12 (2003).
Hope, I.A. et al. Feasibility of genome-scale construction of promoter:reporter gene fusions for expression in Caenorhabditis elegans using a multisite gateway recombination system. Genome Res. 14, 2070–2075 (2004).
Sassi, H.E., Renihan, S., Spence, A.M. & Cooperstock, R.L. Gene CATCHR—gene cloning and tagging for Caenorhabditis elegans using yeast homologous recombination: a novel approach for the analysis of gene expression. Nucleic Acids Res. 33, e163 (2005).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Kamath, R.S., Martinez-Campos, M., Zipperlen, P., Fraser, A.G. & Ahringer, J. Effectiveness of specific RNA-mediated interference through ingested double-stranded RNA in Caenorhabditis elegans. Genome Biol. 2 RESEARCH0002 (2001).
Acknowledgements
We thank M. Valvano for the pSCRhaB2 plasmid; the Caenorhabditis Genetics Center for worm strains; and E. Entchev, L. Pelletier, J. Fu and members of the Genomics Department of TU Dresden for discussions. This work was supported by the European Union and Freistaat Sachsen, by Europäischer Fonds für regionale Entwicklung (EFRE) funding (project code 181) through the Sächsisches Staatsministerium für Wissenschaft und Kunst (SMWK) and Technische Universität Dresden (TUD), and by the Bundesminiterium fuer Bildung und Forschung (BMBF) Proteomics Program.
Author information
Authors and Affiliations
Contributions
M.S. performed the experiments in Figures 1,3–5; M.S., S.S. and A.P. generated the reagents used in Figure 2; A.R. generated the BAC clone map; S.E. performed the bombardment experiments; Y.Z. provided reagents and essential expertise in Red/ET recombineering; M.S, A.A.H. and A.F.S. conceived the project and prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
Y.Z. and A.F.S. are shareholders, and Y.Z. is an employee of Gene Bridges GmbH, which holds the exclusive commercial rights to the Red/ET recombineering methodologies.
Supplementary information
Supplementary Fig. 1
BAC mapping. (PDF 53 kb)
Supplementary Fig. 2
Controlled expression of recombinases from pRedFlp. (PDF 65 kb)
Supplementary Fig. 3
Restriction digest analysis of the 10 tagged transgenes. (PDF 293 kb)
Supplementary Fig. 4
Recombineering outcome with individual clones. (PDF 167 kb)
Supplementary Table 1
Bombardment experiments. (PDF 15 kb)
Rights and permissions
About this article
Cite this article
Sarov, M., Schneider, S., Pozniakovski, A. et al. A recombineering pipeline for functional genomics applied to Caenorhabditis elegans. Nat Methods 3, 839–844 (2006). https://doi.org/10.1038/nmeth933
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth933
This article is cited by
-
Cisplatin toxicity is counteracted by the activation of the p38/ATF-7 signaling pathway in post-mitotic C. elegans
Nature Communications (2023)
-
Modulating p38 MAPK signaling by proteostasis mechanisms supports tissue integrity during growth and aging
Nature Communications (2023)
-
On the benefits of the tryptophan metabolite 3-hydroxyanthranilic acid in Caenorhabditis elegans and mouse aging
Nature Communications (2023)
-
Forward genetic screening identifies novel roles for N-terminal acetyltransferase C and histone deacetylase in C. elegans development
Scientific Reports (2022)
-
Phosphorylated glycosphingolipids essential for cholesterol mobilization in Caenorhabditis elegans
Nature Chemical Biology (2017)