Abstract
Multicolor 3D quantitative imaging of large tissue volumes is necessary to understand tissue development and organization as well as interactions between distinct cell types in situ. However, tissue imaging remains technically challenging, particularly imaging of bone and marrow. Here, we describe a pipeline to reproducibly generate high-dimensional quantitative data from bone and bone marrow that may be extended to any tissue. We generate thick bone sections from adult mouse femurs with preserved tissue microarchitecture and demonstrate eight-color imaging using confocal microscopy without linear unmixing. We introduce XiT, an open-access software for fast and easy data curation, exploration and quantification of large imaging data sets with single-cell resolution. We describe how XiT can be used to correct for potential artifacts in quantitative 3D imaging, and we use the pipeline to measure the spatial relationship between hematopoietic cells, bone matrix and marrow Schwann cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Schroeder, T. Imaging stem-cell-driven regeneration in mammals. Nature 453, 345–351 (2008).
Coutu, D.L. & Schroeder, T. Probing cellular processes by long-term live imaging—historic problems and current solutions. J. Cell Sci. 126, 3805–3815 (2013).
Flores-Figueroa, E., Essers, M. & Bowman, T.V. Hematopoiesis “awakens”: evolving technologies, the force behind them. Exp. Hematol. 44, 101–105 (2016).
Rieger, M.A. & Schroeder, T. Hematopoiesis. Cold Spring Harb. Perspect. Biol. 4, a008250 (2012).
Bianco, P. Bone and the hematopoietic niche: a tale of two stem cells. Blood 117, 5281–5288 (2011).
Beerman, I., Luis, T.C., Singbrant, S., Lo Celso, C. & Méndez-Ferrer, S. The evolving view of the hematopoietic stem cell niche. Exp. Hematol. 50, 22–26 (2017).
Acar, M. et al. Deep imaging of bone marrow shows non-dividing stem cells are mainly perisinusoidal. Nature 526, 126–130 (2015).
Kunisaki, Y. et al. Arteriolar niches maintain haematopoietic stem cell quiescence. Nature 502, 637–643 (2013).
Li, W., Germain, R.N. & Gerner, M.Y. Multiplex, quantitative cellular analysis in large tissue volumes with clearing-enhanced 3D microscopy (Ce3D). Proc. Natl. Acad. Sci. USA 114, E7321–E7330 (2017).
Richardson, D.S. & Lichtman, J.W. Clarifying tissue clearing. Cell 162, 246–257 (2015).
Schnell, U., Dijk, F., Sjollema, K.A. & Giepmans, B.N. Immunolabeling artifacts and the need for live-cell imaging. Nat. Methods 9, 152–158 (2012).
Hama, H. et al. Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain. Nat. Neurosci. 14, 1481–1488 (2011).
Ma, Y.D. et al. Defects in osteoblast function but no changes in long-term repopulating potential of hematopoietic stem cells in a mouse chronic inflammatory arthritis model. Blood 114, 4402–4410 (2009).
Nombela-Arrieta, C. et al. Quantitative imaging of haematopoietic stem and progenitor cell localization and hypoxic status in the bone marrow microenvironment. Nat. Cell Biol. 15, 533–543 (2013).
Yamazaki, S. et al. Nonmyelinating Schwann cells maintain hematopoietic stem cell hibernation in the bone marrow niche. Cell 147, 1146–1158 (2011).
Gruber, H.E. & Ingram, J.A. In Handbook of Histology Methods for Bone and Cartilage (eds. An, Y.H. & Martin, K.L.) 281–286 (Humana Press, 2003).
Kiel, M.J. et al. SLAM family receptors distinguish hematopoietic stem and progenitor cells and reveal endothelial niches for stem cells. Cell 121, 1109–1121 (2005).
Méndez-Ferrer, S. et al. Mesenchymal and haematopoietic stem cells form a unique bone marrow niche. Nature 466, 829–834 (2010).
Wang, L. et al. Identification of a clonally expanding haematopoietic compartment in bone marrow. EMBO J. 32, 219–230 (2013).
Kusumbe, A.P., Ramasamy, S.K. & Adams, R.H. Coupling of angiogenesis and osteogenesis by a specific vessel subtype in bone. Nature 507, 323–328 (2014).
Li, X.-M., Hu, Z., Jorgenson, M.L. & Slayton, W.B. High levels of acetylated low-density lipoprotein uptake and low tyrosine kinase with immunoglobulin and epidermal growth factor homology domains-2 (Tie2) promoter activity distinguish sinusoids from other vessel types in murine bone marrow. Circulation 120, 1910–1918 (2009).
Fischer, A.H., Jacobson, K.A., Rose, J. & Zeller, R. Cryosectioning tissues. CSH Protoc. 2008, 4991 (2008).
Staudt, T., Lang, M.C., Medda, R., Engelhardt, J. & Hell, S.W. 2,2′-thiodiethanol: a new water soluble mounting medium for high resolution optical microscopy. Microsc. Res. Tech. 70, 1–9 (2007).
Bria, A., Iannello, G., Onofri, L. & Peng, H. TeraFly: real-time three-dimensional visualization and annotation of terabytes of multidimensional volumetric images. Nat. Methods 13, 192–194 (2016).
Pietzsch, T., Saalfeld, S., Preibisch, S. & Tomancak, P. BigDataViewer: visualization and processing for large image data sets. Nat. Methods 12, 481–483 (2015).
Gerner, M.Y., Kastenmuller, W., Ifrim, I., Kabat, J. & Germain, R.N. Histo-cytometry: a method for highly multiplex quantitative tissue imaging analysis applied to dendritic cell subset microanatomy in lymph nodes. Immunity 37, 364–376 (2012).
Coutu, D.L., Kokkaliaris, K.D., Kunz, L. & Schroeder, T. Three-dimensional map of non-hematopoetic bone and bone-marrow cells and molecules. Nat. Biotech. http://dx.doi.org/10.1038/nbt.4006 (2017).
Mignone, J.L., Kukekov, V., Chiang, A.-S., Steindler, D. & Enikolopov, G. Neural stem and progenitor cells in nestin-GFP transgenic mice. J. Comp. Neurol. 469, 311–324 (2004).
Maes, C. et al. Osteoblast precursors, but not mature osteoblasts, move into developing and fractured bones along with invading blood vessels. Dev. Cell 19, 329–344 (2010).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Ara, T. et al. Long-term hematopoietic stem cells require stromal cell-derived factor-1 for colonizing bone marrow during ontogeny. Immunity 19, 257–267 (2003).
Bilic-Curcic, I. et al. Visualizing levels of osteoblast differentiation by a two-color promoter-GFP strategy: Type I collagen-GFPcyan and osteocalcin-GFPtpz. Genesis 43, 87–98 (2005).
Huettl, R.-E. & Huber, A.B. In Laser Scanning Microscopy and Quantitative Image Analysis of Neuronal Tissue (eds. Bakota, L. & Brandt, R.) 145–162 (Humana Press, 2014).
Acknowledgements
D.L.C. would like to dedicate this article to Ian B.I. Copland (Emory University), who unexpectedly passed away on June 15th 2015 (http://www.hattrick4ian.com). The authors would like to thank T. Horn, A. Ponti and E.Montani for advice, technical support and useful discussions. This study was supported in part by the SystemsX StemSysMed grant to T. Schroeder.
Author information
Authors and Affiliations
Contributions
D.L.C. started the project and developed the method with K.D.K. and L.K. D.L.C., K.D.K. and L.K. planned and performed experiments and analyses. L.K. programed the XiT software with input from D.L.C. and K.D.K. All authors wrote the manuscript. T.S. funded and supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Experimental workflow for bone imaging.
Processing of individual samples from dissection to imaging can take up to 30 days, depending on the complexity of the immunostaining. However, a single user can easily process 10-20 samples simultaneously until imaging, which is the limiting step. The authors routinely use 0.5-1x104h of imaging time per year using two microscopes.
Supplementary Figure 3 Eight colors immunostaining validation.
Acquisition setup and snapshots from the full bone scan shown in figure 3. The table should be read by rows and shows that the white arrowheads only points at a specific structure in the channel dedicated to this marker/fluorophore (red box). Snapshots from the other channels show lack of bleed-through between channels. When markers are highly correlated, and detected using spectrally similar fluorophores and/or detected simultaneously (e.g. of CD105 and Sca1, magenta box), appearance of bleed-through can be misleading. However, the lack of bleed-through of AlexaFluor 555 into the CF633 channel (Ch7) is confirmed previously (green box). Of note, in the settings for acquisition scan 2 we also illustrate the excitation and emission spectra of Qdot655. As can be seen, the 488nm laser can excite Qdot655 and lead to significant bleed-through into the Chromeo494 channel (Ch4). Hence, the detection settings for Chromeo494 have to be selected carefully.
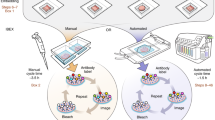
Supplementary Figure 4 Strategies used for complex, indirect immunofluorescent staining using primary antibodies of the same species.
a) Normal indirect immunofluorescent staining is followed by sequential blocking with IgGs and monovalent F(ab)s, and then a second round of immunostaining. b) Alternatively, detection of the primary antibody is directly performed with conjugated monovalent F(ab)s, before a second round of immunostaining (c).
Supplementary Figure 5 Workflow for optimization of complex immunofluorescence staining.
Perfect imaging quality is required for quantification. Thus, if FMO and other controls are not satisfactory repeat the process. When working with well-known antibodies, two to three iterations are typically enough to obtain a satisfactory staining/imaging quality. When working with poorly validated antibodies or for staining with more than five to six colors, up to 10 iterations might be required.
Supplementary Figure 6 Image preprocessing using third party software.
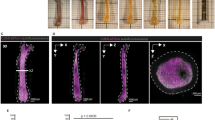
Image processing example for measuring HSPC to bone and neuron distance. a, b) Femur stained for c-Kit, Sca1, lineage markers, collagen 1 and GFAP in four replicates. c, d) c-Kit is used for the initial segmentation (iso). Adjacent cells are often split accurately (white arrows) but splitting errors can occur (white arrowhead). e) After segmentation of collagen 1 and GFAP (not shown), distance transformation (DT) generates a channel representing distances (distance to collagen 1 shown). Statistics of the segmented cells are then exported for analysis with XiT. Typically, 3 - 5 x 104 c-Kit+ cells are segmented per 250μm section. f) A typical image acquisition consists of 1024x1024 pixels fields of view acquired for 4-8 channels at each of 150 or more z planes. To cover the whole organ, 100 to 150 xy positions are acquired. In the example shown, a full femur imaged with four channels consists of 6x105 individual image frames at 1024x1024 resolution. Imaris significantly reduces the size of raw data but segmentation of 2-3 cell types or structures increases file size drastically. Saving segmentation results as a XiT workspace renders data more manageable for fast computational analyses.
Supplementary Figure 7 Validation of XiT gating performed using Imaris.
The Imaris Vantage module was used to display 100 randomly chosen cells from each of four quadrants (Sca1+Lin-, Sca1+Lin+, Sca1-Lin+, Sca1-Lin-). The Vantage module is useful for quick screening of cells and to exclude artefacts.
Supplementary Figure 8 Export function of XiT.
a) The distance between any segmented objects can be measured (here minimum and mean distance of GFAP+ glial cells and collagen 1+ bone matrix). b) Quantifying very large datasets such as 8 color scans presented in figure 3 is greatly facilitated by XiT.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Note 1 and Supplementary Tables 2–3. (PDF 1892 kb)
Life Sciences Reporting Summary
Life Sciences Reporting Summary (PDF 130 kb)
Supplementary Protocol
Supplementary Protocol (PDF 303 kb)
Supplementary Table 1
Details of mounting media and clearing agents tested for this study. (XLSX 11 kb)
Supplementary Table 4
Comparison of data analysis time using Imaris and XiT on various computers. (XLSX 15 kb)
Supplementary Table 5
All antibodies tested in this study. (XLSX 30 kb)
Supplementary Table 6
List and details of all antibodies used in this study. (XLSX 13 kb)
Full bone scan image preprocessing and validation.
Example showing one of the four full femur sections used in figure 5 with c-Kit isosurface (green) and one cKit+Sca1+Lin- cell (magenta) identified using XiT, as well as the distance transform channels from GFAP and collagen 1 (jet color). Markers shown are: c-Kit (green), Sca1 (red), Lin (blue), GFAP (yellow), collagen 1 (grey). (MP4 6774 kb)
Rights and permissions
About this article
Cite this article
Coutu, D., Kokkaliaris, K., Kunz, L. et al. Multicolor quantitative confocal imaging cytometry. Nat Methods 15, 39–46 (2018). https://doi.org/10.1038/nmeth.4503
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4503
This article is cited by
-
Open-source personal pipetting robots with live-cell incubation and microscopy compatibility
Nature Communications (2022)
-
Multispectral confocal 3D imaging of intact healthy and tumor tissue using mLSR-3D
Nature Protocols (2022)
-
Quantitative 3D imaging of the cranial microvascular environment at single-cell resolution
Nature Communications (2021)
-
Mammary tumour cells remodel the bone marrow vascular microenvironment to support metastasis
Nature Communications (2021)
-
Modality attention and sampling enables deep learning with heterogeneous marker combinations in fluorescence microscopy
Nature Machine Intelligence (2021)